Abstract
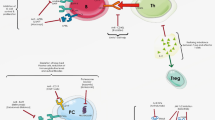
STAT4 is a transcription factor that has been implicated in the pathogenesis of rheumatoid arthritis (RA) and systemic lupus erythematosus (SLE). Recently, several reports has documented that a STAT4 haplotype is associated with RA, SLE and Sjogren’s syndrome. To summarize and review these findings, we conducted a meta-analysis of all relevant reports published before September 2008. Studies on STAT4 rs7574865 single nucleotide polymorphism (SNP) of RA and SLE were identified using PubMed. Meta-analyses were performed for 15,609 patients with RA and 15,793 controls from 14 published studies and for 2,478 patients with SLE and 5,058 controls from 8 published studies. Meta-odds ratios (ORs) and 95% confidence intervals (CIs) based on random effects models were calculated for all available studies. The overall ORs for the minor T allele of STAT4 rs7574865 SNP were 1.27 (95% CI 1.20–1.34) in RA and 1.57 (95% CI 1.44–1.71) in SLE. Asian controls have significantly higher allele frequency (32%) for the minor T allele of STAT4 rs7574865 SNP than population of European origin (22%), however, there was no significant difference of ORs for RA and SLE by ethnicity. No apparent effect of anti-CCP positivity was found in stratified analysis. The risk of STAT4 genotype for SLE was significantly higher than for RA in populations of European origin and Asian. The results of our meta-analysis demonstrated that STAT4 rs7574865 SNP is significantly associated with RA and SLE. In addition to specific alleles of HLA-DRB1, the minor T allele of STAT4 rs7574865 SNP is a common RA risk factor in populations of European origin and Asian.


Similar content being viewed by others
References
Wandstrat A, Wakeland E (2001) The genetics of complex autoimmune diseases: non-MHC susceptibility genes. Nat Immunol 2:802–809. doi:10.1038/ni0901-802
Bowes J, Barton A (2008) Recent advances in the genetics of RA susceptibility. Rheumatology (Oxford) 47:399–402. doi:10.1093/rheumatology/ken005
Yamada R, Yamamoto K (2007) Mechanisms of disease: genetics of rheumatoid arthritis–ethnic differences in disease-associated genes. Nat Clin Pract Rheumatol 3:644–650. doi:10.1038/ncprheum0592
Ikari K, Momohara S, Inoue E et al (2006) Haplotype analysis revealed no association between the PTPN22 gene and RA in a Japanese population. Rheumatology (Oxford) 45:1345–1348. doi:10.1093/rheumatology/kel169
Kyogoku C, Langefeld CD, Ortmann WA et al (2004) Genetic association of the R620W polymorphism of protein tyrosine phosphatase PTPN22 with human SLE. Am J Hum Genet 75:504–507. doi:10.1086/423790
Orozco G, Sanchez E, Gonzalez-Gay MA et al (2005) Association of a functional single-nucleotide polymorphism of PTPN22, encoding lymphoid protein phosphatase, with rheumatoid arthritis and systemic lupus erythematosus. Arthritis Rheum 52:219–224. doi:10.1002/art.20771
Kochi Y, Yamada R, Suzuki A et al (2005) A functional variant in FCRL3, encoding Fc receptor-like 3, is associated with rheumatoid arthritis and several autoimmunities. Nat Genet 37:478–485. doi:10.1038/ng1540
Suzuki A, Yamada R, Chang X et al (2003) Functional haplotypes of PADI4, encoding citrullinating enzyme peptidylarginine deiminase 4, are associated with rheumatoid arthritis. Nat Genet 34:395–402. doi:10.1038/ng1206
Barton A, Bowes J, Eyre S et al (2004) A functional haplotype of the PADI4 gene associated with rheumatoid arthritis in a Japanese population is not associated in a United Kingdom population. Arthritis Rheum 50:1117–1121. doi:10.1002/art.20169
Caponi L, Petit-Teixeira E, Sebbag M et al (2005) A family based study shows no association between rheumatoid arthritis and the PADI4 gene in a white French population. Ann Rheum Dis 64:587–593. doi:10.1136/ard.2004.026831
Martinez A, Sanchez E, Valdivia A et al (2006) Epistatic interaction between FCRL3 and NFkappaB1 genes in Spanish patients with rheumatoid arthritis. Ann Rheum Dis 65:1188–1191. doi:10.1136/ard.2005.048454
Barton A, Thomson W, Ke X et al (2008) Re-evaluation of putative rheumatoid arthritis susceptibility genes in the post-genome wide association study era and hypothesis of a key pathway underlying susceptibility. Hum Mol Genet 17:2274–2279. doi:10.1093/hmg/ddn128
Harley JB, Alarcon-Riquelme ME, Criswell LA et al (2008) Genome-wide association scan in women with systemic lupus erythematosus identifies susceptibility variants in ITGAM, PXK, KIAA1542 and other loci. Nat Genet 40:204–210. doi:10.1038/ng.81
Kobayashi S, Ikari K, Kaneko H et al (2008) Association of STAT4 with susceptibility to rheumatoid arthritis and systemic lupus erythematosus in the Japanese population. Arthritis Rheum 58:1940–1946. doi:10.1002/art.23494
Lee HS, Remmers EF, Le JM et al (2007) Association of STAT4 with rheumatoid arthritis in the Korean population. Mol Med 13:455–460
Orozco G, Alizadeh BZ, Delgado-Vega AM et al (2008) Association of STAT4 with rheumatoid arthritis: a replication study in three European populations. Arthritis Rheum 58:1974–1980. doi:10.1002/art.23549
Palomino-Morales RJ, Rojas-Villarraga A, Gonzalez CI et al (2008) STAT4 but not TRAF1/C5 variants influence the risk of developing rheumatoid arthritis and systemic lupus erythematosus in Colombians. Genes Immun 9:379–382. doi:10.1038/gene.2008.30
Remmers EF, Plenge RM, Lee AT et al (2007) STAT4 and the risk of rheumatoid arthritis and systemic lupus erythematosus. N Engl J Med 357:977–986. doi:10.1056/NEJMoa073003
Taylor KE, Remmers EF, Lee AT et al (2008) Specificity of the STAT4 genetic association for severe disease manifestations of systemic lupus erythematosus. PLoS Genet 4:e1000084. doi:10.1371/journal.pgen.1000084
Zervou MI, Sidiropoulos P, Petraki E et al (2008) Association of a TRAF1 and a STAT4 gene polymorphism with increased risk for rheumatoid arthritis in a genetically homogeneous population. Hum Immunol 69:567–571. doi:10.1016/j.humimm.2008.06.006
Watford WT, Hissong BD, Bream JH et al (2004) Signaling by IL-12 and IL-23 and the immunoregulatory roles of STAT4. Immunol Rev 202:139–156
Kaplan MH, Sun YL, Hoey T et al (1996) Impaired IL-12 responses and enhanced development of Th2 cells in Stat4-deficient mice. Nature 382:174–177
Mathur AN, Chang HC, Zisoulis DG et al (2007) Stat3 and Stat4 direct development of IL-17-secreting Th cells. J Immunol 178:4901–4907
Thierfelder WE, van Deursen JM, Yamamoto K et al (1996) Requirement for Stat4 in interleukin-12-mediated responses of natural killer and T cells. Nature 382:171–174. doi:10.1038/382171a0
Garcia-Sastre A, Biron CA (2006) Type 1 interferons and the virus-host relationship: a lesson in detente. Science 312:879–882. doi:10.1126/science.1125676
Miyagi T, Gil MP, Wang X et al (2007) High basal STAT4 balanced by STAT1 induction to control type 1 interferon effects in natural killer cells. J Exp Med 204:2383–2396. doi:10.1084/jem.20070401
McInnes IB, Schett G (2007) Cytokines in the pathogenesis of rheumatoid arthritis. Nat Rev Immunol 7:429–442. doi:10.1038/nri2094
Ivashkiv LB (2003) Type I interferon modulation of cellular responses to cytokines and infectious pathogens: potential role in SLE pathogenesis. Autoimmunity 36:473–479. doi:10.1080/08916930310001605882
DerSimonian R, Laird N (1986) Meta-analysis in clinical trials. Control Clin Trials 7:177–188. doi:10.1016/0197-2456(86)90046-2
Begg CB, Mazumdar M (1994) Operating characteristics of a rank correlation test for publication bias. Biometrics 50:1088–1101. doi:10.2307/2533446
Egger M, Davey Smith G, Schneider M et al (1997) Bias in meta-analysis detected by a simple, graphical test. BMJ 315:629–634
Duval S, Tweedie R (2000) Trim and fill: a simple funnel-plot-based method of testing and adjusting for publication bias in meta-analysis. Biometrics 56:455–463. doi:10.1111/j.0006-341X.2000.00455.x
Lee YH, Rho YH, Choi SJ et al (2007) The PTPN22 C1858T functional polymorphism and autoimmune diseases–a meta-analysis. Rheumatology (Oxford) 46:49–56. doi:10.1093/rheumatology/kel170
Ueda H, Howson JM, Esposito L et al (2003) Association of the T-cell regulatory gene CTLA4 with susceptibility to autoimmune disease. Nature 423:506–511. doi:10.1038/nature01621
Murphy CA, Langrish CL, Chen Y et al (2003) Divergent pro- and antiinflammatory roles for IL-23 and IL-12 in joint autoimmune inflammation. J Exp Med 198:1951–1957. doi:10.1084/jem.20030896
Kirkham BW, Lassere MN, Edmonds JP et al (2006) Synovial membrane cytokine expression is predictive of joint damage progression in rheumatoid arthritis: a two-year prospective study (the DAMAGE study cohort). Arthritis Rheum 54:1122–1131. doi:10.1002/art.21749
Korman BD, Kastner DL, Gregersen PK et al (2008) STAT4: genetics, mechanisms, and implications for autoimmunity. Curr Allergy Asthma Rep 8:398–403. doi:10.1007/s11882-008-0077-8
Hoey T, Zhang S, Schmidt S et al (2003) Distinct requirements for the naturally occurring splice forms Stat4alpha and Stat4beta in IL-12 responses. EMBO J 22:4237–4248. doi:10.1093/emboj/cdg393
Sigurdsson S, Nordmark G, Garnier S et al (2008) A risk haplotype of STAT4 for systemic lupus erythematosus is over-expressed, correlates with anti-dsDNA and shows additive effects with two risk alleles of IRF5. Hum Mol Genet 17:2868–2876. doi:10.1093/hmg/ddn184
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ji, J.D., Lee, W.J., Kong, K.A. et al. Association of STAT4 polymorphism with rheumatoid arthritis and systemic lupus erythematosus: a meta-analysis. Mol Biol Rep 37, 141–147 (2010). https://doi.org/10.1007/s11033-009-9553-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-009-9553-z