Abstract
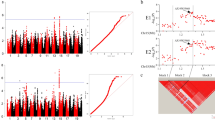
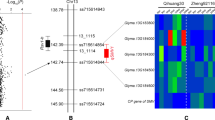
Soybean mosaic disease, caused by soybean mosaic virus (SMV), is one of the most devastating diseases that limit soybean production throughout the world. Soybean isoflavone synthase (IFS) and flavanone 3-hydroxylase (F3H) genes catalyze the production of isoflavones and flavonoids, the increase of which is correlated with increased disease resistance. We have cloned, sequenced, and analyzed the IFS1, IFS2 and F3H genomic regions from 33 Chinese soybean accessions including 16 Glycine soja and 17 Glycine max. High nucleotide diversity and low extent of linkage disequilibrium (LD) in these three genes provided sufficient genetic resolution for association mapping. As a result, a set of single nucleotide polymorphisms (SNPs) with significant (P < 0.05) association to SMV strain SC-3 and SC-7 resistance were discovered in these genes. Among them, the SNP haplotype ‘TCACAACGA-TACA’ in IFS1 gene was found to be extremely significantly (P < 0.01) associated with SMV SC-3 resistance. After 7 days of SC-3 inoculation, the expression level of IFS1 gene in the two SC-3 resistance accessions that have this significant site continued to increased and reach to 30–160 folds high, while in the SC-3 susceptible accession which does not carry the significant site the expression level decreased to near zero. These polymorphisms were corresponding to the trait variance and thus can be considered as the candidate sites for functional molecular markers for future SMV resistance breeding.



Similar content being viewed by others
References
Akashi T, Aoki T, Ayabe S (1999) Cloning and functional expression of a cytochrome P450 cDNA encoding 2-hydroxyisoflavanone synthase involved in biosynthesis of the isoflavonoid skeleton in licorice. Plant Physiol 121:821–828
Andersen JR, Zein I, Wenzel G, Krützfeldt B, Eder J, Ouzunova M, Lübberstedt T (2007) High levels of linkage disequilibrium and associations with forage quality at a Phenylalanine Ammonia-Lyase locus in European maize (Zea mays L.) inbreds. Theor Appl Genet 114:307–319
Aranzana MJ, Kim S, Zhao K, Bakker E, Horton M, Jakob K, Lister C, Molitor J, Shindo C, Tang C, Toomajian C, Traw B, Zheng H, Bergelson J, Dean C, Marjoram P, Nordborg M (2005) Genome-wide association mapping in Arabidopsis identifies previously known flowering time and pathogen resistance genes. PLoS Genet 1:e60
Ashfield T, Danzer JR, Held D, Clayton K, Keim P, Saghai Maroof MA, Webb DM, Innes RW (1998) Rpg1, a soybean gene effective against races of bacterial blight, maps to a cluster of previously identified disease resistance genes. Theor Appl Genet 96:1013–1021
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635
Cheng H, Yu O, Yu D (2008) Polymorphisms of IFS1 and IFS2 gene are associated with isoflavone concentrations in soybean seeds. Plant Sci 175:505–512
Corder EH, Saunders AM, Risch NJ, Strittmatter WJ, Schmechel DE, Gaskell PC, Rimmler JB, Locke PA, Conneally PM, Schmader KE (1994) Protective effect of apolipoprotein E type 2 allele for late onset Alzheimer disease. Nat Genet 7:180–184
Dixon RA (2001) Natural products and plant disease resistance. Nature 411:843–847
Dixon RA, Paiva NL (1995) Stress-induced phenylpropanoid metabolism. Plant Cell 7:1085–1097
Edwards D, Forster J, Chagné D, Batley J (2007) What are SNPs. In: Oraguzie N, Rikkerink E, Gardiner S, Silva HD (eds) Association mapping in plants. Springer, Heidelberg, pp 41–52
Falush D, Stephens M, Pritchard JK (2007) Inference of population structure using multilocus genotype data: dominant markers and null alleles. Mol Ecol Notes 7:574–578
Flint-Garcia SA, Thornsberry JM, Edwards B (2003) Structure of linkage disequilibrium in plants. Annu Rev Plant Biol 54:357–374
Fu S (2006) Construction of soybean genetic map by SSR and its application in soybean disease-resistance and insect-resistance genes mapping. PhD thesis, Nanjing Agricultural University, Nanjing
Fu S, Zhan Y, Zhi H, Gai J, Yu D (2006) Mapping of SMV resistance gene Rsc-7 by SSR markers in soybean. Genetica 128:63–69
Graham TL, Graham MY (1996) Signaling in soybean phenylpropanoid responses (dissection of primary, secondary, and conditioning effects of light, wounding, and elicitor treatments). Plant Physiol 110:1123–1133
Hill JH, Bailey TB, Benner HI, Tachibana H, Durand DP (1987) Soybean mosaic virus: effects of primary disease incidence on yield and seed quality. Plant Dis 71:237–239
Hymowitz T (2004) Speciation and cytogenetics. In: Boerma HR, Specht JE (eds) Soybeans: improvement, production and uses. American society of agromomy, Crop science society of America and Soil science society of America, Madison, pp 97–136
Jung W, Yu O, Lau SM, O’Keefe DP, Odell J, Fader G, McGonigle B (2000) Identification and expression of isoflavone synthase, the key enzyme for biosynthesis of isoflavones in legumes. Nat Biotechnol 18:208–212
Keim P, Olson TC, Shoemaker RC (1988) A rapid protocol for isolating soybean DNA. Soyb Genet Newsl 15:150–152
Kerem B, Rommens JM, Buchanan JA, Markiewicz D, Cox TK, Chakravarti A, Buchwald M, Tsui LC (1989) Identification of the cystic fibrosis gene: genetic analysis. Science 245:1073–1080
Kiihl RAS, Hartwig EE (1979) Inheritance of reaction to soybean mosaic virus in soybeans. Crop Sci 19:372
Knowler WC, Williams RC, Pettitt DJ, Steinberg AG (1988) Gm3; 5, 13, 14 and type 2 diabetes mellitus: an association in American Indians with genetic admixture. Am J Hum Genet 43:520–526
Ma G, Chen P, Buss GR, Tolin SA (1995) Genetic characteristics of two genes for resistance to soybean mosaic virus in PI486355 soybean. Theor Appl Genet 91:907–914
Mansfield JW (2000) Antimicrobial compounds and resistance: the role of phytoalexins and phytoanticipins. In: Slusarenko A, Fraser RSS, Loon LCv (eds) Mechanisms of resistance to plant diseases. Kluwer, Dordrecht, pp 325–370
Moffat AS (2001) Finding new ways to fight plant diseases. Science 292:2270–2273
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Pu ZC, Cao Q, Xie BT (1983) Resistance in soybean varieties to six strains of soybean mosaic virus. J Nanjing Agric Coll 3:41–45
Ralston L, Subramanian S, Matsuno M, Yu O (2005) Partial reconstruction of flavonoid and isoflavonoid biosynthesis in yeast using soybean type I and type II chalcone isomerases. Plant Physiol 137:1375–1388
Ravel C, Praud S, Canaguier A, Dufour P, Giancola S, Balfourier F, Chalhoub B, Brunel D, Linossier L, Dardevet M (2007) DNA sequence polymorphisms and their application to bread wheat quality. Euphytica 158:331–336
Rozas J, Sanchez-DelBarrio JC, Messeguer X, Rozas R (2003) DnaSP, DNA polymorphism analyses by the coalescent and other methods. Bioinformatics 19:2496–2497
Springob K, Nakajima J, Yamazaki M, Saito K (2003) Recent advances in the biosynthesis and accumulation of anthocyanins. Nat Prod Rep 20:288–303
Steele CL, Gijzen M, Qutob D, Dixon RA (1999) Molecular characterization of the enzyme catalyzing the aryl migration reaction of isoflavonoid biosynthesis in soybean. Arch Biochem Biophys 367:146–150
Subramanian S, Hu X, Lu G, Odelland JT, Yu O (2004) The promoters of two isoflavone synthase genes respond differentially to nodulation and defense signals in transgenic soybean roots. Plant Mol Biol 54:623–639
Subramanian S, Stacey G, Yu O (2007) Distinct, crucial roles of flavonoids during legume nodulation. Trends Plant Sci 12:282–285
Szalma SJ, Buckler ES, Snook ME, McMullen MD (2005) Association analysis of candidate genes for maysin and chlorogenic acid accumulation in maize silks. Theor Appl Genet 110:1324–1333
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Wang XQ (2000) Strain identification of soybean mosaic virus in Huang-Huai and Middle and Lower Changjiang River Valleys and the resistant sources as well as the inheritance of resistance in soybeans. Master thesis, Nanjing Agricultural University, Nanjing
Wang Y, Gai J (2002) Study on the ecological regions of soybean in China II. Ecological environment and representative varieties. Ying Yong Sheng Tai Xue Bao 13:71–75
Weir B (1996) Genetic data analysis. Publishing House of Agricultural Science, Beijing
Yang YL (2002) Classification and distribution of strains of soybean mosaic virus in the Middle and Lower Changjiang Valleys and resistance to soybean mosaic virus in soybean. Master thesis, Nanjing Agricultural University, Nanjing
Yu J, Buckler E (2006) Genetic association mapping and genome organization of maize. Curr Opin Biotechnol 17:155–160
Yu O, McGonigle B (2005) Metabolic engineering of isoflavone biosynthesis. Adv Agron 86:147–190
Zhao K, Aranzana MJ, Kim S, Lister C, Shindo C, Tang C, Toomajian C, Zheng H, Dean C, Marjoram P, Nordborg M (2007) An Arabidopsis example of association mapping in structured samples. PLoS Genet 3:e4
Zhu Y, Song Q, Hyten D, Tassell CV, Matukumalli L, Grimm D, Hyatt S, Fickus E, Young N, Cregan P (2003) Single-nucleotide polymorphisms in soybean. Genetics 163:1123–1134
Acknowledgments
We thank Drs. Zhiwu Zhang and Ed Buckler of Cornell University for their advice on association analysis and help on TASSEL software. We also thank two anonymous reviewers for their critical comments and helpful discussions. This work was funded in part by National 973 Project (No. 2004CB117206), National 863 Projects (No. 2006AA10Z1C1, No. 2006AA10A111), and National Natural Science Foundation of China (No. 30771362).
Author information
Authors and Affiliations
Corresponding author
Additional information
The authors Hao Cheng and Hua Yang contributed equally to this paper.
Rights and permissions
About this article
Cite this article
Cheng, H., Yang, H., Zhang, D. et al. Polymorphisms of soybean isoflavone synthase and flavanone 3-hydroxylase genes are associated with soybean mosaic virus resistance. Mol Breeding 25, 13–24 (2010). https://doi.org/10.1007/s11032-009-9305-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-009-9305-8