Abstract
The main study’s purpose is to detect novel natural products (NPs) that are potentially selective MAO-B inhibitors and, additionally, to computationally reposition the marketed drugs with a new therapeutic role for Parkinson’s disease. To reach the goals, 3D similarity search, docking, ADMETox, and drug repurposing approaches were employed. Thus, an unbiased benchmarking dataset was built including selective and nonselective inhibitors for MAO-B compliant with both ligand- and structure-based virtual screening approaches. A retrospective and prospective mining scenario was applied to SPECS NP and DrugBank databases to detect novel scaffolds with potential benefits for Parkinson’s disease patients. Out of the three best selected natural products, cardamomin showed excellently predicted drug-like properties, superior pharmacological profile, and specific interactions with MAO-B active site, indicating a potential selectivity over MAO-B. Two marketed drugs, fenamisal and monobenzone, were proposed as promising candidates repurposed for Parkinson’s disease. The application of shape, physicochemical, and electrostatic similarity searches protocol emerged as a plausible solution to explore MAO-B inhibitors selectivity. This protocol might serve as a rewarding tool in early drug discovery and can be extended to other protein targets.
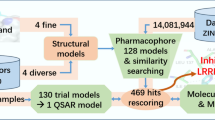
Graphic abstract















Similar content being viewed by others
Abbreviations
- AUC:
-
Area under the curve
- BMF:
-
Bemis–Murcko framework
- CG4:
-
Chemgauss4
- CS:
-
ComboScore
- CoS:
-
ColorScore
- CoT:
-
ColorTanimoto
- FCoTv:
-
FitColorTversky
- FTv:
-
FitTversky
- FTvC:
-
FitTverskyCombo
- FRED:
-
Flexible ligand–rigid protein docking
- HBA:
-
Hydrogen bond acceptor
- HBD:
-
Hydrogen bond donor
- HRM:
-
Harmine
- LB:
-
Ligand based
- MAO-A:
-
Monoamine oxidase A
- MAO-B:
-
Monoamine oxidase B
- MAOIs:
-
Monoamine oxidase inhibitors
- SPECS NP:
-
SPECS natural products
- O:
-
Overlap
- PDB:
-
Protein Data Bank
- q1 :
-
Query 1
- q2 :
-
Query 2
- RBN:
-
Rotatable bond
- RMSD:
-
Root mean squared deviation
- ROC:
-
Receiver operating characteristic
- ROCS:
-
Rapid overlay of chemical structures
- RCoTv:
-
RefColorTversky
- RTv:
-
RefTversky
- RTvC:
-
RefTverskyCombo
- SB:
-
Structure based
- SAG:
-
Safinamide
- ShT:
-
ShapeTanimoto
- SCo:
-
ScaledColor
- TC:
-
TanimotoCombo
- VS:
-
Virtual screening
References
Athauda D, Foltynie T (2018) Drug repurposing in Parkinson’s disease. CNS Drugs 32(8):747–761. https://doi.org/10.1007/s40263-018-0548-y
Mittal R, Debs LH, Patel AP, Nguyen D, Patel K, O’Connor G, Grati M, Mittal J, Yan D, Eshraghi AA, Deo SK, Daunert S, Liu XZ (2017) Neurotransmitters: the critical modulators regulating gut-brain axis. J Cell Physiol 232:2359–2372. https://doi.org/10.1002/jcp.25518
Tripathi RKP, Ayyannan SR (2019) Monoamine oxidase-B inhibitors as potential neurotherapeutic agents: an overview and update. Med Res Rev. https://doi.org/10.1002/med.21561
Ugun-Klusek A, Theodosi TS, Fitzgerald JC, Burté F, Ufer C, Boocock DJ, Yu-Wai-Man P, Bedford L, Billet EE (2019) Monoamine oxidase-A promotes protective autophagy in human SH-SY5Y neuroblastoma cells through Bcl-2 phosphorylation. Redox Biol 20:167–181. https://doi.org/10.1016/j.redox.2018.10.003
Yeung AWK, Georgieva MG, Atanasov AG, Tzvetkov NT (2019) Monoamine oxidases (MAOs) as privileged molecular targets in neuroscience: research literature analysis. Front Mol Neurosci 12:143. https://doi.org/10.3389/fnmol.2019.00143
Finberg JPM, Rabey JM (2016) Inhibitors of MAO-A and MAO-B in psychiatry and neurology. Front Pharmacol 7:340. https://doi.org/10.3389/fphar.2016.00340
Bette S, Shpiner DS, Singer C, Moore H (2018) Safinamide in the management of patients with Parkinson’s disease not stabilized on levodopa: a review of the current clinical evidence. Ther Clin Risk Manag 14:1737–1745. https://doi.org/10.2147/TCRM.S139545
Tanrikulu Y, Krüger B, Proschak E (2013) The holistic integration of virtual screening in drug discovery. Drug Discov Today 18(7–8):358–364. https://doi.org/10.1016/j.drudis.2013.01.007
Lounkine E, Keiser MJ, Whitebread S, Mikhailov D, Hamon J, Jenkins JL, Lavan P, Weber E, Doak AK, Côté S, Shoichet BK, Urban L (2012) Large-scale prediction and testing of drug activity on side-effect targets. Nature 486(7403):361–367. https://doi.org/10.1038/nature11159
Crisan L, Pacureanu L, Bora A, Avram S, Kurunczi L, Simon Z (2013) QSAR study and molecular docking on indirubin inhibitors of Glycogen Synthase Kinase-3. Cent Eur J Chem 11(1):63–77. https://doi.org/10.2478/s11532-012-0133-z
Crisan L, Avram S, Pacureanu L (2017) Pharmacophore-based screening and drug repurposing exemplified on glycogen synthase kinase-3 inhibitors. Mol Divers 21(2):385–405. https://doi.org/10.1007/s11030-016-9724-5
Stumpfe D, Lounkine E, Bajorath J (2011) Molecular test systems for computational selectivity studies and systematic analysis of compound selectivity profiles. Methods Mol Biol 672:503–516. https://doi.org/10.1007/978-1-60761-839-3_20
Bautista-Aguiler OM, Esteban G, Bolea I, Nikolic K, Agbaba D, Moraleda I, Iriepa I, Samadi A, Soriano E, Unzeta M, Marco-Contelles J (2014) Design, synthesis, pharmacological evaluation, QSAR analysis, molecular modeling and ADMET of novel donepezil-indolyl hybrids as multipotent cholinesterase/monoamine oxidase inhibitors for the potential treatment of Alzheimer’s disease. Eur J Med Chem 75:82–95. https://doi.org/10.1016/j.ejmech.2013.12.028
Ogunrombi MO, Malan SF, Terre’Blanche G, Castagnoli N Jr, Bergh JJ, Petzer JP (2008) Structure-activity relationships in the inhibition of monoamine oxidase B by 1-methyl-3-phenylpyrroles. Bioorg Med Chem 16(5):2463–2472. https://doi.org/10.1016/j.bmc.2007.11.059
Ferreira LLG, Santos RN, Oliva G, Andricopulo AD (2015) Molecular docking and structure-based drug design strategies. Molecules 20:13384–13421. https://doi.org/10.3390/molecules200713384
Pacureanu L, Avram S, Crisan L (2020) Comprehensive investigation of selectivity landscape of glycogen synthase kinase-3 inhibitors. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2020.1747544
Bento AP, Gaulton A, Hersey A, Bellis LJ, Chambers J, Davies M, Kruger FA, Light Y, Mak L, McGlinchey S, Nowotka M, Papadatos G, Santos R, Overington JP (2014) The ChEMBL bioactivity database: an update. Nucleic Acids Res 42:D1083–D1090. https://doi.org/10.1093/nar/gkt1031
Mysinger MM, Carchia M, Irwin JJ, Shoichet BK (2012) Directory of useful decoys, enhanced (DUD-E): better ligands and decoys for better benchmarking. J Med Chem 55(14):6582–6594. https://doi.org/10.1021/jm300687e
Lagarde N, Zagury J-F, Montes M (2015) Benchmarking data sets for the evaluation of virtual ligand screening methods: review and perspectives. J Chem Inf Model 55:1297–1307. https://doi.org/10.1021/acs.jcim.5b00090
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (1997) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 23(1):3–25. https://doi.org/10.1016/S0169-409X(96)00423-1
Bemis GW, Murcko MA (1996) The properties of known drugs. 1. Molecular frameworks. J Med Chem 39(15):2887–2893. https://doi.org/10.1021/jm9602928
Good AC, Oprea TI (2008) Optimization of CAMD techniques 3. Virtual screening enrichment studies: a help or hindrance in tool selection. J Comput Aided Mol Des 22(3–4):168–178. https://doi.org/10.1007/s10822-007-9167-2
Réau M, Langenfeld F, Zagury J-F, Lagarde N, Montes M (2018) Decoys selection in benchmarking datasets: overview and perspectives. Front Pharmacol 9:11. https://doi.org/10.3389/fphar.2018.00011
Hawkins PCD, Skillman AG, Warren GL, Ellingson BA, Stahl MT (2010) Conformer generation with OMEGA: algorithm and validation using high quality structures from the protein databank and Cambridge structural database. J Chem Inf Model 50(4):572–584. https://doi.org/10.1021/ci100031x
Halgren TA (1999) MMFF VI. MMFF94s option for energy minimization studies. J Comput Chem 20(7):720–729. https://doi.org/10.1002/(SICI)1096-987X(199905)20:7%3c720:AID-JCC7%3e3.0.CO;2-X
Hawkins PCD, Skillman AG, Nicholls A (2007) Comparison of shape-matching and docking as virtual screening tools. J Med Chem 50(1):74–82. https://doi.org/10.1021/jm0603365
Nel MS, Petzer A, Petzer JP, Legoabe LJ (2016) 2-Heteroarylidene-1-indanone derivatives as inhibitors of monoamine oxidase. Bioorg Chem 69:20–38. https://doi.org/10.1016/j.bioorg.2016.09.004
Binda C, Wang J, Pisani L, Caccia C, Carotti A, Salvati P, Edmondson DE, Mattevi A (2007) Structures of human monoamine oxidase B complexes with selective noncovalent inhibitors: safinamide and coumarin analogs. J Med Chem 50:5848–5852. https://doi.org/10.1021/jm070677y
McGann M (2011) FRED pose prediction and virtual screening accuracy. J Chem Inf Model 51(3):578–596. https://doi.org/10.1021/ci100436p
McGann M (2012) FRED and HYBRID docking performance on standardized datasets. J Comput Aided Mol Des 26(8):897–906. https://doi.org/10.1007/s10822-012-9584-8
Hanley JA, McNeil BJ (1982) The meaning and use of the area under a receiver operating characteristic (ROC) curve. Radiology 143(1):29–36. https://doi.org/10.1148/radiology.143.1.7063747
Xia J, Jin H, Liu Z, Zhang L, Wang XS (2014) An unbiased method to build benchmarking sets for ligand-based virtual screening and its application to GPCRs. J Chem Inf Model 54(5):1433–1450. https://doi.org/10.1021/ci500062f
Avram SI, Crisan L, Bora A, Pacureanu LM, Avram S, Kurunczi L (2013) Retrospective group fusion similarity search based on eROCE evaluation metric. Bioorg Med Chem 21(5):268–1278. https://doi.org/10.1016/j.bmc.2012.12.041
Cabrera-Pérez MÁ, Pham-The H (2018) Computational modelling of human oral bioavailability: What will be next? Expert Opin Drug Discov 13(6):509–521. https://doi.org/10.1080/17460441.2018.1463988
Daina A, Michielin O, Zoete V (2017) SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep 7(1):42717. https://doi.org/10.1038/srep42717
Talevi A (2015) Multi-target pharmacology: possibilities and limitations of the “skeleton key approach” from a medicinal chemist perspective. Front Pharmacol 6:205. https://doi.org/10.3389/fphar.2015.00205
Rohrer SG, Baumann K (2009) Maximum unbiased validation (MUV) data sets for virtual screening based on PubChem bioactivity data. J Chem Inf Model 49(2):169–184. https://doi.org/10.1021/ci8002649
Kumar A, Zhang KYJ (2018) Advances in the development of shape similarity methods and their application in drug discovery. Front Chem 6:315. https://doi.org/10.3389/fchem.2018.00315
Lee Y-H, Lin Y-C, Feng C-H, Tseng W-L, Lua C-Y (2017) A derivatization-enhanced detection strategy in mass spectrometry: analysis of 4-hydroxybenzoates and their metabolites after keratinocytes are exposed to UV radiation. Sci Rep 7:39907. https://doi.org/10.1038/srep39907
Son S, Ma J, Kondou Y, Yoshimura M, Yamashita E, Tsukihara T (2008) Structure of human monoamine oxidase A at 2.2-Å resolution: the control of opening the entry for substrates/inhibitors. Proc Natl Acad Sci USA 105(15):5739–5744. https://doi.org/10.1073/pnas.0710626105
Hubálek F, Binda C, Khalil A, Li M, Mattevi A, Castagnoli N, Edmondson DE (2005) Demonstration of isoleucine 199 as a structural determinant for the selective inhibition of human monoamine oxidase B by specific reversible inhibitors. J Biol Chem 280(16):15761–15766. https://doi.org/10.1074/jbc.M500949200
Geha RM, Chen K, Wouters J, Ooms F, Shih JC (2002) Analysis of conserved active site residues in monoamine oxidase A and B and their three-dimensional molecular modeling. J Biol Chem 27(19):17209–17216. https://doi.org/10.1074/jbc.M110920200
Wishart DS, Knox C, Guo AC, Cheng D, Shrivastava S, Tzur D, Gautam B, Hassanali M (2008) DrugBank: a knowledgebase for drugs, drug actions and drug targets. Nucleic Acids Res 36:D901–D906. https://doi.org/10.1093/nar/gkm958
Acknowledgements
The authors thank ChemAxon Ltd. (Marvin Sketch and Instant JChem), OpenEye Ltd., and BIOVIA software Inc. (Discovery Studio Visualizer) for providing academic license, and Dr. Simona Funar-Timofei, “Coriolan Dragulescu” Institute of Chemistry for providing access to STATISTICA software. The authors wish to thank Schrödinger Inc. for providing an academic trial license to complete the calculations for this paper. Project No. 1.2 of the “Coriolan Dragulescu” Institute of Chemistry, Romanian Academy, Timisoara, financially supported the current work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors indicate no potential conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Crisan, L., Istrate, D., Bora, A. et al. Virtual screening and drug repurposing experiments to identify potential novel selective MAO-B inhibitors for Parkinson’s disease treatment. Mol Divers 25, 1775–1794 (2021). https://doi.org/10.1007/s11030-020-10155-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11030-020-10155-6