Abstract
The orchestration of the events in the cell during the progression of the cell cycle is modulated by various phenomenon which are regulated by structural modules of the cell. The nucleus is a major hub for all these regulatory units which harbour the nuclear matrix, matrix proteins and chromatin. The histone modifications etch a complex code on the chromatin and the matrix proteins in consort with the histone code regulate the gene expression. SMAR1 is a matrix attachment region binding protein that interacts with chromatin modulators like HDAC1, Sin3A and causes chromatin condensation. SMAR1 modulates the chromatin at the Vβ locus and plays a prominent role in V(D)J recombination. Such indispensable function of SMAR1 by the modulation of chromatin in the context of malignancy and V(D)J recombination emphasizes that MAR binding proteins regulate the complex events of the cell and perturbed expression causes disease conditions.
Similar content being viewed by others
Introduction
During the progression of cell cycle, the cell encounters numerous checkpoints where it must decide to go for proliferation, differentiation or apoptosis. For all these processes of the cell to be approached with inexorable certainty, the regulation of the gene expression holds the master key. The central hub for all these regulatory events is the nucleus [1]. The nucleus is a heterogeneous structure with a mosaic of nucleolus, inter-chromatin regions and condensed chromatin, dispersed in the nuclear ground substance traditionally called the ‘nuclear matrix’. Inside the nuclear matrix, the matrix proteins and the matrix attachment region (MAR) binding proteins in consort with the chromatin bound histones modulate the chromatin for the regulation and orchestration of various events of the cell cycle.
Nuclear matrix and matrix proteins
The in situ nuclear matrix is depicted as a dynamic fibrogranular structure in the inter-chromatininic region with at least two main structural as well as functional domains, the chromatin domains and the ribonucleoprotein domains [ 2, 3]. The isolated nuclear matrix is a tripartite structure consisting of 10–25% nuclear proteins and tightly bound DNA and RNA. The nuclear matrix provides the structural milieu for the exposure of functional loops of chromatin and organization of active replicational or transcriptional complex. The potential continuous nature of the matrix system suggests that these two functional domains are interlinked, and matrix serves in an integrative and regulatory role for cascade and assembly of line of regulatory events. The dual aspect of nuclear matrix organization inside the nucleus dictates the heterogeneity of the matrix at both organizational level and hence the functional levels. The functions of the nuclear matrix ranges from DNA loop attachment site to DNA replication [4, 5], to transcriptional association [6], to transcription, to RNA splicing [7], to viral associations and their associated functions [8] and to a vast number of regulatory proteins involved in the functioning and regulation of these functions. One of the best characterized functions of the nuclear matrix is DNA replication.
The nuclear matrix fraction that is formed following nuclease, salt and detergent extraction of the cell nuclei comprises the non-histone nuclear matrix proteins. The nuclear matrix proteins organize the functional domains of the nuclear matrix and form the DNA replication foci [9], the transcription foci [10, 11], coiled bodies [10, 12–15], speckled domains [16] and PML bodies [17]. These nuclear matrix proteins also dictate the cellular type and state of differentiation [18]. Several laboratories have shown that the nuclear matrix protein composition of a malignant cell varies from that of a normal cell. For instance, prostate tumours have nuclear matrix proteins that are present neither in a normal tissue nor in benign prostrate hyperplasia [19]. Six nuclear matrix proteins have been reported in human colon adenocarcinoma tumour samples that were absent in normal colon tissue [20, 21] which in way can be reasoned that these altered nuclear matrix proteins contribute to the malignant state of the cell. Thus, in general, the nuclear matrix with its component proteins, acts as a hub, to integrate various signals and modulate different functional domains in a temporal manner to regulate various events of the cell cycle.
It is with the help of the nuclear matrix and the matrix proteins that the chromatin, which is tailored with the regulatory units called genes, is organized in a hierarchical fashion. The 147 bp DNA is wrapped around the histone octamer (two each of H2A, H2B and two each of H3–H4 dimer) to form a 10-nm fibre [22, 23]. The histone tetramers are organized and packed with the DNA in the form of ‘beads-on-a-string’ and the histone H1, the linker histone that binds to the linker DNA form the 30 nm fibre of chromatin [24–26], which form loops and domains of size ranging from 5 to 200 kb [27]. These loops are believed to be anchored to the supporting nuclear matrix by proteins located at the base of the loops. The regions of the DNA which bind to the nuclear matrix are the scaffold or matrix attachment regions (SARs or MARs), and the proteins that help in the anchorage of the S/MARs to the nuclear matrix are the matrix attachment region binding proteins (MARBPs).
MARs, MAR binding proteins and viruses
Matrix attachment regions (MARs) are sequences of 200–500 bp AT rich segments of the regulatory regions of the gene loci harbouring topoisomerase II cleavage sites [13, 28], supposed to punctuate chromosomal DNA into functional units of topologically constrained loops domains. MARs can be viewed as boundary elements shielding and insulating genes located between them from cis acting elements and torsional stress from neighbouring chromatin domains [29, 30]. About 60% of the MARs colocalize with transcriptional enhancers and 30% of the simple MARs cohabit with origins of replication of the mammalian genome. MAR elements interfere with gene expression when placed between promoter and enhancers. MARs are proposed to enhance the expression of reporter genes by forming a domain and insulation from the position effect variegation of neighbourhood domains [29, 31], by the recruitment of topoisomerase II by the MAR sequence [32], facilitates the displacement of histone H1 repressor from the chromatin and replacement by HMG-I(Y) activator [33], and/or by propagating the enhancer-induced local alteration in chromatin structure to the promoter [34].
The DNA replication units are found anchored to the nuclear matrix and MARs are found to cohabit with the origin of replication sites. From the sequence analysis of origins of replication, ors of the putative origin of replication of monkey genome, autonomously replicating sequences of Saccharomyces cerevisiae [35], and studies on Drosophila scaffold attachment regions in yeast [36, 37] suggested that origins of replication in mammalian cells are in close proximity to MARs and or the structure of DNA at the sites of origins might allowed them to act as MARs. MARs and origin of replications (ORIs) contain AT rich tracts [38]. These AT tracts also form the part of palindromes which facilitate the unwinding of the local regions in the DNA. The inverted repeats or palindromes are able to convert to cruciform structures [39], which can act like efficient reservoirs upon torsional strain in DNA. The AT rich tracts facilitate the interaction with HMG1 and 2, topoisomerase II and Histone H1 [40–42]. These AT rich tracts which harbour DNA unwinding elements can be the sites of origins of DNA replication and can act as MAR/ORI elements. MARs induce the conformation of DNaseI hypersensitive sites in the chromatin. Generally, the regions of active MAR/ORIs are free of nucleosomes and harbour poor nucleosome positioners [43]. MAR regions also act as representative locus control regions (LCR). MARs aid in the cell type specific gene expression by their cohabitation and synergistic collaboration with enhancers towards maintaining some domains into condensed inactive structures and other into transcriptional active looped domains modulated by topoisomerase II.
The proteins of the nuclear matrix are categorized based on the function and affinity to DNA loops as (a) transcription factors with stringent DNA sequence specificity like c-Myc, c-Myb, NMP-1, NMP-2, ATF family, CCAAT etc. [44, 45], (b) factors which bind to the DNA non-stringently exhibiting a cooperative binding like SATB1 [46], SMAR1 [47], SAF-A [48], Matrin 3, HMG1 & 2 [49], Histone H1 [50] etc., (c) factors which does not bind to the DNA, but act as adaptors by binding to other matrix proteins like Rb [51], progesterone [52] etc., (d) some enzymes like CK II [53], HDACs [54] etc., does not bind to the DNA, but are the components of the nuclear matrix. Thus, validating the proteome of the nuclear matrix, it is regarded that nuclear matrix is a sink for all proteins involved in replication, repair, transcription, recombination and splicing, with the interdigitation and crosstalk within these different factors. These factors bind the MARs to dictate the fate of the gene, to be either in the active loop domain or in the inactive domain, dictating the fate of the cell during each successive division.
It has also been reported that the nuclear matrix is involved in the regulation of viral processes. Viruses after entering the cell uses host cell machinery for replication, transcription as well as translation. Retroviruses, a family of RNA viruses after entering the cell, make a copy of DNA and target it for integration into the host genome. Several reports have demonstrated that the nuclear matrix is involved in retroviral DNA replication, transcription and virion assembly [55–57]. An investigation on the proviral sequence integration site preferences has established that the proviral sequence integration is a non-random process exemplified by human immunodeficiency viruses and Murine leukaemia viruses. Cis elements, such as topo II cleavage sites, Alu repeats and MARs are thought to be targets of retroviral sequence integrations [58]. Genome wide analysis revealed that approximately 11% of the MuLV proviral integration sites are flanked by DNaseI hypersensitive sites [59]. Small nuclear genome-containing viruses like HPV16, HBV, SV40, and HTLV-1 have been shown to integrate near MARs [60]. Studies by Kulkarni et al. [61] have shown that HIV proviral sequences integrate next to MAR elements and 485 out of 524 integrations were flanked by MARs and 35.3% of these MARs elements were found within 1 kb of the integration sites (91 MAR elements upstream and 80 MAR elements downstream), while 41.8% of the elements lie at a distance of 2–3 kb of integration sites (106 elements upstream and 97 elements downstream). The number of MARs upstream and downstream of the integration sites on each chromosome are analysed and represented in the Fig. 1. The presence of the MAR elements next to the site of integration might play a critical role in viral gene regulation and transcription. Thus, the nuclear matrix and the matrix components, especially the chromatin domain of the matrix, play a significant role in the integration events of the proviral sequences, thus modulating the activity the virus as well as the cell.
The location of MARs downstream and upstream of the HIV-1 integration sites for individual chromosomes. The distance around the integrations is shown using different colours. The number of integration sites is shown in all the 23 chromosomes (Adopted from Kulkarni et al. [61].)
Chromatin and histones
The histone code, which is being encrypted on the genes, is based on the modifications of the bound histones’ tails whose length is around 25 amino acids, provides an extra layer for the gene regulation. The modifications on these tails include phosphorylation, acetylation, methylation, ubiquitination and ADP-ribosylation [62]. These modifications on the tail dictate the gene either to be in active form or to be repressed forever by forming the heterochromatin. Modifications like phosphorylation and acetylation are reversible, while methylation in most of the cases is irreversible.
Acetylation is one of the major modifications that generally happen at the ε-amino group of the lysine residues of the histones by 10 different histone acetyl transferases (HATs) [63, 64]. Lysines of 1–6 in each histone can undergo acetylation. All the acetylations generally decondense the chromatin and activate the transcription [65]. In some cases, these acetylations crosstalk with phosphorylation and methylation of the corresponding histones as like in H3S10 phosphorylation and H3K9 methylation which effects the binding of HP1 [66, 67]. The acetylation of H3K9, K14 are essential for encrypting the transcriptional activation message. The histone deacetylases (HDACs) remove the acetyl group on the histones and cause repression of the gene. The presence of HATs and HDACs made the acetylation event a very dynamic phenomenon in the regulation of chromatin.
Methylation of histones occurs at arginine and lysine residues. The process of methylation can be either mono, di or tri methylation. Arginine is methylated by Protein R Methyltransferase (PRMT1) [68]. Methylation at lysine residues K4, K9, K27, K36, K79 of H3 and K20 of H4 occur more frequently than arginine methylation and these methylation modifications are generally irreversible phenomenon. These methylation events can be mono-, di-, or tri-methylations. Amongst the many different modifications that were studied, di- and tri-methylation of H3K4 and H3K9 are well studied. Di- and tri-methylation of H3K4, often in combination with H3K14 acetylation has been linked to transcriptional activation [69]. Methylation at H3K4, H3K36 and H3K79 are implicated in transcriptional activation, while methylations at H3K9, H3K27 and H4K20 are involved chromatin condensation and thus linked to transcriptional repression. Heterochromatin protein HP1 specifically binds to tri-methylated H3K9 and this interaction triggers chromatin reorganization which is responsible for formation of constitutive heterochromatin for gene silencing.
Another major post-translational modification of the histones includes phosphorylation of the histones. This event has been contributed to the mitotic chromatin condensation and transcriptional regulation in interphase. Mitotic phosphorylation occurs at Ser10, Ser28 and Thr11 in H3. In meiosis, Ser10 phosphorylation is required for cohesion of sister chromatid exchange rather than condensation [70, 71]. Histone H1 is phosphorylated at Thr18, Thr146, Thr154, Ser172 and Ser187 for relaxation of chromatin structure and thus regulates cell cycle progression. Histone H2A is phosphorylated at Ser1, which causes repression of transcription, Thr119 phosphorylation is pre-requisite for acetylation of H3K14 and H4K5 in meiosis, while phosphorylation at Ser122, Thr126 and Ser129 play an important role in DNA damage response [63, 72]. H2B phosphorylation at Ser14 is correlated with chromatin condensation during apoptosis [73, 74] and double strand DNA breaks, which form foci at later time points [58]. H2B Ser10 is phosphorylated in H2O2-induced cell death, while in drosophila Ser33 phosphorylation is essential for transcriptional activation, cell cycle progression and development [75, 76]. H3S10 phosphorylation, which generally crosstalk’s with methylation and acetylation of other residues of same or other histones, has been observed in transcriptional activation and chromosome condensation during mitosis. Histone H4 Ser1 phosphorylation by CK II is increased during cell cycle [77, 78].
There exist the variants of the histones called the histone variants which are non-allelic forms of conventional histones. These show specific expression, and are cell cycle regulated. H2AX, a variant of H2A, is found bound to the chromatin and regulate gene expression after IR-induced DNA damage, and is indispensable [58, 79]. These variants are found to be profoundly involved in disease scenario, especially cancer. Overall these different histone modifications and variants of the histones crosstalk with either the modifications on the cis-histones or the modifications on the trans-histones and the other histone variants to encrypt a complex histone code, which modulates the chromatin and thus gene expression. Various events of the cell with respect to histone modifications are depicted in the Fig. 2.
Distinct post-translational modifications on the histone tails. a Acetylation of different residues on all the histone tails puts on the transcription, while b methylation of histones causes chromatin condensation and thus transcriptional repression. c Phosphorylation on the histone H1 causes cell cycle progression while on the other histone tails like in H3 chromatin condensation takes place. Phosphorylation on H4 activates genes while on H2A and H2B it is responsive to DNA damage. Crosstalk between phosphorylation and methylation causes chromosome condensation and thus progression through mitosis
The above-mentioned chromatin modification in consort with MAR elements and MAR binding proteins tightly regulate the gene expression. The role of MARs in chromatin dynamics has been tested using an artificial MARBP called MATH20 [80]. This protein has numerous linked DNA binding domains called AT hooks, which preferentially to AT tracks in the MARs. Since this protein could then bind MARs and associated chromatin specifically, this was used to study chromatin condensation in Xenopus oocytes. MATH20 specifically inhibited chromatin conversion without inhibiting condensation resulting abortive mitotic structures and formation of chromatid balls. In a regulated phenomenon, transcription factors, which bind to the matrix attachment regions, recognize the histone code and orchestrate the gene expression phenomenon. Any alteration in the histone code due to mutations in any of the genes involved in the above-mentioned phenomenon can cause aberrant gene expression at that particular loci and causes malignant transformation.
Scaffold/matrix attachment region binding protein 1 (SMAR1) is a MAR-binding transcription factor which tethers chromatin to the nuclear matrix and modulates the architecture of the chromatin by forming inactive loops [47]. SMAR1 acts as a docking site for chromatin modifiers like HDAC1 and modifies histones over the range of 5 kb on the Cyclin D1 promoter [81]. SMAR1 was first identified in mouse double positive thymocytes. It was found attached to the 400 bp upstream of Eβ enhancer in the T-cell receptor locus (TCR) which was found to be a nuclear matrix/scaffold-associated region, referred to as MARβ, which can associate with the known MAR binding proteins SATB1 and Cux. SMAR1 shares 21.4% identity and 64.3% similarity with the MAR binding domain of human and mouse SATB1, 23.9% identity and 56.5% similarity with a region of bright that contains the tetramer domain [82].
SMAR1 modulates chromatin at Vβ locus
MAR sequences at the Ig V loci are present at a frequency higher than in the rest of the genome. The Vβ locus is organized with 11 hypersensitive sites (HS) and SMAR1 along with other MAR binding proteins binds to the MARβ/HS1 site (Fig. 3). Modulation of chromatin by SMAR1 at the MARβ locus occurs at the double positive stage of the thymocytes development. The accessibility of the MARβ as well as the Eβ enhancer is detected only in the T cells, but not in the B cells, implying the lineage specific presence of these two cis elements which are critical for V(D)J rearrangement of TCRβ gene segments. The chromatin of the MARβ is differentially accessible in different stages of the thymocytes development. MARβ which resides at the 5′ end of the TCRβ enhancer is the docking site for three MARBPs—SMAR1, CUX and SATB1. Amongst these, SMAR1 and CUX together regulate the MARβ locus and over expression of both of these proteins induce the HS1 sites within the TCRβ locus. The higher levels of SMAR1 in the DP stage of thymocytes explain the repression of the V(D)J recombination at this stage of the thymocytes development. As other negative regulators, CUX and SATB1, were also found to bind the MARβ, it was found that SMAR1 works in consort with CUX, but not SATB1 to repress the Eβ. Further it was delineated that the 160–350 region of SMAR1 is essential for its interaction with CUX and show strong repressor activity at the Eβ locus. Thus, SMAR1 and CUX form ternary complex with MARβ and regulate T-cell development by controlling V(D)J recombination via modulation of Eβ enhancer [17].
The transgenic mice of SMAR1 in comparison to non-transgenic littermates show abnormal V(D)J recombination. Analysis of the data in a T-cell stage specific population indicate that over expression of the MAR-binding protein, SMAR1 perturbs the T-cell development from DN to DP stage and these cells tend to remain in the early stage. The marked feature exhibited by SMAR1 transgenic mice is the decreased frequency of T cells expressing commonly used Vβs, particularly Vβ5.1, 5.2, 8.1, 8.2 and 8.3 in thymus and lymph nodes. SMAR1 also significantly reduces the frequency of other Vβs including Vβ9, Vβ10b, Vβ11, Vβ12 and Vβ13. There was no gross change in the recombination frequency of Dβ and Jβ, indicating that SMAR1 affects the V to DJ locus, but not D to J locus [17]. Similarly, other MAR binding proteins like Cux/CDP, B-cell regulator of IgH transcription (Bright) and SATB1 regulate the Ig V genes, flanking the TCR Vβ genes. Cux/CDP and SATB1 are associated with repression while Bright is involved in the activation of Ig transcription. These MAR binding proteins interplay, and it is speculated that they can bind to sequences of non-MARs and the potentiality of the factor binding and MAR activity influences the extent of accessibility to V(D)J recombination and thus could play a role in the unequal rearrangement of individual V genes, providing the effective transcription during maturation and/or activation through the promoter as well as enhancer regions by bringing them into close proximity at the nuclear matrix. This overall phenomenon explains that the MAR binding proteins Cux/CDP, Bright, SMAR1 and SATB1 etc., modulate the chromatin through MARs and constitute the functional nuclear matrix [83].
SMAR1-mediated malignancy and malignant phenotype
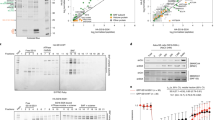
As malignant transformation occurs by means of dysregulation of genes or viral integrations, MARs and the MAR binding proteins play very profound role in this context. Malignancy is often accompanied by alteration in the nuclear structure, size, shape and organization. It is found that the nuclear matrix protein composition of a malignant cell when compared to a normal cell varies. Studies to identify the nuclear matrix binding proteins associated with aggressive cancer phenotype have led to the identification of PARP, Ku, HMG group proteins and SAF-A/G which have higher binding affinity double strand breaks. Unlike all the other MARBPs like SATB1 that is expressed highly in the aggressive cancer cells [84], SMAR1 levels are downregulated in breast cancer derived cell lines and in higher grades of breast cancer. This altered SMAR1 levels are explained, in part, by its modulation of Cyclin D1. Cyclin D1, a G1 phase cyclin which is expressed in redundant fashion in all proliferating cells and collectively control cell-cycle progression along with CDK4/CDK6. Studies have shown that control of Cyclin D1 expression is regulated both at transcriptional and degradation levels. PTEN a tumour suppressor induces cell cycle arrest by reducing the levels of Cyclin D1 and its decreased nuclear availability. MAR-binding protein SMAR1 shows 99% homology with the human BANP and it is mapped to the 16q24 locus. Loss of heterozygosity in this locus, in breast and prostate cancers, has been linked to the dysregulation of Cyclin D1. Our study on SMAR1 by Rampalli et al. has shown that SMAR1 binds to a putative MAR element in the Cyclin D1 promoter and recruits the histone modifying enzyme, HDAC1 along with mSin3A and pocket Rbs and cause deacetylation of the histones H3K9 and H4K8 (Fig. 4). In contrast, knock down of SMAR1 causes increase in histone acetylation. The co-repressor complex that is recruited at the loci of Cyclin D1 leads to chromatin condensation and eventually shutdown of transcription [81].
Regulation of Cyclin D1 expression by SMAR1. a The expression levels of SMAR1 and Cyclin D1 in different cell lines. b The levels of different cyclins in the breast cancer-derived cell line MCF-7. c The mode of regulation of Cyclin D1. SMAR1 modulates the chromatin by recruiting HDAC1 and mSin3a and causes chromatin compaction by deacetylation [81]
It has been established that in the higher grades of breast cancer, the levels of SMAR1 are drastically downregulated and is correlated with higher levels of Cyclin D1. SMAR1 has p53 binding sites and it has been deciphered that this decreased SMAR1 is due to sequestration of acetylated p53 in the heterochromatinic region [85].
It was earlier mentioned that the malignant transformation results in altered phenotype of the nuclear matrix and the cell phenotype as well. The cytoskeletal proteins and cell adhesion molecules play a critical role in the regulation of phenotype of the cell. Cytokeratins are the major components of the intermediate filament molecules that are involved in the structural changes that lead to malignant transformation. Pavithra et al. have shown that MAR-binding protein SMAR1 downregulates the cytokeratin 8 gene expression by displacing the p53 from its cognate sites. The higher levels of this protein cause decreased migration and invasiveness of cells. SMAR1 causes repression of cytokeratin 8 by local chromatin condensation upon genotoxic insults. The upregulation of SMAR1 and the crosstalk between SMAR1 and p53 regulates the CK8 gene expression (Fig. 5). The higher levels of SMAR1 displaces p53 from its cognitive sites on CK8 promoter by 4.5-fold, while the higher levels of p53 displaces SMAR1 from CK8 promoter by 7-fold. The recruitment of SMAR1 results in the loss of recruitment of Pol II by 16-fold, causing transcriptional repression. SMAR1 recruits the repressive machinery identical to that on the Cyclin D1 promoter. The recruitment of HDAC1 on to the promoter is validated by the deacetylation of H3K9 (6-fold) and H4K8 (8-fold) and reduced phosphorylation at H3Ser10 and a prominent tri-methylation mark at H3K9 (10-fold) [86]. This state of chromatin marks the condensed and transcriptional repressive state induced upon the binding of MAR-binding protein SMAR1. This also explains the phenomenon that the nuclear matrix proteins play indispensable role in maintaining the morphology of the cell by regulating the cytoskeletal genes and contributing the nuclear matrix and matrix proteins in the malignant transformation.
SMAR1 downregulates the cytokeratin gene expression by displacing p53 from its cognate sites causing local chromatin modifications like methylation and deacetylation. a p53 drives the transcription of cytokeratin by binding to its cognate sites in the promoter region and causes cell proliferation. b p53 binding site is occupied by SMAR1 in complex with HDAC1 and causes transcriptional shutdown of cytokeratin gene and causes cell cycle arrest (Adopted from Pavithra et al. [86].)
Conclusions and perspectives
Modulation of chromatin is a crucial step in the regulation of gene expression. Nuclear matrix and matrix attachment region binding proteins play a prominent role in orchestrating and decorating chromatin with various modifiers and regulators. The histone code that is etched on the genes dictates the chromatin to be either silent or active. From the current albeit limited knowledge, histone modifications modulate the chromatin organization and eventually the regulation and maintenance of chromatin in the context of heterochromatin, euchromatin, promoters, transcribed regions and sites of DNA damage. The MARs and MAR binding proteins provide and extra layer of gene regulation by forming the active and inactive loops of the chromatin which are being accessed correspondingly by various transcriptional factors. The studies on the MAR-binding protein SMAR1 have clearly earmarked that it causes chromatin condensation at the promoters of the genes by binding to the putative MAR elements. SMAR1 is an abundant and ubiquitous nuclear protein which plays an important role in variety of cellular functions. It binds to the Eβ enhancer in the double positive thymocytes and modulates V(D)J recombination. SMAR1 modulates chromatin structure through direct interaction with HDAC1 and causes deacetylation at active promoters. SMAR1 being a nuclear matrix protein its role in DNA repair, cell death and inflammatory responses have not yet been deciphered and it is likely that it can play a very critical role. The other functional component of the nuclear matrix is involved in the RNA metabolism, which includes the RNA splicing and maturation. SMAR1 has RS domain which is characteristic of most of the RNA regulatory proteins. Deciphering the functional significance of SMAR1 in the RNA regulatory metabolism can contribute to an intriguing phenomenon of cell-cycle regulation.
References
Izaurralde E, Spector DL (2004) Nucleus and gene expression: the interplay of transcriptional and post-transcriptional mechanisms that regulate gene expression. Curr Opin Cell Biol 16(3):219–222
Berezney R, Coffey DS (1977) Nuclear matrix, isolation and characterization of a framework structure from rat liver nuclei. J Cell Biol 73(3):616–637
Berezney R, Mortillaro MJ, Ma H, Wei X, Samarabandu J (1995) The nuclear matrix: a structural milieu for genomic function. Int Rev Cytol 162A:1–65
Smith HC, Berezney R (1980) DNA polymerase alpha is tightly bound to the nuclear matrix of actively replicating liver. Biochem Biophys Res Commun 97(4):1541–1547
Smith HC, Puvion E, Buchholtz LA, Berezney R (1984) Spatial distribution of DNA loop attachment and replicational sites in the nuclear matrix. J Cell Biol 99(5):1794–1802
Jackson DA, Cook PR (1985) Transcription occurs at a nucleoskeleton. EMBO J 4(4):919–925
Zeitlin S, Parent A, Silverstein S, Efstratiadis A (1987) Pre-mRNA splicing and the nuclear matrix. Mol Cell Biol 7(1):111–120
Schirmbeck R, Deppert W (1987) Specific interaction of simian virus 40 large T antigen with cellular chromatin and nuclear matrix during the course of infection. J Virol 61:3561–3569
Djeliova V, Russev G, Anachkova B (2001) Dynamics of association of origins of DNA replication with the nuclear matrix during the cell cycle. Nucleic Acids Res 29(15):3181–3187
Carmo-Fonseca M, Tollervey D, Pepperkok R, Barabino SML, Merdes A, Brunner C, Zamore PD, Green MR, Hurt E, Lamond AI (1991) Mammalian nuclei contain foci which are highly enriched in components of the pre-mRNA splicing machinery. EMBO J 10:195–206
Zhang G, Taneja KL, Singer RH, Green MR (1994) Localization of pre-mRNA splicing in mammalian nuclei. Nature (London) 372:809–812
Blencowe BJ, Carmo-Fonseca M, Behrens SE, Luhrmann R, Lamond AI (1993) Interaction of the human autoantigen p150 with splicing snRNPs. J Cell Sci 105:685–697
Blasquez VC, Sperry AO, Cockerill PN, Garrard WT (1989) Protein: DNA interactions at chromosomal loop attachment sites. Genome 31:503–509
Eliceiri GL, Ryerse JS (1984) Detection of intranuclear clusters of Sm antigens with monoclonal anti-Sm antibodies by immunoelectron microscopy. J Cell Physiol 121:449–451
Fakan S, Lesser G, Martin TE (1984) Ultrastructural distribution of nuclear ribonucleoproteins as visualized by immunocytochemistry on thin sections. J Cell Biol 98:358–363
Carter KC, Bowman D, Carrington W, Fogarty K, McNeil JA, Fay FS, Lawrence JB (1993) A three-dimensional view of precursor messenger RNA metabolism within the mammalian nucleus. Science 259(5099):1330–1335
Dyck J, Maul GG, Miller WH, Chen JD, Kakizuka A, Evans R (1994) A novel macromolecular structure is a target of the premyelocyte-retinoic acid receptor oncoprotein. Cell 76:333–343
Fey EG, Penman S (1988) Nuclear matrix proteins reflect cell type of origin in cultured human cells. Proc Natl Acad Sci USA 85:121–125
Partin AW, Getzenberg RH, CarMichael MJ, Vindivich D, Yoo J, Epstein JI, Coffey DS (1993) Nuclear matrix protein patterns in human benign prostatic hyperplasia and prostrate cancer. Cancer Res 53:744–746
Keesee SK, Meneghini M, Szaro RP, Wu YJ (1994) Nuclear matrix proteins in colon cancer. Proc Natl Acad Sci USA 91:1913–1916
Khanuja PS, Lehr JE, Soule HD, Gehani SK, Noto AC, Choudhary S, Chen E, Pietna KJ (1993) Nuclear matrix proteins in normal and breast cancer cells. Cancer Res 53:3394–3398
Luger K, Mader AW, Richmond RK, Sargent DF, Richmond TJ (1997) Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 389:251–260
Richmond TJ, Finch JT, Rushton B, Rhodes D, Klug A (1984) Structure of the nucleosome core particle at 7 Å resolution. Nature (London) 311:532–537
Rattner JB, Hamkalo BA (1979) Nucleosome packing in interphase chromatin. J Cell Biol 81:453–457
Rattner JB, Saunders C, Davie JR, Hamkalo BA (1982) Ultrastructural organization of yeast chromatin. J Cell Biol 93:217–222
Thoma F, Koller T (1977) Influence of histone H1 on chromatin structure. Cell 12:101–107
Jackson DA, Dickinson P, Cook PR (1990) The size of chromatin loops in HeLa cells. EMBO J 9:567–571
Levy-Wilson B, Fortier C (1989) The limits of the DNase I-sensitive domain of the human apolipoprotein B gene coincide with the locations of chromosomal anchorage loops and define the 5′ and 3′ boundaries of the gene. J Biol Chem 264:21196–21204
Breyne P, Van Montagu M, Depicker A, Gheysen G (1992) Characterization of a plant scaffold attachment region in a DNA fragment that normalizes transgene expression in tobacco. Plant cell 4:463–471
Luderus MEE, de Graaf A, Mattia E, den Blaauwen JL, Grande MA, de Jong L, Van Driel R (1992) Binding of matrix attachment regions to lamin B1. Cell 70:949–959
Allen GC, Hall GE, Childs LC, Weissinger AK, Spiker S, Thompson WF (1993) Scaffold attachment regions increase reporter gene expression in stably transformed plant cells. Plant cell 5:603–613
Blasquez VC, Xu M, Moses SC, Garrard WT (1989) Immunoglobulin kappa gene expression after stable integration. I. Role of the intronic MAR and enhancer in plasmacytoma cells. J Biol Chem 264:21183–21189
Polijak L, Seum C, Mattioni T, Laemmli UK (1994) SARs stimulate but do not confer position independent gene expression. Nucleic Acids Res 22:4386–4394
Forrester WC, Van Genderen C, Jenuwein T, Grosschedl R (1994) Dependence of enhancer-mediated transcription of the immunoglobulin μ gene of nuclear matrix attachment regions. Science 265:1221–1225
Rao BS, Zannis-Hadjopoulos M, Price GB, Reitman M, Martin RG (1990) Sequence similarities among monkey ori-enriched (ors) fragments. Gene 87:233–242
Amati B, Gasser SM (1990) Drosophila Scaffold-attached regions bind nuclear scaffolds and can function as ARS elements in both budding and fission yeasts. Mol Cell Biol 10:5442–5454
Amati B, Pick L, Laroche T, Grasser SM (1990) Nuclear scaffold attachment stimulates, but is not essential for ARS activity in Saccharomyces cerevisiae: Analysis of the Drosophila ftz SAR. EMBO J 9:4007–4016
Moreau J, Marcaud L, Maschat F, Kejzlarova-Lepesant JA, Scherrer K (1982) A + T rich linkers define functional domains in eukaryotic DNA. Nature (London) 295:260–262
Panayotatos N, Wells RD (1981) Cruciform structures in supercoiled DNA. Nature (London) 289:466–470
Berrios M, Osheroff N, Fisher PA (1985) In situ localization of DNA topoisomerase II, a major polypeptide component of the Drosophila nuclear matrix fraction. Proc Natl Acad Sci USA 82:4142–4146
Izaurralde E, Kas E, Laemmli UK (1989) Highly preferential nucleation of histone H1 assembly on scaffold-associated regions. J Mol Biol 210:573–585
Kas E, Izaurralde E, Laemmli UK (1989) Specific inhibition of DNA binding to nuclear scaffolds and histone H1 by distamycin. The role of oligo (dA). oligo (dT) tracts. J Mol Biol 210:587–599
Archer TK, Cordingley MG, Wolford RG, Hager GL (1991) Transcription factor access is mediated by accurately positioned nucleosomes on the mouse mammary tumor virus promoter. Mol Cell Biol 11:688–698
Stein GS, Lian JB, Dworetzky SI, Owen TA, Bortell R, Bidwell JP, Van Winjnen AJ (1991) Regulation of transcription-factor activity during growth and differentiation: involvement of the nuclear matrix in concentration and localization of promoter binding proteins. J Cell Biochem 47:300–305
Waitz W, Loidl P (1991) Cell cycle dependent association of c-myc protein with the nuclear matrix. Oncogene 6(1):29–35
Dickinson LA, Joh T, Kohwi Y, Kohwi-Shigematsu T (1992) A tissue-specific MAR/SAR DNA binding protein with unusual binding site recognition. Cell 70(4):631–645
Chattopadhyay S, Kaul R, Charest A, Houseman D, Chen J (1998) SMAR1, a novel alternatively spliced gene product, binds to scaffold/matrix associated region at TCRβ locus. Genomics 68(1):93–96
Romig H, Fackelmayer FO, Renz A, Ramsperger U, Richter A (1992) Characterization of SAF-A, a novel nuclear DNA binding protein from HeLa cells with high affinity for nuclear matrix/scaffold attachment DNA elements. EMBO J 11:3431–3440
Ivanchenko M, Avramova Z (1992) Interaction of MAR sequences with nuclear matrix proteins. J Cell Biochem 50:190–200
Adachi Y, Kas E, Laemmli UK (1989) Preferential, cooperative binding of DNA topoisomerase II to scaffold-associated regions. EMBO J 8:3997–4006
Mancini MA, Shan B, Nickerson JA, Penman S, Lee WH (1994) The retinoblastoma gene product is a cell cycle-dependent, nuclear matrix associated protein. Proc Natl Acad Sci USA 91:418–422
Schuchard M, Subramaniam M, Ruesink T, Spelsberg TC (1991) Nuclear matrix localization and specific matrix DNA binding by receptor binding factor 1 of the avian oviduct progesterone receptor. Biochemistry 30:9516–9522
Li H, Rouz SJ (1992) Casein kinase II protein kinase is bound to lamina-matrix and phosphorylates lamin like protein in isolated pea nuclei. Proc Natl Acad Sci USA 849:8434–8438
Hendzel MJ, Delcuve GP, Davie JR (1991) Histone deacetylases is a component of the internal nuclear matrix. J Biol Chem 266:21936–21942
Berezney R (1984) Organization and functions of the nuclear matrix. In: Hnilica LS (ed) Chromosomal nonhistone proteins, vol 4. CRC press, Boca Raton, pp 119–180
Hancock R (1982) Topological organization of interphase DNA: the nuclear matrix and other skeletal structures. Biol Cell 46:105–122
Nelson WG, Pienta KJ, Barrack ER, Coffey DS (1986) The role of the nuclear matrix in the organization and function of DNA. Annu Rev Biophys Chem 15:457–475
Fernandez-Capetillo O, Chen HT, Celeste A, Ward I, Romanienko PJ, Morales JC, Naka K, Xia Z, Camerini-Otero RD, Motoyama N, Carpenter PB, Bonner WM, Chen J, Nussenzweig A (2002) DNA damage induced G2M checkpoint activation by histone H2AX and 53Bp1. Nat Cell Biol 4(12):993–997
Lewinski MK, Yamashita M, Emerman M (2006) Retroviral DNA integration: viral and cellular determinants of target-site selection. PLoS Pathog 2(6):e60
Shera KA, Shera CA, McDougall JK (2001) Small tumor virus genomes are integrated near nuclear matrix attachment regions in transformed cells. J Virol 75:12339–12346
Kulkarni A, Pavithra L, Rampalli S, Mogare D, Babu K, Shiekh G, Ghosh S, Chattopadhyay S (2004) HIV-1 integration sites are flanked by potential MARs that alone can act as promoters. Biochem Biophys Res Commun 322:7672–7677
Peterson CL, Laniel MA (2004) Histones and histone modifications. Curr Biol 14:R546–R551
Roth SY, Denu JM, Allis CD (2001) Histone acetyltransferases. Annu Rev Biochem 70:81–120
Sterner DE, Berger SL (2000) Acetylation of histones and transcription-related factors. Microbiol Mol Biol Rev 64:435–459
Allfrey VG, Faulkner R, Mirsky AE (1964) Acetylation and methylation of histones and their possible role in the regulation of RNA synthesis. Proc Natl Acad Sci USA 51:786–794
Fischle W, Tseng BS, Dorman HL, Ueberheide BM, Garcia BA, Shabanowitz J, Hunt DF, Funabiki H, Allis CD (2005) Regulation of HP1-chromatin binding by histone H3 methylation and phosphorylation. Nature 438:1116–1122
Kouzarides T (2007) Chromatin modifications and their function. Cell 128:693–705
Chen D, Ma H, Hong H, Koh SS, Haung SM, Schurter BT, Aswad DW, Stallcup MR (1999) Regulation of transcription by protein methyltransferase. Science 284:2174–2177
Santos-Rosa H, Schneider R, Bannister AJ, Sherriff J, Bernstein BE, Emre NC, Schreiber SL, Mellor J, Kouzarides T (2002) Active genes are tri-methylated at K4 of histone H3. Nature 419:407–411
Hendzel MJ, Wei Y, Mancini MA, Van Hooser A, Ranali T, Brinkley BR, Bazett-Jones DP, Allis CD (1997) Mitosis specific phosphorylation of histone H3 initiates primarily within pericentromeric heterochromatin during G2 and spreads in an ordered fashion coincident with mitotic chromosome condensation. Chromosoma 106(6):348–360
Preuss U, Landsberg G, Scheidtmann KH (2003) Novel mitotic specific phosphorylation of histone H3 and Thr11 mediated by Dlk/ZIP kinase. Nucleic Acids Res 31(3):878–885
Langan TA, Gautier J, Lohka M, Hollingsworth R, Moreno S, Nurse P, Maller J, Sclafani RA (1989) Mammalian growth associated H1 histone kinase. A homologue of cdc2+/CDC28 protein kinases controlling mitotic entry in yeast and frog cells. Mol Cell Biol 9(9):3860–3868
Ajiro K (2000) Histone H2B phosphorylation in mammalian apoptotic cells. An association with DNA fragmentation. J Biol Chem 275(1):439–443
Cheung WL, Turner FB, Krishnamoorthy T, Wolner B, Ahn SH, Foley M, Dorsey JA, Peterson CL, Berger SL, Allis CD (2003) Apoptotic phosphorylation of histone H2B is mediated by mammalian sterile twenty kinase. Cell 113(4):507–517
Ahn SH, Cheung WL, Hsu JY, Diaz RL, Smith MM, Allis CD (2005) Sterile 20 kinase phosphorylates histone H2B at serine 10 during hydrogen peroxide-induced apoptosis in S. cerevisiae. Cell 120(1):25–36
Maile T, Kwoczynski S, Katzenberger RJ, Wassarman DA, Sauer F (2004) TAF1 activates transcription by phosphorylation of serine 33 in histone H2B. Science 304(5673):1010–1014
Barber CM, Turner FB, Wang Y, Hagstrom K, Taverna SD, Mollah S, Ueberheide B, Meyer BJ, Hunt DF, Cheung P, Allis CD (2004) The enhancement of histone H4 and H2A serine 1 phosphorylation during mitosis and S-phase is evolutionarily conserved. Chromosoma 112(7):360–371
Ruiz-Carrillo A, Wangh LJ, Allfrey VG (1975) Processing of newly synthesized histone molecules. Science 190(4210):117–128
Nakamura TM, Du LL, Redon C, Russell P (2004) Histone H2A phosphorylation controls Crb2 recruitment at DNA breaks, maintains checkpoint arrest, influences DNA repair in fission yeast. Mol Cell Biol 24(14):6215–6230
Strick R, Laemmli UK (1995) SARs are cis DNA elements of chromosome dynamics: synthesis of a SAR repressor protein. Cell 83(7):1137–1148
Rampalli S, Pavithra L, Bhatt A, Kundu TK, Chattopadhyay S (2005) Tumor suppressor SMAR1 mediates cyclin D1 repression by recruitment of SIN3/HDAC1 complex. Mol Cell Biol 25(19):8415–8429
Kaul-Ghanekar R, Jalota A, Pavithra L, Tucker P, Chattopadhyay S (2004) SMAR1 and Cux/CDP modulate chromatin and act as negative regulators of the TCRβ enhancer (Eβ). Nucleic Acids Res 32(16):4862–4875
Goebel P, Montalbano A, Ayers N, Kompfner E, Dickinson L, Web CF, Feeney AJ (2002) High frequency of matrix attachment regions and cut-like protein x/CCAAT-displacement protein and B cell regulator of IgH transcription binding sites flanking Ig V region genes. J Immunol 169:2477–2487
Han HJ, Russo J, Kohwi Y, Kohwi-Shigematsu T (2008) SATB1 reprogrammes gene expression to promote breast tumour growth and metastasis. Nature 452:187–193
Singh K, Mogare D, Giridharagopalan RO, Gogiraju R, Pande G, Chattopadhyay S (2007) p53 target gene SMAR1 is dysregulated in breast cancer: its role in cancer cell migration and invasion. PLoS ONE 2(7):e660
Pavithra L, Singh S, Sreenath K, Chattopadhyay S (2008) Tumor suppressor SMAR1 downregulates Cytokeratin 8 expression by displacing p53 from its cognate site. Int J Biochem Cell Biol 41(4):862–871
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Nakka, K.K., Chattopadhyay, S. Modulation of chromatin by MARs and MAR binding oncogenic transcription factor SMAR1. Mol Cell Biochem 336, 75–84 (2010). https://doi.org/10.1007/s11010-009-0262-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-009-0262-7