Abstract
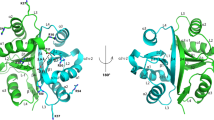
Protein domain family PF06855 (DUF1250) is a family of small domains of unknown function found only in bacteria, and mostly in the order Bacillales and Lactobacillales. Here we describe the solution NMR or X-ray crystal structures of three representatives of this domain family, MW0776 and MW1311 from Staphyloccocus aureus and yozE from Bacillus subtilis. All three proteins adopt a four-helix motif similar to sterile alpha motif (SAM) domains. Phylogenetic analysis classifies MW1311 and yozE as functionally equivalent proteins of the UPF0346 family of unknown function, but excludes MW0776, which likely has a different biological function. Our structural characterization of the three domains supports this separation of function. The structures of MW0776, MW1311, and yozE constitute the first structural representatives from this protein domain family.

Similar content being viewed by others
Abbreviations
- NESG:
-
Northeast Structural Genomics Consortium
- PDB:
-
Protein Data Bank
- rmsd:
-
Root-mean-square-deviation
References
Tatusov RL, Koonin EV, Lipman DJ (1997) Science 278:631–637
Finn RD, Mistry J, Tate J, Coggill P, Heger A, Pollington JE, Gavin OL, Gunasekaran P, Ceric G, Forslund K, Holm L, Sonnhammer EL, Eddy SR, Bateman A (2010) Nucleic Acids Res 38:D211–D222
Xiao R, Anderson S, Aramini J, Belote R, Buchwald WA, Ciccosanti C, Conover K, Everett JK, Hamilton K, Huang YJ, Janjua H, Jiang M, Kornhaber GJ, Lee DY, Locke JY, Ma LC, Maglaqui M, Mao L, Mitra S, Patel D, Rossi P, Sahdev S, Sharma S, Shastry R, Swapna GV, Tong SN, Wang D, Wang H, Zhao L, Montelione GT, Acton TB (2010) J Struct Biol 172:21–33
Jansson M, Li YC, Jendeberg L, Anderson S, Montelione GT, Nilsson B (1996) J Biomol NMR 7:131–141
Doublie S, Kapp U, Aberg A, Brown K, Strub K, Cusack S (1996) FEBS Lett 384:219–221
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) J Biomol NMR 6:277–293
Moseley HN, Monleon D, Montelione GT (2001) Methods Enzymol 339:91–108
Vuister GW, Bax A (1993) J Am Chem Soc 115:7772–7777
Neri D, Szyperski T, Otting G, Senn H, Wuthrich K (1989) Biochemistry 28:7510–7516
Moseley HN, Sahota G, Montelione GT (2004) J Biomol NMR 28:341–355
Herrmann T, Guntert P, Wuthrich K (2002) J Mol Biol 319:209–227
Pascal SM, Muhandiram DR, Yamazaki T, Forman-Kay JD, Kay LE (1994) J Magn Reson, Ser B 103:197–201
Cornilescu G, Delaglio F, Bax A (1999) J Biomol NMR 13:289–302
Linge JP, Williams MA, Spronk CA, Bonvin AM, Nilges M (2003) Proteins 50:496–506
Brunger AT, Adams PD, Clore GM, DeLano WL, Gros P, Grosse-Kunstleve RW, Jiang JS, Kuszewski J, Nilges M, Pannu NS, Read RJ, Rice LM, Simonson T, Warren GL (1998) Acta Crystallogr D Biol Crystallogr 54(Pt 5):905–921
Huang YJ, Tejero R, Powers R, Montelione GT (2006) Proteins 62:587–603
Tejero R, Monleon D, Celda B, Powers R, Montelione GT (1999) J Biomol NMR 15:251–264
Luthy R, Bowie JU, Eisenberg D (1992) Nature 356:83–85
Sippl MJ (1993) Proteins 17:355–362
Laskowski RA, MacArthur MW, Moss DS, Thornton JM (1993) J Appl Crystallogr 26:283–291
Davis IW, Leaver-Fay A, Chen VB, Block JN, Kapral GJ, Wang X, Murray LW, Arendall WB III, Snoeyink J, Richardson JS, Richardson DC (2007) Nucleic Acids Res 35:W375–W383
Bhattacharya A, Tejero R, Montelione GT (2007) Proteins 66:778–795
Huang YJ, Powers R, Montelione GT (2005) J Am Chem Soc 127:1665–1674
Luft JR, Wolfley JR, Said MI, Nagel RM, Lauricella AM, Smith JL, Thayer MH, Veatch CK, Snell EH, Malkowski MG, Detitta GT (2007) Protein Sci 16:715–722
Otwinowski Z, Minor W (1997) Methods Enzymol 276:307–326
Matthews BW (1968) J Mol Biol 33:491–497
Weeks CM, Miller R (1999) Acta Crystallogr D Biol Crystallogr 55:492–500
Terwilliger TC (2003) Methods Enzymol 374:22–37
Cowtan EP (2004) Acta Crystallogr D Biol Crystallogr D60:2126–2132
Engh RA, Huber R (1991) Acta Crystallogr A 47:392–400
Schulze-Gahmen U, Pelaschier J, Yokota H, Kim R, Kim SH (2003) Proteins 50:526–530
Masayama A, Kuwana R, Takamatsu H, Hemmi H, Yoshimura T, Watabe K, Moriyama R (2007) J Bacteriol 189:2369–2375
Beres SB, Richter EW, Nagiec MJ, Sumby P, Porcella SF, DeLeo FR, Musser JM (2006) Proc Natl Acad Sci USA 103:7059–7064
Johnson TJ, Kariyawasam S, Wannemuehler Y, Mangiamele P, Johnson SJ, Doetkott C, Skyberg JA, Lynne AM, Johnson JR, Nolan LK (2007) J Bacteriol 189:3228–3236
Murzin AG, Brenner SE, Hubbard T, Chothia C (1995) J Mol Biol 247:536–540
Holm L, Kaariainen S, Rosenstrom P, Schenkel A (2008) Bioinformatics 24:2780–2781
Keel AY, Rambo RP, Batey RT, Kieft JS (2007) Structure 15:761–772
Tahirov TH, Temiakov D, Anikin M, Patlan V, McAllister WT, Vassylyev DG, Yokoyama S (2002) Nature 420:43–50
Qiao F, Bowie JU (2005) Sci STKE 2005:re7
Chenna R, Sugawara H, Koike T, Lopez R, Gibson TJ, Higgins DG, Thompson JD (2003) Nucleic Acids Res 31:3497–3500
Baker NA, Sept D, Joseph S, Holst MJ, McCammon JA (2001) Proc Natl Acad Sci USA 98:10037–10041
Acknowledgments
The authors thank J.M. Aramini, M.C. Baran, C. Ciccosanti, K.E. Cunningham, J.K. Everett, Y. Fang, H. Janjua, L.-C. Ma, B. Rost, K. Shetty, and D. Wang for valuable scientific discussions and/or technical support, A. Lauicella and G. DeTitta for performing initial crystallization screening, and R. Abramowitz and J. Schwarnof for access to beamline X4A at Brookhaven National Laboratory. The NESG expression vectors SR391-21.2, ZR218-21.1, and ZR215-21.1 have been deposited in the PSI Materials Repository (http://psimr.asu.edu/). This research was supported by the Protein Structure Initiative of the US National Institutes of Health, grants U54 GM074958 and U54 GM094597.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Swapna, G.V.T., Rossi, P., Montelione, A.F. et al. Three structural representatives of the PF06855 protein domain family from Staphyloccocus aureus and Bacillus subtilis have SAM domain-like folds and different functions. J Struct Funct Genomics 13, 163–170 (2012). https://doi.org/10.1007/s10969-012-9134-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10969-012-9134-6