Abstract
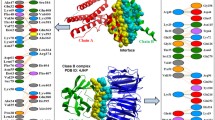
We analyze the characteristics of protein–protein interfaces using the largest datasets available from the Protein Data Bank (PDB). We start with a comparison of interfaces with protein cores and non-interface surfaces. The results show that interfaces differ from protein cores and non-interface surfaces in residue composition, sequence entropy, and secondary structure. Since interfaces, protein cores, and non-interface surfaces have different solvent accessibilities, it is important to investigate whether the observed differences are due to the differences in solvent accessibility or differences in functionality. We separate out the effect of solvent accessibility by comparing interfaces with a set of residues having the same solvent accessibility as the interfaces. This strategy reveals residue distribution propensities that are not observable by comparing interfaces with protein cores and non-interface surfaces. Our conclusions are that there are larger numbers of hydrophobic residues, particularly aromatic residues, in interfaces, and the interactions apparently favored in interfaces include the opposite charge pairs and hydrophobic pairs. Surprisingly, Pro-Trp pairs are over represented in interfaces, presumably because of favorable geometries. The analysis is repeated using three datasets having different constraints on sequence similarity and structure quality. Consistent results are obtained across these datasets. We have also investigated separately the characteristics of heteromeric interfaces and homomeric interfaces.














Similar content being viewed by others
Abbreviations
- PDB:
-
Protein Data Bank
- PQS:
-
Protein Quaternary Structure
- ΔASA:
-
Changes in solvent accessible surface area
- rASA:
-
Relative solvent accessibility
- RIP:
-
Raw interface propensity
- NIP:
-
Normalized interface propensity
References
Chothia C, Janin J (1975) Nature 256:705–708
Wodak SJ, Janin J (2002) Adv Protein Chem 61:9–73
Deremble C, Lavery R (2005) Curr Opin Struct Biol 15:171–175
Ponstingl H, Kabir T, Gorse D, Thornton JM (2005) Progr Biophys Mol Biol 89:9–35
Reichmann D, Rahat O, Cohen M, Neuvirth H, Schreiber G (2007) Curr Opin Struct Biol 17:67–76
Sheinerman FB, Norel R, Honig B (2000) Curr Opin Struct Biol 10:153–159
Heifetz A, Katchalski-Katzir E, Eisenstein M (2002) Protein Sci 11:571–587
Vizcarra CL, Mayo SL (2005) Curr Opin Chem Biol 9:622–626
Glaser F, Steinberg DM, Vakser IA, Ben-Tal N (2001) Proteins 43:89–102
Ofran Y, Rost B (2003) J Mol Biol 325:377–387
Miyazawa S, Jernigan RL (1985) Macromolecules 18:534–552
Keskin O, Bahar I, Badretinov AY, Ptitsyn OB, Jernigan RL (1998) Protein Sci 7:2578–2586
Young L, Jernigan RL, Covell DG (1994) Protein Sci 3:717–729
Berchanski A, Shapira B, Eisenstein M (2004) Proteins 56:130–142
Ben-Naima A (2006) J Chem Phys 125:24901
Keskin O, Mab B, Nussinov R (2005) J Mol Biol 345:1281–1294
Jones S, Thornton JM (1996) Proc Natl Acad Sci USA 93:13–20
Peter Block JP, Hülermeier E, Sanschagrin P, Sotriffer CA, Klebe G (2006) Proteins 65:607–622
Chakrabarti P, Janin J (2002) Proteins 47:334–343
Kyu-il Cho KL, Lee KH, Kim D, Lee D (2006) Proteins 65:593–606
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) Nucl Acids Res 28:235–242
Henrick K, Thornton JM (1998) Trends Biochem Sci 23:358–361
Hubbard SJ (1993) NACCESS, department of biochemistry and molecular biology. University College, London
Gutteridge A, Bartlett GJ, Thornton JM (2003) J Mol Biol 330:719–734
Kyte J, Doolittle RF (1982) J Mol Biol 157:105–132
Nooren IMA, Thornton JM (2003) J Mol Biol 325:991–1016
Kabsch W, Sander C (1983) Biopolymers 22:2577–2637
McCoy AJ, Chandana Epa V, Colman PM (1997) J Mol Biol 268:570–584
Lo Conte L, Chothia C, Janin J (1999) J Mol Biol 285:2177–2198
Bahadur RP, Chakrabarti P, Rodier F, Janin J (2003) Proteins 53:708–719
Jones S, Thornton JM (1997) J Mol Biol 272:121–132
Prasad Bahadur R, Chakrabarti P, Rodier F, Janin J (2004) J Mol Biol 336:943–955
Raih MF, Ahmad S, Zheng R, Mohamed R (2005) Biophys Chem 114:63–69
Creighton T (1997) Protein structures and molecular properties. WH Freeman, New York
Bahar I, Jernigan RL (1997) J Mol Biol 266:195–244
Acknowledgements
This Research was supported in part by a grant from the National Institutes of Health (GM 066387) to VH, DD, and RLJ.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yan, C., Wu, F., Jernigan, R.L. et al. Characterization of Protein–Protein Interfaces. Protein J 27, 59–70 (2008). https://doi.org/10.1007/s10930-007-9108-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-007-9108-x