Abstract
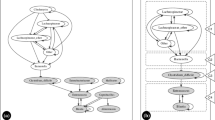
Agent-based modeling is a computational modeling method that represents system-level behavior as arising from multiple interactions between the multiple components that make up a system. Biological systems are thus readily described using agent-based models (ABMs), as multi-cellular organisms can be viewed as populations of interacting cells, and microbial systems manifest as colonies of individual microbes. Intersections between these two domains underlie an increasing number of pathophysiological processes, and the intestinal tract represents one of the most significant locations for these inter-domain interactions, so much so that it can be considered an internal ecology of varying robustness and function. Intestinal infections represent significant disturbances of this internal ecology, and one of the most clinically relevant intestinal infections is Clostridium difficile infection (CDI). CDI is precipitated by the use of broad-spectrum antibiotics, involves the depletion of commensal microbiota, and alterations in bile acid composition in the intestinal lumen. We present an example ABM of CDI (the C. difficile Infection ABM, or CDIABM) to examine fundamental dynamics of the pathogenesis of CDI and its response to treatment with anti-CDI antibiotics and a newer treatment therapy, fecal microbial transplant. The CDIABM focuses on one specific mechanism of potential CDI suppression: commensal modulation of bile acid composition. Even given its abstraction, the CDIABM reproduces essential dynamics of CDI and its response to therapy, and identifies a paradoxical zone of behavior that provides insight into the role of intestinal nutritional status and the efficacy of anti-CDI therapies. It is hoped that this use case example of the CDIABM can demonstrate the usefulness of both agent-based modeling and the application of abstract functional representation as the biomedical community seeks to address the challenges of increasingly complex diseases with the goal of personalized medicine.







Similar content being viewed by others
References
An G et al (2009) Agent-based models in translational systems biology. Wiley Interdiscip Rev Syst Biol Med. doi:10.1002/wsbm.45
Bankes SC (2002) Agent-based modeling: a revolution? Proc Natl Acad Sci USA 99(Suppl 3):7199–7200
Bonabeau E (2002) Agent-based modeling: methods and techniques for simulating human systems. Proc Natl Acad Sci USA 99(Suppl 3):7280–7287
Hunt CA et al (2009) At the biological modeling and simulation frontier. Pharm Res 26(11):2369–2400
Walker DC, Southgate J (2009) The virtual cell—a candidate co-ordinator for ‘middle-out’ modeling of biological systems. Brief Bioinform 10(4):450–461
Zhang L, Athale CA, Deisboeck TS (2007) Development of a three-dimensional multiscale agent-based tumor model: simulating gene-protein interaction profiles, cell phenotypes and multicellular patterns in brain cancer. J Theor Biol 244(1):96–107
Santoni D, Pedicini M, Castiglione F (2008) Implementation of a regulatory gene network to simulate the TH1/2 differentiation in an agent-based model of hypersensitivity reactions. Bioinformatics 24(11):1374–1380
Fallahi-Sichani M et al (2011) Multiscale computational modeling reveals a critical role for TNF-alpha receptor 1 dynamics in tuberculosis granuloma formation. J Immunol 186(6):3472–3483
Hoehme S, Drasdo D (2010) A cell-based simulation software for multi-cellular systems. Bioinformatics 26(20):2641–2642
Adra S et al (2010) Development of a three dimesional multiscale computational model of the human epidermis. PLoS One 5(1):e8511
Christley S, Alber MS, Newman SA (2007) Patterns of mesenchymal condensation in a multiscale, discrete stochastic model. PLoS Comput Biol 3(4):e76
Grimm V et al (2005) Pattern-oriented modeling of agent-based complex systems: lessons from ecology. Science 310:987–991
Macy M, Willer R (2002) From factors to actors: compuational sociology and agent-based modeling. Annu Rev Sociol 28:143–166
Tesfatsion L (2002) Agent-based computational economics: growing economies from the bottom up. Artificial Life 8(1):55–82
Parker J, Epstein J (2011) A distributed platform for global-scale agent-based models of disease transmission. ACM Trans Model Comput Simul 22(1):2
An G (2001) Agent-based computer simulation and sirs: building a bridge between basic science and clinical trials. Shock 16(4):266–273
An G (2004) In silico experiments of existing and hypothetical cytokine-directed clinical trials using agent-based modeling. Crit Care Med 32(10):2050–2060
Hunt CA et al (2006) Physiologically based synthetic models of hepatic disposition. J Pharmacokinet Pharmacodyn 33(6):737–772
An G (2008) Introduction of an agent-based multi-scale modular architecture for dynamic knowledge representation of acute inflammation. Theor Biol Med Model 5(1):11
Mansury Y, Diggory M, Deisboeck TS (2006) Evolutionary game theory in an agent-based brain tumor model: exploring the ‘Genotype-Phenotype’ link. J Theor Biol 238(1):146–156
Engelberg JA, Ropella GE, Hunt CA (2008) Essential operating principles for tumor spheroid growth. BMC Syst Biol 2(1):110
Deisboeck TS et al (2001) Pattern of self-organization in tumour systems: complex growth dynamics in a novel brain tumour spheroid model. Cell Prolif 34(2):115–134
Chen S, Ganguli S, Hunt CA (2004) An agent-based computational approach for representing aspects of in vitro multi-cellular tumor spheroid growth. Conf Proc IEEE Eng Med Biol Soc 1:691–694
Thorne BC et al (2006) Modeling blood vessel growth and leukocyte extravasation in ischemic injury: an integrated agent-based and finite element analysis approach. J Crit Care 21(4):346
Tang J, Ley KF, Hunt CA (2007) Dynamics of in silico leukocyte rolling, activation, and adhesion. BMC Syst Biol 1:14
Tang J et al (2004) Simulating leukocyte-venule interactions–a novel agent-oriented approach. Conf Proc IEEE Eng Med Biol Soc 7:4978–4981
Bailey AM, Thorne BC, Peirce SM (2007) Multi-cell agent-based simulation of the microvasculature to study the dynamics of circulating inflammatory cell trafficking. Ann Biomed Eng 35(6):916–936
Bailey AM et al (2009) Agent-based model of therapeutic adipose-derived stromal cell trafficking during ischemia predicts ability to roll on P-selectin. PLoS Comput Biol 5(2):e1000294
Mi Q et al (2007) Agent-based model of inflammation and wound healing: insights into diabetic foot ulcer pathology and the role of transforming growth factor-beta1. Wound Repair Regen 15(5):671–682
Walker DC et al (2004) Agent-based computational modeling of wounded epithelial cell monolayers. IEEE Trans Nanobioscience 3(3):153–163
An G (2009) Dynamic knowledge representation using agent-based modeling: ontology instantiation and verification of conceptual models. Methods Mol Biol 500:445–468
Kirschner DE et al (2007) Toward a multiscale model of antigen presentation in immunity. Immunol Rev 216:93–118
Reynolds CW (1987) Flocks, herds, and schools: a distributed behavioral model in computer graphics. In: SIGGRAPH ‘87 Conf. Proc, vol. 21(4), pp. 25-34
Lipniacki T et al (2006) Stochastic regulation in early immune response. Biophys J 90(3):725–742
Lipniacki T et al (2006) Transcriptional stochasticity in gene expression. J Theor Biol 238(2):348–367
Carbo A et al (2013) Predictive computational modeling of the mucosal immune responses during Helicobacter pylori infection. PLoS One 8(9):e73365
Kim M et al (2012) Immature oxidative stress management as a unifying principle in the pathogenesis of necrotizing enterocolitis: insights from an agent-based model. Surg Infect 13(1):18–32
Seal JB et al (2011) Agent-based dynamic knowledge representation of Pseudomonas aeruginosa virulence activation in the stressed gut: towards characterizing host-pathogen interactions in gut-derived sepsis. Theor Biol Med Model 8:33
Carrie M (2010) Agent based model of Salmonella typhimurium infection in the gut, in Computer Science. 2010, University of Aberdeen: Online. p. 87
Wendelsdorf KV et al (2012) ENteric Immunity SImulator: a tool for in silico study of gastroenteric infections. IEEE Trans Nanobioscience 11(3):273–288
Wilensky U (1999) NetLogo. Center for Connected Learning, Northwestern University, Evanston, IL. http://ccl.northwestern.edu/netlogo/. Accessed 1 July 2013
Britton RA, Young VB (2012) Interaction between the intestinal microbiota and host in Clostridium difficile colonization resistance. Trends Microbiol 20(7):313–319
Eyre DW et al (2013) Diverse sources of C. difficile infection identified on whole-genome sequencing. N Engl J Med 369(13):1195–1205
Vedantam G et al (2012) Clostridium difficile infection: toxins and non-toxin virulence factors, and their contributions to disease establishment and host response. Gut Microbes 3(2):121–134
McCune VL, Struthers JK, Hawkey PM (2014) Faecal transplantation for the treatment of Clostridium difficile infection: a review. Int J Antimicrob Agents 43(3):201–206
Pattani R et al (2013) Probiotics for the prevention of antibiotic-associated diarrhea and Clostridium difficile infection among hospitalized patients: systematic review and meta-analysis. Open Med 7(2):e56–e67
Sorg J, Sonenshein A (2008) Bile salts and glycine as cogerminants for Clostridium difficile spore germination. J Bacteriol 190:2505
Wilson K, Kennedy M, Fekety F (1982) Use of sodium taurocholate to enhance spore recovery on a medium selective for Clostridium difficile. J Clin Microbiol 15(3):443–446
Wilson K, Perini F (1988) Role of competition for nutrients in suppression of Clostridium difficile by colonic microflora. Infect Immun 56(10):2610–2614
An G (2010) Closing the scientific loop: bridging correlation and causality in the petaflop age. Sci Transl Med 2(41):41ps34
An G (2009) A model of TLR4 signaling and tolerance using a qualitative, particle-event-based method: introduction of spatially configured stochastic reaction chambers (SCSRC). Math Biosci 217(1):43–52
Balci O (1998) Verification, validation and testing. In: Banks J (ed) Handbook of simulation: principles, methodology, advances, applications, and practice. John Wley & Sons, New York, pp 335–396
Underwood S et al (2009) Characterization of the sporulation initiation pathway of Clostridium difficile and its role in toxin production. J Bacteriol 191(23):7296–7305
An G, Christley S (2011) Agent-based modeling and biomedical Ontologies: a roadmap. Wiley Interdiscip Rev Comput Stat 3(4):343–356
An G, Wilensky U (2009) From artificial life to in silico medicine: NetLogo as a means of translational knowledge representation in biomedical research. In: Adamatsky A, Komosinski M (eds) Artificial life in software, 2nd edn. Springer-Verlag, London, pp 183–209
Seok J et al (2013) Genomic responses in mouse models poorly mimic human inflammatory diseases. Proc Natl Acad Sci USA 110(9):3507–3512
Pound P, Bracken MB (2014) Is animal research sufficiently evidence based to be a cornerstone of biomedical research? BMJ 348:g3387
An GC, Faeder JR (2009) Detailed qualitative dynamic knowledge representation using a BioNetGen model of TLR-4 signaling and preconditioning. Math Biosci 217(1):53–63
Acknowledgments
This work was supported, in part, by the National Institutes of Health, Grants NIGMS P50GM53789 and NIDDK P30DK42086.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Peer, X., An, G. Agent-based model of fecal microbial transplant effect on bile acid metabolism on suppressing Clostridium difficile infection: an example of agent-based modeling of intestinal bacterial infection. J Pharmacokinet Pharmacodyn 41, 493–507 (2014). https://doi.org/10.1007/s10928-014-9381-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10928-014-9381-1