Abstract
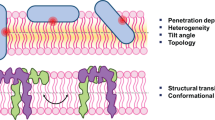
Cellular membranes have relevant roles in processes related to proteases like human kallikreins and cathepsins. As enzyme and substrate may interact with cell membranes and associated co-factors, it is important to take into account the behavior of peptide substrates in the lipid environment. In this paper we report an study based on energy transfer in two bradykinin derived peptides labeled with the donor-acceptor pair Abz/Eddnp (ortho-aminobenzoic acid/N-[2,4-dinitrophenyl]-ethylenediamine). Time-resolved fluorescence experiments were performed in phosphate buffer and in the presence of large unilamelar vesicles of phospholipids, and of micelles of sodium dodecyl sulphate (SDS). The decay kinetics were analyzed using the program CONTIN to obtain end-to-end distance distribution functions f(r). Despite of the large difference in the number of residues the end-to-end distance of the longer peptide (9 amino acid residues) is only 20 % larger than the values obtained for the shorter peptide (5 amino acid residues). The proline residue, in position 4 of the bradykinin sequence promotes a turn in the longer peptide chain, shortening its end-to-end distance. The surfactant SDS has a strong disorganizing effect, substantially broadening the distance distributions, while temperature increase has mild effects in the flexibility of the chains, causing small increase in the distribution width. The interaction with phospholipid vesicles stabilizes more compact conformations, decreasing end-to-end distances in the peptides. Anisotropy experiments showed that rotational diffusion was not severely affected by the interaction with the vesicles, suggesting a location for the peptides in the surface region of the bilayer, a result consistent with small effect of lipid phase transition on the peptides conformations.





Similar content being viewed by others
References
Carmona AK, Schwager SL, Juliano MA, Juliano L, Sturrock ED (2007) A continuous fluorescence resonance energy transfer angiotensin I-converting enzyme assay. Nat Protoc 1:1971–1976
Korkmaz B, Attucci S, Juliano MA, Kalupov T, Jourdan ML, Juliano L, Gauthier F (2008) Measuring elastase, proteinase 3 and cathepsin G activities at the surface of human neutrophils with fluorescence resonance energy transfer substrates. Nat Protoc 3:991–1000
Chagas JR, Portaro FC, Hirata IY, Almeida PC, Juliano MA, Juliano L, Prado ES (1995) Determinants of the unusual cleavage specificity of lysyl-bradykinin-releasing kallikreins. Biochem J 306:63–69
Del Nery E, Chagas JR, Juliano MA, Prado ES, Juliano L (1995) Evaluation of the extent of the binding site in human tissue kallikrein by synthetic substrates with sequences of human kininogen fragments. Biochem J 312:233–238
Henkhaus RS, Roy UKB, Cavallo-Medved D, Sloane BF, Gerner EW, Ignatenko NA (2008) Caveolin-1-Mediated Expression and Secretion of Kallikrein 6 in Colon Cancer Cells. Neoplasia 10:140–148
Takara M, Ito AS (2005) General and specific solvent effects in optical spectra of ortho-aminobenzoic acid. J Fluoresc 15:171–177
Turchiello RF, Lamy-Freund MT, Hirata IY, Juliano L, Ito AS (2002) Ortho-Aminobenzoic Acid-Labeled Bradykinins in Interaction with Lipid Vesicles: Fluorescence Study. Biopolymers 65:336–346
Turchiello RF, Juliano L, Ito AS, Lamy-Freund MT (2000) How bradykinin alters the lipid membrane structure. A spin label comparative study with bradykinin fragments and other cations. Biopolymers 54:211–221
Regoli D, Barabe J (1980) Pharmacology of bradykinin and related kinins. Pharmacol Rev 32:1–46
Stewart JM (1995) Bradykinin antagonists: development and applications. Biopolymers 37:143–155
Cann JR, Stewart JM, Matsueda GR (1973) A circular dichroism study of the secondary structure of bradykinin. Biochemistry 12:3780–3788
Denys L, Bothner-By AA, Fisher GH, Ryan JW (1982) Conformational diversity of bradykinin in aqueous solution. Biochemistry 21:6531–6536
Cann JR, Liu X, Stewart JM, Gera J, Kotovych GA (1994) CD and NMR study of multiple bradykinin conformations in aqueous tri-£uoroethanol solutions. Biopolymers 34:869–878
Kotovych G, Cann JR, Stewart JM, Yamamoto H (1998) NMR and CD conformational studies of bradykinin and its agonists and antagonists: application to receptor binding. Biochem Cell Biol 76:257–266
Ito AS, Souza ES, Barbosa SR, Nakaie CR (2001) Fluorescence study of conformational properties of melanotropins labeled with aminobenzoic acid. Biophys J 81:1180–1189
Pimenta DC, Nantes IL, Souza ES, le Boniec B, Ito AS, Tersariol ILS, Oliveira V, Juliano MA, Juliano L (2002) Interaction of heparin with internally quenched fluorogenic peptides derived from heparin-binding consensus sequences, kallistatin and anti-thrombin III. Biochem J 366:435–446
Souza ES, Hirata IY, Juliano L, Ito AS (2000) End-to-end distribution distances in bradykinin observed by fluorescence energy transfer. Bioch Biophys Acta 1474:251–261
Ito AS, Turchiello RF, Hirata IY, Cezari MH, Meldal M, Juliano L (1998) Fluorescent properties of amino acids labeled with orthoaminobenzoic. Biospectroscopy 4:395–402
Provencher SW (1982) CONTIN: a general purpose constrained regularization program for inverting noisy linear algebraic and integral equations, Computer Phys. Commun 27:229–242
Turchiello RF, Lamy-Freund MT, Hirata IY, Juliano L, Ito AS (1998) ortho-aminobenzoic acid as a fluorescent probe for the interaction between peptides and micelles. Biophys Chem 73:217–225
Takara M, Eisenhut JK, Hirata IY, Juliano L, Ito AS (2009) Solvent effects in optical spectra of ortho-Aminobenzoic acid derivatives. J Fluoresc 19:1053–1060
Acknowledgments
We thank the Brazilian agencies FAPESP and CNPq for financial support. MB and LMR thanks FAPESP and CNPq for fellowships. Support from INCT-FCx, Brazil, is also acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Montaldi, L.R., Berardi, M., Souza, E.S. et al. End-to-end Distance Distribution in Fluorescent Derivatives of Bradykinin in Interaction with Lipid Vesicles. J Fluoresc 22, 1151–1158 (2012). https://doi.org/10.1007/s10895-012-1054-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10895-012-1054-0