Abstract
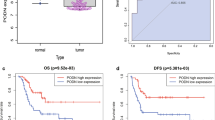
Osteosarcoma represents one of the most aggressive tumors of bone among adolescents and young adults. Despite improvements in treatment, osteosarcoma has a grave prognosis. The identification of prognostic factors is still in its infancy. Weighted gene correlation network analysis (WGCNA) was conducted on mRNA-sequencing and clinical information (gender, survival and metastasis) of osteosarcoma patients from the TARGET database to obtain genes in modules associated with metastasis of osteosarcoma. The Cox regression analysis was then performed on the gene expression profile from TARGET to screen genes associated with patients’ survival. Known genes related to osteosarcoma were obtained by intersecting osteosarcoma-related genes from DisGeNET and DiGSeE, followed by the construction of PPI network of osteosarcoma-related genes and survival-related genes in modules. The screened key genes were subject to multi-factor Cox proportional hazards model, and osteosarcoma patients were classified into high- and low- risk groups according to the risk score to evaluate the potential of key genes to predict the survival of osteosarcoma patients. The WGCNA showed that 4 genes in tan and 19 genes in pink modules were related to the survival of osteosarcoma patients. Osteosarcoma-related known genes (9) were obtained in intersection of DisGeNET and DiGSeE. PPI network identified 4 key genes (KRT5, HIPK2, MAP3K5 and CD5) closely associated with survival of osteosarcoma patients. HIPK2, MAP3K5 and CD5 expression was inversely correlated with survival risk, while KRT5 expression was positively correlated with survival risk. These results show KRT5, HIPK2, MAP3K5 and CD5 serve as prognostic factors of osteosarcoma patients.






Similar content being viewed by others
References
Abashev TM, Metzler MA, Wright DM, Sandell LL (2017) Retinoic acid signaling regulates Krt5 and Krt14 independently of stem cell markers in submandibular salivary gland epithelium. Dev Dyn 246(2):135–147. https://doi.org/10.1002/dvdy.24476
Angulo P, Kaushik G, Subramaniam D, Dandawate P, Neville K, Chastain K, Anant S (2017) Natural compounds targeting major cell signaling pathways: a novel paradigm for osteosarcoma therapy. J Hematol Oncol 10(1):10. https://doi.org/10.1186/s13045-016-0373-z
Burgueno-Bucio E, Mier-Aguilar CA, Soldevila G (2019) The multiple faces of CD5. J Leukoc Biol 105(5):891–904. https://doi.org/10.1002/JLB.MR0618-226R
Chen JT, Fong YC, Li TM, Liu JF, Hsu CW, Chang CS, Tang CH (2008) DDTD, an isoflavone derivative, induces cell apoptosis through the reactive oxygen species/apoptosis signal-regulating kinase 1 pathway in human osteosarcoma cells. Eur J Pharmacol 597(1-3):19–26. https://doi.org/10.1016/j.ejphar.2008.08.036
Fisher JN, Terao M, Fratelli M, Kurosaki M, Paroni G, Zanetti A, Gianni M, Bolis M, Lupi M, Tsykin A, Goodall GJ, Garattini E (2015) MicroRNA networks regulated by all-trans retinoic acid and Lapatinib control the growth, survival and motility of breast cancer cells. Oncotarget 6(15):13176–13200. https://doi.org/10.18632/oncotarget.3759
Gao L, Lin P, Chen P, Gao RZ, Yang H, He Y, Chen JB, Luo YG, Xu QQ, Liang SW, Gu JH, Huang ZG, Dang YW, Chen G (2019) A novel risk signature that combines 10 long noncoding RNAs to predict neuroblastoma prognosis. J Cell Physiol 235:3823–3834. https://doi.org/10.1002/jcp.29277
Garufi A, Traversi G, Cirone M, D'Orazi G (2019) HIPK2 role in the tumor-host interaction: impact on fibroblasts transdifferentiation CAF-like. IUBMB life 71(12):2055–2061. https://doi.org/10.1002/iub.2144
Giulietti M, Occhipinti G, Principato G, Piva F (2017) Identification of candidate miRNA biomarkers for pancreatic ductal adenocarcinoma by weighted gene co-expression network analysis. Cell Oncol (Dordr) 40(2):181–192. https://doi.org/10.1007/s13402-017-0315-y
Hattinger CM, Patrizio MP, Tavanti E, Luppi S, Magagnoli F, Picci P, Serra M (2018) Genetic testing for high-grade osteosarcoma: a guide for future tailored treatments? Expert rev Mol Diagn 18(11):947–961. https://doi.org/10.1080/14737159.2018.1535903
Hill KE, Kelly AD, Kuijjer ML, Barry W, Rattani A, Garbutt CC, Kissick H, Janeway K, Perez-Atayde A, Goldsmith J, Gebhardt MC, Arredouani MS, Cote G, Hornicek F, Choy E, Duan Z, Quackenbush J, Haibe-Kains B, Spentzos D (2017) An imprinted non-coding genomic cluster at 14q32 defines clinically relevant molecular subtypes in osteosarcoma across multiple independent datasets. J Hematol Oncol 10(1):107. https://doi.org/10.1186/s13045-017-0465-4
Hoggarth ZE, Osowski DB, Freeberg BA, Garrett SH, Sens DA, Sens MA, Zhou XD, Zhang KK, Somji S (2018) The urothelial cell line UROtsa transformed by arsenite and cadmium display basal characteristics associated with muscle invasive urothelial cancers. PLoS One 13(12):e0207877. https://doi.org/10.1371/journal.pone.0207877
Kim M, Kim DJ (2018) GFRA1: a novel molecular target for the prevention of osteosarcoma Chemoresistance. Int J Mol Sci 19(4). https://doi.org/10.3390/ijms19041078
Ma XB, Zhong YP, Zheng Y, Jiang J, Wang YP (2018) Coexpression of CD5 and CD43 predicts worse prognosis in diffuse large B-cell lymphoma. Cancer med 7(9):4284–4295. https://doi.org/10.1002/cam4.1674
Messerschmitt PJ, Garcia RM, Abdul-Karim FW, Greenfield EM, Getty PJ (2009) Osteosarcoma. J Am Acad Orthop Surg 17(8):515–527. https://doi.org/10.5435/00124635-200908000-00005
Nodale C, Sheffer M, Jacob-Hirsch J, Folgiero V, Falcioni R, Aiello A, Garufi A, Rechavi G, Givol D, D'Orazi G (2012) HIPK2 downregulates vimentin and inhibits breast cancer cell invasion. Cancer Biol Ther 13(4):198–205. https://doi.org/10.4161/cbt.13.4.18694
Petrvalska O, Kosek D, Kukacka Z, Tosner Z, Man P, Vecer J, Herman P, Obsilova V, Obsil T (2016) Structural insight into the 14-3-3 protein-dependent inhibition of protein kinase ASK1 (apoptosis signal-regulating kinase 1). J Biol Chem 291(39):20753–20765. https://doi.org/10.1074/jbc.M116.724310
Pinero J, Bravo A, Queralt-Rosinach N, Gutierrez-Sacristan A, Deu-Pons J, Centeno E, Garcia-Garcia J, Sanz F, Furlong LI (2017) DisGeNET: a comprehensive platform integrating information on human disease-associated genes and variants. Nucleic Acids Res 45(D1):D833–D839. https://doi.org/10.1093/nar/gkw943
Poos K, Smida J, Maugg D, Eckstein G, Baumhoer D, Nathrath M, Korsching E (2015) Genomic heterogeneity of osteosarcoma - shift from single candidates to functional modules. PLoS one 10(4):e0123082. https://doi.org/10.1371/journal.pone.0123082
Qin Y, Hu Q, Ji S, Xu J, Dai W, Liu W, Xu W, Sun Q, Zhang Z, Ni Q, Yu X, Zhang B, Xu X (2019) Homeodomain-interacting protein kinase 2 suppresses proliferation and aerobic glycolysis via ERK/cMyc axis in pancreatic cancer. Cell Prolif 52(3):e12603. https://doi.org/10.1111/cpr.12603
Ricciardelli C, Lokman NA, Pyragius CE, Ween MP, Macpherson AM, Ruszkiewicz A, Hoffmann P, Oehler MK (2017) Keratin 5 overexpression is associated with serous ovarian cancer recurrence and chemotherapy resistance. Oncotarget 8(11):17819–17832. https://doi.org/10.18632/oncotarget.14867
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13(11):2498–2504. https://doi.org/10.1101/gr.1239303
Sizemore GM, Sizemore ST, Seachrist DD, Keri RA (2014) GABA(a) receptor pi (GABRP) stimulates basal-like breast cancer cell migration through activation of extracellular-regulated kinase 1/2 (ERK1/2). J Biol Chem 289(35):24102–24113. https://doi.org/10.1074/jbc.M114.593582
Subramanian K, Dierckx T, Khouri R, Menezes SM, Kagdi H, Taylor GP, Farre L, Bittencourt A, Kataoka K, Ogawa S, Van Weyenbergh J (2019) Decreased RORC expression and downstream signaling in HTLV-1-associated adult T-cell lymphoma/leukemia uncovers an antiproliferative IL17 link: a potential target for immunotherapy? Int J Cancer 144(7):1664–1675. https://doi.org/10.1002/ijc.31922
Szklarczyk D, Franceschini A, Wyder S, Forslund K, Heller D, Huerta-Cepas J, Simonovic M, Roth A, Santos A, Tsafou KP, Kuhn M, Bork P, Jensen LJ, von Mering C (2015) STRING v10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res 43(database issue):D447–D452. https://doi.org/10.1093/nar/gku1003
Verdina A, Di Rocco G, Virdia I, Monteonofrio L, Gatti V, Policicchio E, Bruselles A, Tartaglia M, Soddu S (2017) HIPK2-T566 autophosphorylation diversely contributes to UV- and doxorubicin-induced HIPK2 activation. Oncotarget 8(10):16744–16754. https://doi.org/10.18632/oncotarget.14421
Wada T (2018) Downregulation of CD5 and dysregulated CD8(+) T-cell activation. Pediatr Int 60(9):776–780. https://doi.org/10.1111/ped.13636
Wan X, Shi L, Ma X, Tang H, Wu G (2018) The elevated ASK1 expression inhibits proliferation and invasion in gastric Cancer HGC-27 cells. Anat rec (Hoboken) 301(11):1815–1819. https://doi.org/10.1002/ar.23906
Wang J, Ni J, Song D, Ding M, Huang J, Li W, He G (2019) MAT1 facilitates the lung metastasis of osteosarcoma through upregulation of AKT1 expression. Life Sci 234:116771. https://doi.org/10.1016/j.lfs.2019.116771
Wang Y, Fu R, Liu H, Wang H, Zhang T, Ding S, Zhang J, Gao S, Liu C, Wang J, Xing L, Wang H, Li L, Liu H, Ruan E, Song J, Wu Y, Guan J, Qu W, Shao Z (2014) memory B (CD5(+) CD19(+) CD27(+)) lymphocyte in patients with immune-related pancytopenia. Zhonghua Xue ye Xue Za Zhi 35(8):719–723. https://doi.org/10.3760/cma.j.issn.0253-2727.2014.08.011
Yin M, Zhou HJ, Zhang J, Lin C, Li H, Li X, Li Y, Zhang H, Breckenridge DG, Ji W, Min W (2017) ASK1-dependent endothelial cell activation is critical in ovarian cancer growth and metastasis. JCI Insight 2(18). https://doi.org/10.1172/jci.insight.91828
Zhao GS, Gao ZR, Zhang Q, Tang XF, Lv YF, Zhang ZS, Zhang Y, Tan QL, Peng DB, Jiang DM, Guo QN (2018) TSSC3 promotes autophagy via inactivating the Src-mediated PI3K/Akt/mTOR pathway to suppress tumorigenesis and metastasis in osteosarcoma, and predicts a favorable prognosis. J Exp Clin Cancer res 37(1):188. https://doi.org/10.1186/s13046-018-0856-6
Zhao Z, Jin G, Yao K, Liu K, Liu F, Chen H, Wang K, Gorja DR, Reddy K, Bode AM, Guo Z, Dong Z (2019) Aurora B kinase as a novel molecular target for inhibition the growth of osteosarcoma. Mol Carcinog 58(6):1056–1067. https://doi.org/10.1002/mc.22993
Acknowledgements
We give our sincere gratitude to the reviewers for their valuable suggestions.
Availability of data
The datasets generated/analysed during the current study are available.
Author information
Authors and Affiliations
Contributions
Yuan Li and Fengxiao Ge participated in the conception and design of the study. Yuan Li, Fengxiao Ge and Shuaihua Wang performed the analysis and interpretation of data. Yuan Li, Fengxiao Ge and Shuaihua Wang contributed to drafting the article. Yuan Li and Fengxiao Ge revised it critically for important intellectual content. Shuaihua Wang is the GUARANTOR for the article who accepts full responsibility for the work and/or the conduct of the study, had access to the data, and oversaw the decision to publish.
Corresponding author
Ethics declarations
Competing interest
The author declares no competing interest exists.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Li, Y., Ge, F. & Wang, S. Four genes predict the survival of osteosarcoma patients based on TARGET database. J Bioenerg Biomembr 52, 291–299 (2020). https://doi.org/10.1007/s10863-020-09836-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10863-020-09836-6