Abstract
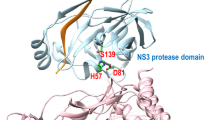
Due to the inherently flexible nature of a protein–protein interaction surface, it is difficult both to inhibit the association with a small molecule, and to predict how it might bind to the surface. In this study, we have examined small molecules that mediate the interaction between a WWI motif on the C-helix of HIV-1 glycoprotein-41 (gp41) and a deep hydrophobic pocket contained in the interior N-helical trimer. Association between these two components of gp41 leads to virus–cell and cell–cell fusion, which could be abrogated in the presence of an inhibitor that binds tightly in the pocket. We have studied a comprehensive combinatorial library of α-helical peptidomimetics, and found that compounds with strongly hydrophobic side chains had the highest affinity. Computational docking studies produced multiple possible binding modes due to the flexibility of both the binding site and the peptidomimetic compounds. We applied a transferred paramagnetic relaxation enhancement experiment to two selected members of the library, and showed that addition of a few experimental constraints enabled definitive identification of unique binding poses. Computational docking results were extremely sensitive to side chain conformations, and slight variations could preclude observation of the experimentally validated poses. Different receptor structures were required for docking simulations to sample the correct pose for the two compounds. The study demonstrated the sensitivity of predicted poses to receptor structure and indicated the importance of experimental verification when docking to a malleable protein–protein interaction surface.











Similar content being viewed by others
References
Ou HD, May AP, O’Shea CC (2011) Wiley Interdiscip Rev Syst Biol Med 3:48
Liu S, Wu S, Jiang S (2007) Curr Pharm Des 13:143
Welch BD, VanDemark AP, Heroux A, Hill CP, Kay MS (2007) Proc Natl Acad Sci USA 104:16828
Welch BD, Francis JN, Redman JS, Paul S, Weinstock MT, Reeves JD, Lie YS, Whitby FG, Eckert DM, Hill CP, Root MJ, Kay MS (2010) J Virol 84:11235
Eckert DM, Malashkevich VN, Hong LH, Carr PA, Kim PS (1999) Cell 99:103
Naider F, Anglister J (2009) Curr Opin Struct Biol 19:473
Allen KA, Rizzo M, Sadosty AT (2012) J Emerg Med 43:93
Jiang S, Lu H, Liu S, Zhao Q, He Y, Debnath AK (2004) Antimicrob Agents Chemother 48:4349
Liu K, Lu H, Hou L, Qi Z, Teixeira C, Barbault F, Fan BT, Liu S, Jiang S, Xie L (2008) J Med Chem 51:7843
Teixeira C, Barbault F, Rebehmed J, Liu K, Xie L, Lu H, Jiang S, Fan B, Maurel F (2008) Bioorg Med Chem 16:3039
Katritzky AR, Tala SR, Lu H, Vakulenko AV, Chen QY, Sivapackiam J, Pandya K, Jiang S, Debnath AK (2009) J Med Chem 52:7631
Jiang S, Debnath AK (2000) Biochem Biophys Res Commun 270:153
He Y, Liu S, Jing W, Lu H, Cai D, Chin DJ, Debnath AK, Kirchhoff F, Jiang S (2007) J Biol Chem 282:25631
Jahnke W, Rudisser S, Zurini M (2001) J Am Chem Soc 123:3149
Balogh E, Wu D, Zhou G, Gochin M (2009) J Am Chem Soc 131:2821
Gochin M, Zhou G, Phillips AH (2010) ACS Chem Biol 6:267
Gochin M, Guy RK, Case MA (2003) Angew Chem Int Ed 42:5325
Gochin M, Savage R, Hinckley S, Cai L (2006) Biol Chem 387:477
Cai L, Gochin M (2007) Antimicrob Agents Chemother 51:2388
Gochin M, Cai L (2009) J Med Chem 52:4338
Cai L, Balogh E, Gochin M (2009) Antimicrob Agents Chemother 53:2444
Gochin M (2012) Assay Drug Dev Technol 10:407
Shaginian A, Whitby LR, Hong S, Hwang I, Farooqi B, Searcey M, Chen J, Vogt PK, Boger DL (2009) J Am Chem Soc 131:5564
Whitby LR, Boyle KE, Cai L, Yu X, Gochin M, Boger DL (2012) Bioorg Med Chem Lett 22:2861
Cheng S, Tarby CM, Comer DD, Williams JP, Caporale LH, Myers PL, Boger DL (1996) Bioorg Med Chem 4:727
Boger DL, Tarby CM, Myers PL, Caporale LH (1996) J Am Chem Soc 118:2109
Cheng S, Comer DD, Williams JP, Myers PL, Boger DL (1996) J Am Chem Soc 118:2567
Boger DL, Desharnais J, Capps K (2003) Angew Chem Int Ed 42:4138
Wexler-Cohen Y, Shai Y (2007) Faseb J 21:3677
Platt EJ, Wehrly K, Kuhmann SE, Chesebro B, Kabat D (1998) J Virol 72:2855
Wei X, Decker JM, Liu H, Zhang Z, Arani RB, Kilby JM, Saag MS, Wu X, Shaw GM, Kappes JC (2002) Antimicrob Agents Chemother 46:1896
Ciminale V, Felber BK, Campbell M, Pavlakis GN (1990) AIDS Res Hum Retroviruses 6:1281
Meiboom S, Gill D (1958) Rev Sci Instrum 29:688
Carr HY, Purcell EM (1954) Phys Rev 94:630
Hoult D (1976) J Magn Reson 21:337
Tosner Z, Skoch A, Kowalewski J (2010) ChemPhysChem 11:638
Mulder FA, Skrynnikov NR, Hon B, Dahlquist FW, Kay LE (2001) J Am Chem Soc 123:967
Aue W, Bartholdi E, Ernst R (1976) J Chem Phys 64:2229
Solomon I, Bloembergen N (1956) J Chem Phys 25:261
Trott O, Olson AJ (2010) J Comput Chem 31:455
Caffrey M (2001) Biochim Biophys Acta 1536:116
Stewart KD, Huth JR, Ng TI, McDaniel K, Hutchinson RN, Stoll VS, Mendoza RR, Matayoshi ED, Carrick R, Mo H, Severin J, Walter K, Richardson PL, Barrett LW, Meadows R, Anderson S, Kohlbrenner W, Maring C, Kempf DJ, Molla A, Olejniczak ET (2010) Bioorg Med Chem Lett 20:612
Kleywegt GJ, Henrick K, Dodson EJ, van Aalten DM (2003) Structure 11:1051
Schwieters CD, Kuszewski JJ, Tjandra N, Clore GM (2003) J Magn Reson 160:65
Iwahara J, Schwieters CD, Clore GM (2004) J Am Chem Soc 126:5879
Weissenhorn W, Dessen A, Harrison SC, Skehel JJ, Wiley DC (1997) Nature 387:426
Chan DC, Fass D, Berger JM, Kim PS (1997) Cell 89:263
Caffrey M, Cai M, Kaufman J, Stahl SJ, Wingfield PT, Covell DG, Gronenborn AM, Clore GM (1998) EMBO J 17:4572
Springman EB personal communication
Penfold BR, White JCB (1959) Acta Cryst 12:130
Acknowledgments
We gratefully acknowledge the financial support of the National Institutes of Health (NS059403, GM087998, MG), (CA078045, DLB). We thank E. Balogh and D. Wu for technical assistance in the collection of data for Fig. 1 and Table 1. Molecular graphics images were produced using the UCSF Chimera package from the Resource for Biocomputing, Visualization, and Informatics at the University of California, San Francisco (supported by NIH P41 RR-01081). The authors also gratefully acknowledge use of the UC Berkeley Biomolecular NMR facility. The authors thank Dr. Eric Springman at Locus Pharmaceuticals (Ansaris) for providing the coordinates of 3p7k prior to publication.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gochin, M., Whitby, L.R., Phillips, A.H. et al. NMR-assisted computational studies of peptidomimetic inhibitors bound in the hydrophobic pocket of HIV-1 glycoprotein 41. J Comput Aided Mol Des 27, 569–582 (2013). https://doi.org/10.1007/s10822-013-9662-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-013-9662-6