Abstract
Purpose
Non-obstructive azoospermia (NOA) is an essential cause of male infertility for which treatment options are limited. The pathogenic mechanism of NOA, especially idiopathic NOA, remains unclear. Gene variations are associated with the occurrence of NOA. Our study was performed to investigate the genetic causes of NOA.
Methods
Whole exome sequencing (WES) was performed in two probands diagnosed with NOA from a Chinese family. Sanger sequencing was applied to verify the pathogenic variants. A minigene assay was carried out to identify the effect of the splicing variants.
Results
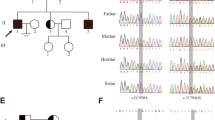
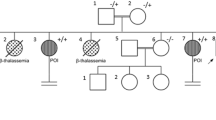
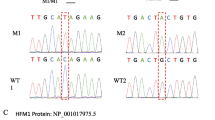
We detected a novel homozygous variant (c.2681-3 T > A) in the HFM1 gene in the two siblings diagnosed with NOA, and their parents carried heterozygous mutations in the same gene. The results of the minigene assay revealed this splicing variant results in exon25 of HFM1 being skipped, leading to a protein truncation (p.Trp894Cysfs*44).
Conclusion
Our results showed that a deleterious splicing variant in HFM1 was related to NOA in these two patients. This novel variant of HFM1 may serve as a potential genetic biomarker for NOA patients.



Similar content being viewed by others
Data availability
The authors declare the availability of data upon request.
References
Agarwal A, Baskaran S, Parekh N, Cho CL, Henkel R, Vij S, Arafa M, Panner Selvam MK, Shah R. Male infertility. Lancet. 2021;397(10271):319–33.
Tournaye H, Krausz C, Oates RD. Novel concepts in the aetiology of male reproductive impairment. Lancet Diabetes Endocrinol. 2017;5(7):544–53.
Cioppi F, Rosta V, Krausz C. Genetics of azoospermia. Int J Mol Sci. 2021;22(6):3264.
Ozturk S. Genetic variants underlying spermatogenic arrests in men with non-obstructive azoospermia. Cell Cycle. 2023;22(9):1021–61.
Chen S, Wang G, Zheng X, Ge S, Dai Y, Ping P, Chen X, Liu G, Zhang J, Yang Y, Zhang X, Zhong A, Zhu Y, Chu Q, Huang Y, Zhang Y, Shen C, Yuan Y, Yuan Q, Pei X, Cheng CY, Sun F. Whole-exome sequencing of a large Chinese azoospermia and severe oligospermia cohort identifies novel infertility causative variants and genes. Hum Mol Genet. 2020;29(14):2451–9.
Kherraf ZE, Cazin C, Bouker A, Fourati Ben Mustapha S, Hennebicq S, Septier A, Coutton C, Raymond L, Nouchy M, Thierry-Mieg N, Zouari R, Arnoult C, Ray PF. Whole-exome sequencing improves the diagnosis and care of men with non-obstructive azoospermia. Am J Hum Genet. 2022;109(3):508–17.
Kasak L, Punab M, Nagirnaja L, Grigorova M, Minajeva A, Lopes AM, Punab AM, Aston KI, Carvalho F, Laasik E, Smith LB, Conrad DF, Laan M. Bi-allelic recessive loss-of-function variants in FANCM cause non-obstructive azoospermia. Am J Hum Genet. 2018;103(2):200–12.
Li Y, Wu Y, Zhou J, Zhang H, Zhang Y, Ma H, Jiang X, Shi Q. A recurrent ZSWIM7 mutation causes male infertility resulting from decreased meiotic recombination. Hum Reprod. 2021;36(5):1436–45.
Wang W, Meng L, He J, Su L, Li Y, Tan C, Xu X, Nie H, Zhang H, Du J, Lu G, Luo M, Lin G, Tu C, Tan YQ. Bi-allelic variants in SHOC1 cause non-obstructive azoospermia with meiosis arrest in humans and mice. Mol Hum Reprod. 2022;28(6):gaac015.
Fan S, Jiao Y, Khan R, Jiang X, Javed AR, Ali A, Zhang H, Zhou J, Naeem M, Murtaza G, Li Y, Yang G, Zaman Q, Zubair M, Guan H, Zhang X, Ma H, Jiang H, Ali H, Dil S, Shah W, Ahmad N, Zhang Y, Shi Q. Homozygous mutations in C14orf39/SIX6OS1 cause non-obstructive azoospermia and premature ovarian insufficiency in humans. Am J Hum Genet. 2021;108(2):324–36.
Guiraldelli MF, Eyster C, Wilkerson JL, Dresser ME, Pezza RJ. Mouse HFM1/Mer3 is required for crossover formation and complete synapsis of homologous chromosomes during meiosis. PLoS Genet. 2013;9(3):e1003383.
Xie X, Murtaza G, Li Y, Zhou J, Ye J, Khan R, Jiang L, Khan I, Zubair M, Yin H, Jiang H, Liu W, Shi B, Hou X, Gong C, Fan S, Wang Y, Jiang X, Zhang Y, Zhang H, Ma H, Shi Q. Biallelic HFM1 variants cause non-obstructive azoospermia with meiotic arrest in humans by impairing crossover formation to varying degrees. Hum Reprod. 2022;37(7):1664–77.
Saebnia N, Ebrahimzadeh-Vesal R, Haddad-Mashhadrizeh A, Gholampour-Faroji N, Schinzel A, Neshati Z, Azimi-Nezhad M. Identification of a new splice-acceptor mutation in HFM1 and functional analysis through molecular docking in nonobstructive azoospermia. J Assist Reprod Genet. 2022;39(5):1195–203.
Tang D, Lv M, Gao Y, Cheng H, Li K, Xu C, Geng H, Li G, Shen Q, Wang C, He X, Cao Y. Novel variants in helicase for meiosis 1 lead to male infertility due to non-obstructive azoospermia. Reprod Biol Endocrinol. 2021;19(1):129.
Tang F, Gao Y, Li K, Tang D, Hao Y, Lv M, Wu H, Cheng H, Fei J, Jin Z, Wang C, Xu Y, Wei Z, Zhou P, Zhang Z, He X, Cao Y. Novel deleterious splicing variant in HFM1 causes gametogenesis defect and recurrent implantation failure: concerning the risk of chromosomal abnormalities in embryos[J]. J Assist Reprod Genet. 2023;40(7):1689–702.
Yu L, Li M, Zhang H, He Q, Wan F, Zhang C, Wang F. Novel pathogenic splicing variants in helicase for meiosis 1 (HFM1) are associated with diminished ovarian reserve and poor pregnancy outcomes. J Assist Reprod Genet. 2022;39(9):2135–41.
Liu H, Wei X, Sha Y, Liu W, Gao H, Lin J, Li Y, Tang Y, Wang Y, Wang Y, Su Z. Whole-exome sequencing in patients with premature ovarian insufficiency: early detection and early intervention. J Ovarian Res. 2020;13(1):114.
Zhang W, Song X, Ni F, Cheng J, Wu BL, Jiang H. Association analysis between HFM1 variations and idiopathic azoospermia or severe oligozoospermia in Chinese Men. Sci China Life Sci. 2017;60(3):315–8.
Acknowledgements
We would like to thank all the researchers for their contributions.
Funding
This article was funded by the National Natural Science Foundation of China (grant numbers: 82071638; 82001618).
Author information
Authors and Affiliations
Contributions
Ninghong Song and Jinzhao Ma designed the project. Liangyu Yao and Yifeng Ge drafted the manuscript. Tian Du participated in the collection of clinical information and revision work. Liangyu Yao carried out the experiments. Ninghong Song and Jinzhao Ma guided the experiment directions and edited the manuscript. All authors read and approved the final manuscript version.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
10815_2023_2907_MOESM2_ESM.pdf
Supplementary file2 Supplementary Fig. S1: Splicing effect of the c.2681-3T>A variant in HFM1 was scored by varSEAK. (PDF 1194 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yao, L., Ge, Y., Du, T. et al. A novel splicing mutation in helicase for meiosis 1 leads to non-obstructive azoospermia. J Assist Reprod Genet 40, 2493–2498 (2023). https://doi.org/10.1007/s10815-023-02907-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10815-023-02907-8