Abstract
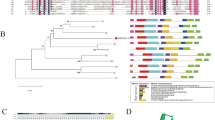
Nitrate reductase (NR) is the key rate-limiting enzyme of the nitrogen (N) assimilation process in plants, which has not been characterized in bread wheat under nitrogen stress, especially with respect to their homeologues. Total 9 NRs were identified and classified into three groups, which showed a close relationship with different wheat ancestors. The occurrence of N-responsive cis-acting regulatory elements like MYB, MYC, G-Box, and GATA-motif confirmed their N-responsiveness. Expression of all three groups of NR under N-stress revealed the NR 6-1ABD group to be the most N-responsive, which was characterized further in detail. The study was carried out in two genotypes contrasting for their N-responsiveness (HD 2967: Highly responsive to applied N, and Choti Lerma: Less responsive to applied N) selected on the basis of field evaluation. Homeologous differences within a genotype were found much more than the genotypic differences of a specific homeologue coding sequence. Among the three homeologues, though NR 6-1D homeologue was found to be most responsive to N-stress, the contribution was maximum for this homeologue, followed by NR 6-1A and least by NR 6-1B. We found that the expression of homeologues was linked to the presence of N-responsive cis-elements. All the homeologues of NR 6-1 in Choti Lerma were found less responsive to N-stress in comparison to HD 2967, which might also be linked to N-use efficiency. Homeologous expression of NR 6-1ABD revealed the negligible contribution of B-homeologue to N-stress. Homeologous differences of NR 6-1ABD was found much more than the genotypic differences. Hence, our study on wheat NR will be helpful in manipulating the specific homeologue of the NR gene in the future.













Similar content being viewed by others
Data availability
The CDS sequences of the genes were submitted to GenBank NCBI and the corresponding gene accession numbers are provided in the text and can be obtained in the NCBI nucleotide Database (https://www.ncbi.nlm.nih.gov/nucleotide/). All data generated or analyzed during the present study are included in this manuscript and its supplementary information files.
Abbreviations
- CDS:
-
Coding sequence
- mmol/L:
-
Millimolar/litre
- h:
-
Hour
- cDNA:
-
Complementary deoxy-ribonucleic acid
- dNTPs:
-
Deoxynucleoside triphosphates
- PCR:
-
Polymerase chain reaction
- qPCR:
-
Quantitative-polymerase chain reaction
- Tm :
-
Melting temperature
- °C:
-
Degree celsius
- ORF:
-
Open reading frame
- MW:
-
Molecular weight
- pI:
-
Isoelectric point
- NCBI:
-
National Center for Biotechnology Information
- µl:
-
Microlitre
- ng:
-
Nanogram
- bp:
-
Basepair
- UTR:
-
Untranslated region
References
Alipour H, Bihamta MR, Mohammadi V, Peyghambari SA, Bai G, Zhang G (2017) Genotyping-by-sequencing (GBS) revealed molecular genetic diversity of Iranian wheat landraces and cultivars. Front Plant Sci 8:1293
Balotf S, Kavoosi G, Kholdebarin B (2016) Nitrate reductase, nitrite reductase, glutamine synthetase, and glutamate synthase expression and activity in response to different nitrogen sources in nitrogen-starved wheat seedlings. Biotechnol Appl Biochem 63(2):220–229
Bharati A, Tehlan G, Nagar CK, Sinha SK, Venkatesh K, Mandal PK (2022) Nitrogen stress-induced TaDof1 expression in diverse wheat genotypes and its relation with nitrogen use efficiency. Cereal Res Commun 50:637–645
Blackwood GC, Hallam R (1979) Nitrate reductase activity in wheat (Triticum aestivum L.) II. The correlation with yield. New Phytol 82(2):417–425
Botella MA, Cruz C, Martins-Louçao MA, Cerdá A (1993) Nitrate reductase activity in wheat seedlings as affected by NO3-/NH4+ ratio and salinity. J Plant Physiol 142(5):531–536
Bottley A, Xia GM, Koebner RMD (2006) Homoeologous gene silencing in hexaploid wheat. Plant J 47(6):897–906
Campbell WH (1996) Nitrate reductase biochemistry comes of age. Plant Physiol 111(2):355
Campbell WH (2001) Structure and function of eukaryotic NAD (P) H: nitrate reductase. Cell Mol Life Sci CMLS 58:194–204
Carillo P, Mastrolonardo G, Nacca F, Fuggi A (2005) Nitrate reductase in durum wheat seedlings as affected by nitrate nutrition and salinity. Funct Plant Biol 32(3):209–219
Chandna R, Kaur G, Iqbal M, Khan I, Ahmad A (2012) Differential response of wheat genotypes to applied nitrogen: biochemical and molecular analysis. Arch Agron Soil Sci 58(8):915–929
Chattaraj S, Chakraborty D, Garg RN, Singh GP, Gupta VK, Singh S, Singh R (2013) Hyperspectral remote sensing for growth-stage-specific water use in wheat. Field Crop Res 144:179–191
Choi HK, Kleinhofs A, An G (1989) Nucleotide sequence of rice nitrate reductase genes. Plant Mol Biol 13:731–733
Daniel-Vedele F, Dorbe M-F, Caboche M, Rouzé P (1989) Cloning and analysis of the tomato nitrate reductase-encoding gene: protein domain structure and amino acid homologies in higher plants. Gene 85(2):371–380
dos Santos RL, Freire FJ, de Oliveira ECA, dos Freire MBGS, West JB, de Barbosa JA, de Moura MJA, da Bezerra PC (2019) Nitrate reductase activity and nitrogen and biomass accumulation in sugarcane under molybdenum and nitrogen fertilization. Revista Brasileira de Ciência Do Solo 43:e0180171
Fa CUI, Chun-Hua Z, Yin-Guang BAO, Hao Z, Yu-Hai W, Qing-Zhuan W, Bin DU, Hang-Yun MA, Hong-Gang W (2010) Genetic differences in homoeologous group 1 of seven types of winter wheat Aimengniu. Acta Agron Sin 36(9):1450–1456
Fan X, Naz M, Fan X, Xuan W, Miller AJ, Xu G (2017) Plant nitrate transporters: from gene function to application. J Exp Bot 68(10):2463–2475
Foulkes MJ, Hawkesford MJ, Barraclough PB, Holdsworth MJ, Kerr S, Kightley S, Shewry PR (2009) Identifying traits to improve the nitrogen economy of wheat: recent advances and future prospects. Field Crop Res 114(3):329–342
Gaju O, Allard V, Martre P, Snape JW, Heumez E, LeGouis J, Moreau D, Bogard M, Griffiths S, Orford S (2011) Identification of traits to improve the nitrogen-use efficiency of wheat genotypes. Field Crop Res 123(2):139–152
Gao S, Li L, Han X, Liu T, Jin P, Cai L, Xu M, Zhang T, Zhang F, Chen J (2021) Genome-wide identification of the histone acetyltransferase gene family in Triticum aestivum. BMC Genomics 22:1–17
Garcia-Oliveira AL, Benito C, Prieto P, de Andrade Menezes R, Rodrigues-Pousada C, Guedes-Pinto H, Martins-Lopes P (2013) Molecular characterization of TaSTOP1 homoeologues and their response to aluminium and proton (H+) toxicity in bread wheat (Triticum aestivum L.). BMC Plant Biol 13:1–13
Gayatri, Rani M, Mahato AK, Sinha SK, Dalal M, Singh NK, Mandal PK (2019) Homeologue specific gene expression analysis of two vital carbon metabolizing enzymes-citrate synthase and NADP-isocitrate dehydrogenase-from wheat (Triticum aestivum L.) under nitrogen stress: homeologue specific gene expression of CS and NADP-ICDH. Appl Biochem Biotechnol 188:569–584
Gayatri, Venkatesh K, Sinha SK, Roy P, Mandal PK (2021) Molecular characterization of GS2 and Fd-GOGAT homeologues and their biased response to nitrogen stress in bread wheat (Triticum aestivum L.). J Plant Growth Regul 41:2555–2569
Golberg AD, Jonas OA, Pereyra MC, Cabeza C, Ledent JF (1995) Nitrate reductase activity in nitrogen and water-stressed plants of bread wheat. Cereal Res Commun 23:433–439
Good AG, Shrawat AK, Muench DG (2004) Can less yield more? Is reducing nutrient input into the environment compatible with maintaining crop production? Trends Plant Sci 9(12):597–605
Guerrero MG, Vega JM, Losada M (1981) The assimilatory nitrate-reducing system and its regulation. Annu Rev Plant Physiol 32(1):169–204
Hageman RH (1979) Integration of nitrogen assimilation in relation to yield. In: Hewitt EJ, Cutting CV (ed) Nitrogen Assimilation of Plants. Academic Press, London, pp 591–611
Hawkesford MJ (2014) Reducing the reliance on nitrogen fertilizer for wheat production. J Cereal Sci 59(3):276–283
Hawkesford MJ, Griffiths S (2019) Exploiting genetic variation in nitrogen use efficiency for cereal crop improvement. Curr Opin Plant Biol 49:35–42
He X, Zhang H, Ye X, Hong J, Ding G (2021) Nitrogen assimilation related genes in Brassica napus: systematic characterization and expression analysis identified hub genes in multiple nutrient stress responses. Plants 10(10):2160
Himi E, Noda K (2004) Isolation and location of three homoeologous dihydroflavonol-4-reductase (DFR) genes of wheat and their tissue-dependent expression. J Exp Bot 55(396):365–375
Hirel B, Le Gouis J, Ney B, Gallais A (2007) The challenge of improving nitrogen use efficiency in crop plants: towards a more central role for genetic variability and quantitative genetics within integrated approaches. J Exp Bot 58(9):2369–2387
Hiscox JD, Israelstam GF (1979) A method for the extraction of chlorophyll from leaf tissue without maceration. Can J Bot 57(12):1332–1334
Hu Z, Yu Y, Wang R, Yao Y, Peng H, Ni Z, Sun Q (2011) Expression divergence of TaMBD2 homoeologous genes encoding methyl CpG-binding domain proteins in wheat (Triticum aestivum L.). Gene 471(1–2):13–18
Hurali DT, Bhurta R, Tyagi S, Sathee L, Sandeep AB, Singh D, Mallick N, Vinod X, Jha SK (2022) Analysis of NIA and GSNOR family genes and nitric oxide homeostasis in response to wheat-leaf rust interaction. Sci Rep 12(1):803
Islam MQ, Hasan MN, Hoque H, Jewel NA, Bhuiyan MFH, Prodhan SH (2021) Characterization of transcription factor MYB59 and expression profiling in response to low K+ and NO3− in indica rice (Oryza sativa L.). J Genet Eng Biotechnol 19(1):1–14
Juery C, Concia L, De Oliveira R, Papon N, Ramírez-González R, Benhamed M, Uauy C, Choulet F, Paux E (2021) New insights into homoeologous copy number variations in the hexaploid wheat genome. Plant Genome 14(1):e20069
Kashkush K, Feldman M, Levy AA (2002) Gene loss, silencing and activation in a newly synthesized wheat allotetraploid. Genetics 160(4):1651–1659
Kavoosi G, Balotf S, Eshghi H, Hasani H (2014) Analysis of nitrate reductase mRNA expression and nitrate reductase activity in response to nitrogen supply. Mol Biol Res Commun 3(2):75
Konishi M, Yanagisawa S (2010) Identification of a nitrate-responsive cis-element in the Arabidopsis NIR1 promoter defines the presence of multiple cis-regulatory elements for nitrogen response. Plant J 63(2):269–282
Konishi M, Yanagisawa S (2013) Arabidopsis NIN-like transcription factors have a central role in nitrate signalling. Nat Commun 4(1):1617
Kumar A, Kumar S, Venkatesh K, Singh NK, Mandal PK, Sinha SK (2022) Physio-molecular traits of contrasting bread wheat genotypes associated with 15N influx exhibiting homeolog expression bias in nitrate transporter genes under different external nitrate concentrations. Planta 255(5):104
Kumari S (2011) Yield response of uniculm wheat (Triticum aestivum L.) to early and late application of nitrogen: flag leaf development and senescence. J Agric Sci 3(1):170
Lea PJ, Miflin BJ (2003) Glutamate synthase and the synthesis of glutamate in plants. Plant Physiol Biochem 41(6–7):555–564
Lea US, Slimestad R, Smedvig P, Lillo C (2007) Nitrogen deficiency enhances expression of specific MYB and bHLH transcription factors and accumulation of end products in the flavonoid pathway. Planta 225:1245–1253
Lejay L, Tillard P, Lepetit M, Olive FD, Filleur S, Daniel-Vedele F, Gojon A (1999) Molecular and functional regulation of two NO3–uptake systems by N-and C-status of Arabidopsis plants. Plant J 18(5):509–519
Lescot M, Déhais P, Thijs G, Marchal K, Moreau Y, Van de Peer Y, Rouzé P, Rombauts S (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30(1):325–327
Li H, Hu B, Chu C (2017) Nitrogen use efficiency in crops: lessons from Arabidopsis and rice. J Exp Bot 68(10):2477–2488
Lowry OH, Rosenbrough NJ, Farr AL, Randall RJ (1951) Protein measurement with Folin reagent. J Biol Chem 193:265–273
Lv Z, Li Z, Wang M, Zhao F, Zhang W, Li C, Gong L, Zhang Y, Mason AS, Liu B (2021) Conservation and trans-regulation of histone modification in the A and B subgenomes of polyploid wheat during domestication and ploidy transition. BMC Biol 19(1):1–16
Mălinaş A, Vidican R, Rotar I, Mălinaş C, Moldovan CM, Proorocu M (2022) Current status and future prospective for nitrogen use efficiency in wheat (Triticum aestivum L.). Plants 11(2):217
Marcussen T, Sandve SR, Heier L, Spannagl M, Pfeifer M, International Wheat Genome Sequencing Consortium, Jakobsen KS, Wulff BBH, Steuernagel B, Mayer KFX (2014) Ancient hybridizations among the ancestral genomes of bread wheat. Science 345(6194):1250092
Minotti PL, Jackson WA (1970) Nitrate reduction in the roots and shoots of wheat seedlings. Planta 95:36–44
Mochida K, Yamazaki Y, Ogihara Y (2004) Discrimination of homoeologous gene expression in hexaploid wheat by SNP analysis of contigs grouped from a large number of expressed sequence tags. Mol Genet Genomics 270:371–377
Moll RH, Kamprath EJ, Jackson WA (1982) Analysis and interpretation of factors which contribute to efficiency of nitrogen utilization 1. Agron J 74(3):562–564
Morenikeji OB, Capria AL, Ojurongbe O, Thomas BN (2020) SNP diversity in CD14 gene promoter suggests adaptation footprints in trypanosome tolerant N’Dama (Bos taurus) but not in susceptible White Fulani (Bos indicus) cattle. Genes 11(1):112
Nagar CK, Gayatri AB, Mandal PK (2018) Nitrogen stress induced changes in root system architecture (RSA) in diverse wheat (T. aestivum L.) genotypes at seedling stage. Wheat Barley Res 10(2):93–101
Nathawat NS, Kuhad MS, Goswami CL, Patel AL, Kumar R (2005) Nitrogen-metabolizing enzymes: effect of nitrogen sources and saline irrigation. J Plant Nutr 28(6):1089–1101
Ng PC, Henikoff S (2003) SIFT: Predicting amino acid changes that affect protein function. Nucleic Acids Res 31(13):3812–3814
Nikolic M, Cesco S, Monte R, Tomasi N, Gottardi S, Zamboni A, Pinton R, Varanini Z (2012) Nitrate transport in cucumber leaves is an inducible process involving an increase in plasma membrane H+-ATPase activity and abundance. BMC Plant Biol 12:1–12
Nomura T, Ishihara A, Yanagita RC, Endo TR, Iwamura H (2005) Three genomes differentially contribute to the biosynthesis of benzoxazinones in hexaploid wheat. Proc Natl Acad Sci 102(45):16490–16495
Panchy N, Lehti-Shiu M, Shiu S-H (2016) Evolution of gene duplication in plants. Plant Physiol 171(4):2294–2316
Raghuram N, Sharma N (2019) Improving crop nitrogen use efficiency. In: Moo-Young M (ed) Comprehensive Biotechnology, vol 4. Pergamon, Elsevier, pp 211–220
Rathour M, Sharma A, Kaur A, Upadhyay SK (2020) Genome-wide characterization and expression and co-expression analysis suggested diverse functions of WOX genes in bread wheat. Heliyon 6(12):e05762
Rawal N, Pande KR, Shrestha R, Vista SP (2022) Nutrient use efficiency (NUE) of wheat (Triticum aestivum L.) as affected by NPK fertilization. PLoS ONE 17(1):e0262771
Rimbert H, Darrier B, Navarro J, Kitt J, Choulet F, Leveugle M, Duarte J, Rivière N, Eversole K, Consortium, I. W. G. S (2018) High throughput SNP discovery and genotyping in hexaploid wheat. PLoS ONE 13(1):e0186329
Rolly NK, Yun B-W (2021) Regulation of nitrate (NO3) transporters and glutamate synthase-encoding genes under drought stress in arabidopsis: the regulatory role of AtbZIP62 transcription factor. Plants 10(10):2149
Roorkiwal M, Nayak SN, Thudi M, Upadhyaya HD, Brunel D, Mournet P, This D, Sharma PC, Varshney RK (2014) Allele diversity for abiotic stress responsive candidate genes in chickpea reference set using gene based SNP markers. Front Plant Sci 5:248
Rosales EP, Iannone MF, Groppa MD, Benavides MP (2011) Nitric oxide inhibits nitrate reductase activity in wheat leaves. Plant Physiol Biochem 49(2):124–130
Sharma N, Sinha VB, Prem Kumar NA, Subrahmanyam D, Neeraja CN, Kuchi S, Jha A, Parsad R, Sitaramam V, Raghuram N (2021) Nitrogen use efficiency phenotype and associated genes: roles of germination, flowering, root/shoot length and biomass. Front Plant Sci 11:587464
Sherrard JH, Dalling MJ (1979) In vitro stability of nitrate reductase from wheat leaves: I. Stability of highly purified enzyme and its component activities. Plant Physiol 63(2):346–353
Shitsukawa N, Tahira C, Kassai K, Hirabayashi C, Shimizu T, Takumi S, Mochida K, Kawaura K, Ogihara Y, Murai K (2007) Genetic and epigenetic alteration among three homoeologous genes of a class E MADS box gene in hexaploid wheat. Plant Cell 19(6):1723–1737
Sinha SK, Rani M, Bansal N, Gayatri, Venkatesh K, Mandal PK (2015) Nitrate starvation induced changes in root system architecture, carbon: nitrogen metabolism, and miRNA expression in nitrogen-responsive wheat genotypes. Appl Biochem Biotechnol 177:1299–1312
Sinha SK, Rani M, Kumar A, Kumar S, Venkatesh K, Mandal PK (2018) Natural variation in root system architecture in diverse wheat genotypes grown under different nitrate conditions and root growth media. Theor Exp Plant Physiol 30:223–234
Sinha SK, Kumar A, Tyagi A, Venkatesh K, Paul D, Singh NK, Mandal PK (2020) Root architecture traits variation and nitrate-influx responses in diverse wheat genotypes under different external nitrogen concentrations. Plant Physiol Biochem 148:246–259
Srivastava HS (1980) Regulation of nitrate reductase activity in higher plants. Phytochemistry 19(5):725–733
Stitt M, Müller C, Matt P, Gibon Y, Carillo P, Morcuende R, Scheible W, Krapp A (2002) Steps towards an integrated view of nitrogen metabolism. J Exp Bot 53(370):959–970
Tang X, Peng Y, Li Z, Guo H, Xia X, Li B, Yin W (2022) The regulation of nitrate reductases in response to abiotic stress in Arabidopsis. Int J Mol Sci 23(3):1202
Tovkach A, Ryan PR, Richardson AE, Lewis DC, Rathjen TM, Ramesh S, Tyerman SD, Delhaize E (2013) Transposon-mediated alteration of TaMATE1B expression in wheat confers constitutive citrate efflux from root apices. Plant Physiol 161(2):880–892
Tyagi BS, Foulkes J, Singh G, Sareen S, Kumar P, Broadley MR, Gupta V, Krishnappa G, Ojha A, Khokhar JS (2020) Identification of wheat cultivars for low nitrogen tolerance using multivariable screening approaches. Agronomy 10(3):417
Vaucheret H, Kronenberger J, Rouze P, Caboche M (1989) Complete nucleotide sequence of the two homeologous tobacco nitrate reductase genes. Plant Mol Biol 12(5):597–600
Wang P, Du Y, Song C-P (2011) Phosphorylation by MPK6: a conserved transcriptional modification mediates nitrate reductase activation and NO production? Plant Signal Behav 6(6):889–891
Wang Y, Pan F, Chen D, Chu W, Liu H, Xiang Y (2017) Genome-wide identification and analysis of the Populus trichocarpa TIFY gene family. Plant Physiol Biochem 115:360–371
Winfield MO, Wilkinson PA, Allen AM, Barker GLA, Coghill JA, Burridge A, Hall A, Brenchley RC, D’Amore R, Hall N (2012) Targeted re-sequencing of the allohexaploid wheat exome. Plant Biotechnol J 10(6):733–742
Witko A (2003). In World (Vol. 3, Issue February 2004)
Zhang X, Zong J, Liu J, Yin J, Zhang D (2010) Genome-wide analysis of WOX gene family in rice, sorghum, maize, Arabidopsis and poplar. J Integr Plant Biol 52(11):1016–1026
Zhang C, Hou Y, Hao Q, Chen H, Chen L, Yuan S, Shan Z, Zhang X, Yang Z, Qiu D (2015) Genome-wide survey of the soybean GATA transcription factor gene family and expression analysis under low nitrogen stress. PLoS ONE 10(4):e0125174
Zhao X-Q, Nie X-L, Xiao X-G (2013) Over-expression of a tobacco nitrate reductase gene in wheat (Triticum aestivum L.) increases seed protein content and weight without augmenting nitrogen supplying. PLoS ONE 8(9):e74678
Zhou J, Kleinhofs A (1996) Molecular evolution of nitrate reductase genes. J Mol Evol 42:432–442
Zhou C, Dong Z, Zhang T, Wu J, Yu S, Zeng Q, Han D, Tong W (2020) Genome-scale analysis of homologous genes among subgenomes of bread wheat (Triticum aestivum L.). Int J Mol Sci 21(8):3015
Acknowledgements
We would like to thank to the Department of Biotechnology, Government of India (BT/IN/UK-VNC/43/KV/2015-16) for funding under the project “Indo-UK Centre for the Improvement of Nitrogen Use Efficiency in Wheat (INEW)” and Director, NIPB, New Delhi for providing all the facilities. Authors also want to thank P.G. School, ICAR-Indian Agricultural Research Institute since part of the work is from Mr. Megavath Ravi’s M.Sc. thesis, submitted to this Institute.
Funding
We are thankful to the Department of Biotechnology, Government of India (BT/IN/UK-VNC/43/KV/2015-16) for funding under the project “Indo-UK Centre for the Improvement of Nitrogen Use Efficiency in Wheat (INEW)”.
Author information
Authors and Affiliations
Contributions
G and MR designed and performed the experiments. G wrote the manuscript. HC performed the hydroponics experiment. EM constructed circos. SP edited the manuscript. KV provided seeds of HD 2967 and CL genotypes and all relevant data generated from the field conditions of both the genotypes, edited the manuscript. PKM conceptualized the research objectives, designed the experiments, supervised the entire work, and revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Additional information
Communicated by Ben Zhang.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gayatri, Ravi, M., Chauhan, H. et al. Characterization and expression analysis of nitrate reductase 6-1ABD gene in hexaploid bread wheat under different nitrogen regimes. Plant Growth Regul 103, 81–100 (2024). https://doi.org/10.1007/s10725-023-01082-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-023-01082-1