Abstract
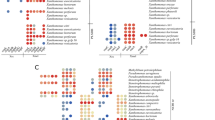
In this study, we explored the multiple heavy metal-resistant yeast isolated from heavy metal-polluted environment. The isolated yeast showed maximum growth at 30 °C, pH 7.0, and the strain was identified as Candida tropicalis through 18S ribosomal RNA (rRNA) gene sequence analysis. Yeast cells grew well in medium containing different concentrations of heavy metal ions [CdCl2, Pb(NO3)2, NaAsO2, CuSO4 and K2Cr2O7]. Minimum inhibitory concentration (MIC) against different metal ions was ranged from 5 to 19 mM, and the metal resistance value against each metal observed by yeast cells was 5 mM (Cr), 10 mM (Cd), 15 mM (As), 14 mM (Cu) and 19 mM (Pb) and increased in the following order: Pb > Cu > As ≥ Cd > Cr. The total cellular glutathione, GSH/GSSG redox couple and metallothioneins like protein (MT) were assayed by growing cultures for 24 h and exposed to 100 mg/L of each heavy metal ion. Remarkable increase in γ-glutamylcysteinylglycine (GSH) level was determined in arsenic and cadmium treatment followed by chromium, lead and copper. Stressed cells had much more oxidized GSH than unstressed cells. GSH/GSSG ratio was significantly increased in cadmium and copper treatment in contrast to chromium, arsenic and lead. Statistical analysis revealed significantly higher cysteine level in all metal-treated samples as compared to control. Antioxidant glutathione transferase activity was not detected in metal-treated and untreated yeast samples. One-dimensional electrophoresis of proteins revealed marked differences in banding pattern of heavy metal-exposed yeast samples. A prominent 20 kDa band was observed in all treated samples suggesting that some differential proteins could be over-expressed during heavy metal treatment and might be involved in cell resistance mechanisms.




Similar content being viewed by others
References
Balsalobre, L., De-Siloniz, M. I., Validerrama, M. J., Benito, T., Larrea, M. T., & Peinado, J. M. (2003). Occurrence of yeasts in municipal wastes and their behaviour in presence of cadmium copper and zinc. Journal of Basic Microbiology, 43, 185–193.
Berdicevsky, I., Duek, L., Merzbach, D., & Yannai, S. (1993). Susceptibility of different yeast species to environmental toxic metals. Environmental Pollution, 80(1), 41–44.
Blackwell, K. J., Singlenton, I., & Tobin, J. M. (1995). Metal cation uptake by yeast: a review. Applied Journal of Microbiology and Biotechnology, 43, 579–584.
Bradford, M. M. (1976). Rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Analytical Biochemistry, 72, 248–254.
Cuozzo, J. W., & Kaiser, C. A. (1999). Competition between glutathione and protein thiols for disufide bond formation. Nature Cell Biology, 1, 130–135.
Edward Raja, C., Anbazhagan, K., & Selvam, G. S. (2006). Isolation and characterization of a metal resistant Pseudomonas aeruginosa strain. World Journal of Microbiology and Biotechnology, 22, 577–586.
Fauchon, M., Lagniel, G., Aude, J. C., Lombardia, L., Soularue, P., Petat, C., Marguerie, G., Sentenac, A., Werner, M., & Labarre, J. (2002). Sulfur sparing in the yeast proteome in response to sulfur demand. Molecular Cell, 9, 713–723.
Filomeni, G., Rotilio, G., & Ciriolo, M. R. (2002). Cell signalling and the glutathione redox system. Biochemical Pharmacology, 64, 1057–1064.
Freeman, M. L., Huntley, S. A., Meredith, M. J., Senisterra, G. A., & Lepock, J. (1997). Destabilization and denaturation of cellular protein by glutathione depletion. Cell Stress and Chaperones, 2, 191–198.
Fujs, S., Gazdag, Z., Poijsak, B., Stibilj, V., Milacic, R., Pesti, M., Raspor, P., & Batic, M. (2005). The oxidative stress response of the yeast Candida intermedia to copper, zinc, and selenium exposure. Journal of Basic Microbiology, 45, 125–135.
Gardiner, C. S., & Reed, D. J. (1994). Status of glutathione during oxidant-induced oxidative stress in the pre implantation mouse embryo. Biology of Reproduction, 51, 1307–1314.
Gharieb, M. M., & Gadd, G. M. (2004). Role of glutathione in detoxification of metal (loid)s by Saccharomyces cerevisiae. Biometals, 17, 183–188.
Grant, C. M., MacIver, F. H., & Dawes, I. W. (1997). Glutathione synthetase is dispensable for growth under both normal and oxidative stress conditions in the yeast Saccharomyces cerevisiae due to an accumulation of the dipeptide-glutamylcysteine. Molecular Biology of the Cell, 8, 1699–1707.
Habig, W. H., Pabst, M. J., & Jakoby, W. B. (1974). Glutathione S-transferases. The first enzymatic step in mercapturic acid formation. Journal of Biological Chemistry, 249, 7130–7139.
Huang, K. P., & Huang, F. L. (2002). Glutathionylation of proteins by glutathione disulfide S-oxide. Biochemical Pharmacology, 64(5–6), 1049–1056.
Ilyas, S., Rehman, A., Varela, A. C., & Sheehan, D. (2014). Redox proteomics changes in the fungal pathogen Trichosporon asahii on arsenic exposure: identification of protein responses to metal-induced oxidative stress in an environmentally-sampled isolate. PloS One, 9(7), e102340.
Israr, M., Sahi, S. V., & Jain, J. (2006). Cadmium accumulation and antioxidant responses in the Sesbania drummondii callus. Archives of Environmental Contamination and Toxicology, 50, 121–127.
Kim, H. S., Kwack, S. J., & Lee, B. M. (2005). Alteration of cytochrome P-450 and glutathione S-transferase activity in normal and malignant human stomach. Journal of Toxicology and Environmental Health, 68, 1611–1620.
Kothavade, R. J., Kura, M. M., Valand, A. G., & Panthaki, M. H. (2010). Candida tropicalis: its prevalence, pathogenicity and increasing resistance to fluconazole. Journal of Medical Microbiology, 59(8), 873–880.
Kumagai, H., Tamaki, H., Koshino, Y., Suzuki, H., & Tochikura, T. (1988). Distribution, formation and stabilization of yeast glutathione S-transferase. Agricaltural and Biological Chemistry, 52,1377–1382.
Laemmli, U. K. (1970). Cleavage of structural proteins during assembly of the head of the bacteriophage T4. Nature, 227, 680–685.
Lafaye, A., Junot, C., Pereira, Y., Lagniel, G., Tabet, J. C., Ezan, E., & Labarre, J. (2005). Combined proteome and metabolite-profiling analyses reveal surprising insights into yeast sulfur metabolism. Journal of Biological Chemistry, 280, 24723–24730.
Li, Z. J., & Yuan, H. L. (2006). Characterization of cadmium removal by Rhodotorula sp Y11. Applied Journal of Microbiology and Biotechnology, 73, 458–463.
Li, Z. S., Lu, Y. P., Zhen, R. G., Szczypka, M., Thiele, D. J., & Rea, P. A. (1997). A new pathway for vacuolar cadmium sequestration in Saccharomyces cerevisiae: YCF1-catalyzed transport of bis(glutathionato)cadmium. Proceedings of National Academy of Sciences USA, 94, 42–47.
Masneuf-Pomarède, I., Le Jeune, C., Durrens, P., Lollier, M., Aigle, M., & Dubourdieu, D. (2007). Molecular typing of wine yeast strains Saccharomyces bayanus var. uvarum using microsatellite markers. Systematic and Applied Microbiology, 30(1), 75–82.
Matés, J. M., & Sánchez-Jiménez, F. M. (2000). Role of reactive oxygen species in apoptosis: implications for cancer therapy. The International Journal of Biochemistry & Cell Biology, 32(2), 157–170.
Menezes, R. A., Amaral, C., Batista-Nascimento, L., Santos, C., Ferreira, R. B., Devaux, F., Eleutherio, E. C., & Rodrigues-Pousada, C. (2008). Contribution of Yap1 towards Saccharomyces cerevisiae adaptation to arsenic-mediated oxidative stress. Journal of Biochemistry, 414, 301–311.
Mozafar, A., Ruh, R., Klingel, P., Gamper, H., Egli, S., & Frossard, E. (2002). Effect of heavy metal contaminated shooting range soils on mycorrhizal colonization of roots and metal uptake by leek. Environmental Monitoring and Assessment, 79(2), 177–191.
Pal, A., Choudhuri, P., Dutta, S., Mukherjee, P. K., & Paul, A. K. (2004). Isolation and characterization of nickel-resistant microflora from serpentine soils of Andaman. World Journal of Microbiology and Biotechnology, 20(9), 881–886.
Pena-Llopis, S., Ferrando, M. D., & Pena, J. B. (2002). Impaired glutathione redox status is associated with decreased survival in two organophosphate-poisoned marine bivalves. Chemosphere, 47(5), 485–497.
Rehman, A., & Anjum, M. S. (2011). Multiple metal tolerance and biosorption of cadmium by Candida tropicalis isolated from industrial effluents: glutathione as detoxifying agent. Environmental Monitoring and Assessment, 174, 585–595.
Penninckx, M. J., (2002). An overview on glutathione in Saccharomyces versus non-conventional yeasts. FEMS Yeast Research, 2(3), 295–305
Rehman, A., Farooq, H., & Shakoori, A. R. (2007). Copper tolerant yeast, Candida tropicalis, isolated from industrial effluents: its potential use in wastewater treatment. Pakistan Journal of Zoology, 39(6), 405.
Schafer, F. Q., & Buettner, G. R. (2001). Redox environment of the cell as viewed through the redox state of the glutathione disulfide/glutathione couple. Free Radical Biology and Medicine, 30, 1191–1212.
Sheehan, D., & Casey, J. P. (1993). Evidence for Alpha and Mu class glutathione S-transferases in a number of fungal species. Comparative Biochemistry and Physiology Part B: Comparative Biochemistry, 104(1), 7–13.
Sies, H. (1999). Glutathione and its role in cellular functions. Free Radical Biology and Medicine, 27, 916–921.
Suzuki, Y., Ono, Y., & Hirabayashi, Y. (1998). Rapid and specific reactive oxygen species generation via NADPH oxidase activation during Fas-mediated apoptosis. FEBS Letters, 425(2), 209–212.
Thorsen, M., Lagniel, G., Kristiansson, E., Junot, C., Nerman, O., Labarre, J., & Tamas, M. J. (2007). Quantitative transcriptome, proteome and sulfur metabolite profiling of the Saccharomyces cerevisiae response to arsenite. Physiological Genomics, 30, 35–43.
Vido, K., Spector, D., Lagniel, G., Lopez, S., Toledano, M. B., & Labarre, J. (2001). A proteome analysis of the cadmium response in Saccharomyces cerevisiae. Journal of Biological Chemistry, 276, 8469–8474.
Conflict of interest
We declare that we have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ilyas, S., Rehman, A. Oxidative stress, glutathione level and antioxidant response to heavy metals in multi-resistant pathogen, Candida tropicalis . Environ Monit Assess 187, 4115 (2015). https://doi.org/10.1007/s10661-014-4115-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10661-014-4115-9