Abstract
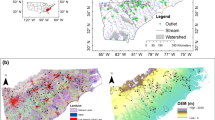
Statistical models of microbial water quality inform risk management for water recreation. Current research focuses on resource-intensive, location-specific data collection and water quality modeling, but this approach may be cost-prohibitive for risk managers responsible for numerous recreation sites. As an alternative, we tested the ability of two data-driven models, tree regression and random forests with conditional inference trees, to select readily available hydrometeorological variables for use in linear mixed effects (LME) models predicting bacterial density. The study included the Chicago Area Waterway System (CAWS) and Lake Michigan beaches and harbors in Chicago, Illinois, at which Escherichia coli and enterococci were measured seasonally in 2007–2009. Tree regression node variables reduced data dimensionality by >50 %. Variable importance ranks from random forests were used in a forward-step selection based on R 2 and root mean squared prediction error (RMSPE). We found two to three variables explained bacteria densities well relative to random forests with all variables. LME models with tree- or forest-selected variables performed reasonably well (0.335 < R 2 < 0.658). LME models for Lake Michigan had good prediction accuracy with respect to the single sample maximum standard (72–77 %), but limited sensitivity (23–62 %). Results suggest that our alternative approach is feasible and performs similarly to more resource-intensive approaches.

Similar content being viewed by others
References
Auret, L., & Aldrich, C. (2011). Empirical comparison of tree ensemble variable importance measures. Chemometrics and Intelligent Laboratory Systems, 105, 157–170.
Boehm, A. B., Whitman, R. L., Nevers, M. B., Hou, D., & Weisberg, S. B. (2007). Nowcasting recreational water quality. In L. J. Wymer (Ed.), Statistical framework for recreational water quality criteria and monitoring (pp. 179–210). Wiley: New York.
Breiman, L. (2001a). Statistical modeling, The two cultures. Statistical Science, 16, 199–231.
Breiman, L. (2001b). Random forests. Machine Learning, 45, 5–32.
Diaz-Uriarte, R., & Alvarez de Andres, S. (2006). Gene selection and classification of microarray data using random forest. BMC Bioinformatics, 7, 3. doi:10.1186/1471-2105-73-3.
Dorevitch, S., Pratap, P., Wroblewski, M., Hryhorczuk, D. O., Li, H., Liu, L. C., et al. (2012). Health risks of limited-contact water recreation. Environmental Health Perspectives, 120, 192. doi:10.1289/ehp.1103934.
Dunkerley, D. (2008). Identifying individual rain events from pluviograph records: a review with analysis from an Australian dryland site. Hydrologic Processes, 22, 5024–5036.
Edwards, P. J., Headley, A. S., Machin, F. H., & Scarr, A. M. (2003). Factors affecting microbiological water quality at sixteen beaches in South-West Wales. Journal of CIWEM, 17, 45–50.
Eleria, A., & Vogel, R. M. (2005). Predicting fecal coliform bacterial levels in the Charles River, Massachusetts, USA. Journal of the American Water Resources Association, 41, 1195–1209.
Frick, W. E., Ge, Z., & Zepp, R. G. (2008). Nowcasting and forecasting concentrations of biological contaminants at beaches: a feasibility and case study. Environmental Science & Technology, 42, 4818–4824.
He, Y., Wang, J., Lek-Ang, S., & Lek, S. (2010). Predicting assemblages and species richness of endemic fish in the upper Yangtze River. Science of the Total Environment, 408, 4211–4220.
Hou, D., Ravinovici, S. J. M., & Boehm, A. B. (2006). Enterococci predictions from partial least squares regression models in conjunction with a single-sample standard improve the efficacy of beach management advisories. Environmental Science & Technology, 40, 1737–1743.
Jiang, H., Deng, Y., Chen, H. S., Tao, L., Sha, Q., Chen, J., et al. (2004). Joint analysis of two microarray gene-expression data sets to select lung adenocarcinoma marker genes. BMC Bioinformatics, 5, 81. doi:10.1186/1471-2105-5-81.
Kampichler, C., Wieland, R., Calme, S., Weissenberger, H., & Arriaga-Weiss, S. (2010). Classification in conservation biology: a comparison of five machine-learning methods. Ecological Informatics, 5, 441–450.
Liaw, A., & Wiener, M. (2002). Classification and regression by random forest. R News, 2(3), 18–22.
Maimone, M., Crockett, C. S., & Cesanek, W. E. (2007). PhillyRiverCast: a real-time bacteria forecasting model and web application for the Schuylkill River. Journal of Water Resources, Planning & Management, 133, 542–549.
Nevers, M. B., & Whitman, R. L. (2005). Nowcast modeling of Escherichia coli concentrations at multiple urban beaches of southern Lake Michigan. Water Research, 39, 5250–5260.
Nevers, M. B., & Whitman, R. L. (2008). Coastal strategies to predict Escherichia coli concentrations for beaches along a 35 km stretch of southern Lake Michigan. Environmental Science & Technology, 42, 4454–4460.
Noble, R. T., Lee, I. M., & Schiff, K. C. (2004). Inactivation of indicator micro-organisms from various sources of faecal contamination in seawater and freshwater. Journal of Applied Microbiology., 96, 464–472.
Olyphant, G. A., & Whitman, R. L. (2004). Elements of a predictive model for determining beach closures on a real time basis: the case of 63rd Street beach Chicago. Environmental Monitoring & Assessment, 98, 175–190.
Parkhurst, D. F., Brenner, K. P., Dufour, A. P., & Wymer, L. J. (2005). Indicator bacteria at five swimming beaches—Analysis using random forests. Water Research, 39, 1354–1360.
Prasad, A. M., Iverson, L. R., & Liaw, A. (2006). Newer classification and regression tree techniques: bagging and random forests for ecological prediction. Ecosystems, 9, 181–199.
Rijal, G., Petropoulou, C., Tolson, J. K., DeFlaun, M., Gerba, C., Gore, R., et al. (2009). Dry and wet weather microbial characterization of the Chicago Area Waterway System. Water Science & Technology, 60, 1847–1855.
Roser, D. J., Davies, C. M., Ashbolt, N. J., & Morison, P. (2006). Microbial exposure assessment of an urban recreational lake: a case study of the application of new risk-based guidelines. Water Science & Technology, 54, 245–252.
Schets, F. M., vanWijnen, J. H., Schijven, J. F., Schoon, H., & de RodaHusman, A. M. (2008). Monitoring of waterborne pathogens in surface waters in Amsterdam, the Netherlands, and the potential health risk associated with exposure to Cryptosporidium and Giardia in these waters. Applied Environmental Microbiology, 74, 2069–2078.
Sinton, L. W., Hall, C. H., Lynch, P. A., & Davies-Colley, R. J. (2002). Sunlight inactivation of fecal indicator bacteria and bacteriophages from waste stabilization pond effluent in fresh and saline waters. Applied Environmental Microbiology, 68, 1122–1131.
Smith, A., Sterba-Boatwright, B., & Mott, J. (2010). Novel application of a statistical technique, Random Forests, in a bacterial source tracking study. Water Research, 44, 4067–4076.
Strobl, C., Boulesteix, A. L., Zeileis, A., & Hothorn, T. (2007). Bias in random forest variable importance measures: illustrations, sources and a solution. BMC Bionformatics, 8, 25. doi:10.1186/1471/2105-8-25.
Strobl, C., Boulesteix, A. L., Kneib, T., Hothorn, T., & Zeileis, A. (2008). Conditional variable importance for random forests. BMC Bioinformatics, 9, 307. doi:10.1186/1471-2105-9-307.
Strobl, C, Hothorn, T., & Zeileis, A. (2009) Party on! A new, conditional variable importance measure for random forests available in the party package. Technical Report Number 050, Department of Statistics, University of Munich.
Svetnik, V., Liaw, A., Tong, C., & Wang, T. (2004). Using Breiman’s random forest to modeling structure–activity relationships of pharmaceutical molecules. Multiple classifier systems, Fifth international workshop, MCS2004, proceedings, 9–11 June, 2004, Caligari, Italy. Lecture notes in computer science, Springer. 3007, 334-343.
Telech, J. W., Brenner, K. P., Haughland, R., Sams, E., Dufour, A. P., Wymer, L., et al. (2009). Modeling enterococcus densities measured by quantitative polymerase chain reaction and membrane filtration using environmental conditions at four Great Lakes beaches. Water Research, 43, 4947–4955.
US EPA. (1986). Ambient water quality criteria for beaches—1986. EPA 440/5-84-002, http://water.epa.gov/scitech/swguidance/standards/criteria/ health/recreation/ upload/2009_04_13_beaches_1986crit.pdf. Accessed on April 12, 2011.
Wie, C. L., Rowe, G. T., Escobar-Briones, E., Boetius, A., Soltwedel, T., Caley, et al. (2010). Global patterns and predictions of seafloor biomass using random forests. PLoS ONE, 5, e15323. doi:10.1371/journal.pone.0015323.
Wilkes, G., Edge, T., Gannon, V., Jokinen, C., Lyautey, E., Medeiros, D., et al. (2009). Seasonal relationships among indicator bacteria, pathogenic bacteria, Cryptosporidium oocysts, Giardia cysts, and hydrological indices for surface waters within an agricultural landscape. Water Research, 43, 2209–2223.
Wong, M., Kumar, L., Jenkins, T. M., Xagoraraki, I., Phanikumar, M. S., & Rose, J. B. (2009). Evaluation of public health risks at recreational beaches in Lake Michigan via detection of enteric viruses and a human-specific bacteriological marker. Water Research, 43, 1137–1149.
Acknowledgments
We would like to acknowledge the contributions of the CHEERS sample collection and data management team, particularly, Mr. Ross Gladding, Dr. Margit Javor, Ms. Chiping Nieh, Dr. Peter Scheff, and Ms. Ember Vannoy. The map was created by Mr. Raja Kaliappan. The CHEERS study was funded by the Metropolitan Water Reclamation District of Greater Chicago.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 6057 kb)
Rights and permissions
About this article
Cite this article
Jones, R.M., Liu, L. & Dorevitch, S. Hydrometeorological variables predict fecal indicator bacteria densities in freshwater: data-driven methods for variable selection. Environ Monit Assess 185, 2355–2366 (2013). https://doi.org/10.1007/s10661-012-2716-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10661-012-2716-8