Abstract
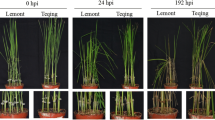
Sclerotinia disease, caused by Sclerotinia sclerotiorum, is one of the most serious plant diseases in China. Research on the mechanism of disease resistance to S. sclerotiorum will help solve control problems. In this study, near-isogenic lines were first used in combination with the proteomic technique. A comparison of protein expression profiles in a susceptible line with those in a resistant line during the interaction of adult Brassica napus with S. sclerotiorum resulted in the identification of 20 important proteins related to disease resistance. Those proteins were then determined to be involved in various functions, including pathogen resistance, antioxidation, and transcription regulation. Our finding showed that some proteins involved in defence—a glycine rich protein (GRP); a trypsin inhibitor protein (TIP); two heat shock proteins (HSPs); and a thiol methyltransferase (TMT)—were upregulated or expressed specially in the resistant B. napus lines. These proteins contribute to ROS (reactive oxygen species) elimination and pathogen-defence in the resistant line, which would help the host defend itself against S. sclerotiorum. As a consequence, the onset of PCD (programmed cell death) was delayed, and the spread of S. sclerotiorum was slowed in the resistant line. Presented results underline the role of specific proteins in the disease process. By building on these results, future research may help determine the genes that are important in conveying resistance to S. sclerotiorum in B. napus.






Similar content being viewed by others
References
Attieh, J., Djiana, R., Koonjul, P., Etienne, C., Sparace, S. A., & Saini, H. S. (2002). Cloning and functional expression of two plant thiol methyltransferases: a new class of enzymes involved in the biosynthesis of sulfur volatiles. Plant Molecular Biology, 50(3), 511–521.
Attieh, J. M., Hanson, A. D., & Saini, H. S. (1995). Purification and characterization of a novel methyltransferase responsible for biosynthesis of halomethanes and methanethiol in Brassica oleracea. Journal of Biological Chemistry, 270(16), 9250–9257.
Banzet, N., Richaud, C., Deveaux, Y., Kazmaier, M., Gagnon, J., & Triantaphylides, C. (1998). Accumulation of small heat shock proteins, including mitochondrial HSP22, induced by oxidative stress and adaptive response in tomato cells. Plant Journal, 13(4), 519–527.
Barna, B., Fodor, J., Harrach, B. D., Pogany, M., & Kiraly, Z. (2012). The Janus face of reactive oxygen species in resistance and susceptibility of plants to necrotrophic and biotrophic pathogens. Plant Physiology and Biochemistry, 59, 37–43. doi:10.1016/j.plaphy.2012.01.014.
Bateman, D. F., & Beer, S. V. (1965). Simultaneous production and synergistic action of oxalic acid and polygalacturonase during pathogenesis by Sclerotium rolfsii. Phytopathology, 55, 204–211.
Bradford, M. M. (1976). A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Analytical Biochemistry, 72, 248–254.
Bressano, M., Curetti, M., Giachero, L., Gil, S. V., Cabello, M., March, G., et al. (2010). Mycorrhizal fungi symbiosis as a strategy against oxidative stress in soybean plants. Journal of Plant Physiology, 167(18), 1622–1626. doi:10.1016/j.jplph.2010.06.024.
Coll, N. S., Epple, P., & Dangl, J. L. (2011). Programmed cell death in the plant immune system. Cell Death and Differentiation, 18(8), 1247–1256. doi:10.1038/cdd.2011.37.
Fulda, S., Mikkat, S., Stegmann, H., & Horn, R. (2011). Physiology and proteomics of drought stress acclimation in sunflower (Helianthus annuus L.). Plant Biology (Stuttgart, Germany), 13(4), 632–642. doi:10.1111/j.1438-8677.2010.00426.x.
Gill, S. S., & Tuteja, N. (2010). Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiology and Biochemistry, 48(12), 909–930. doi:10.1016/j.plaphy.2010.08.016.
Gonzalez-Fernandez, R., Prats, E., & Jorrin-Novo, J. V. (2010). Proteomics of plant pathogenic fungi. Journal of Biomedicine and Biotechnology, 2010, 932527. doi:10.1155/2010/932527.
Gonzalez, I., Romero, J., Rodriguez, B. L., Perez-Castro, R., & Rojas, A. (2012). The immunobiology of the receptor of advanced glycation end-products: trends and challenges. Immunobiology. doi:10.1016/j.imbio.2012.09.005.
Gorg, A., Drews, O., Luck, C., Weiland, F., & Weiss, W. (2009). 2-DE with IPGs. Electrophoresis, 30(Suppl 1), S122–S132. doi:10.1002/elps.200900051.
Gorovits, R., Akad, F., Beery, H., Vidavsky, F., Mahadav, A., & Czosnek, H. (2007). Expression of stress-response proteins upon whitefly-mediated inoculation of tomato yellow leaf curlvirus in susceptible and resistant tomato plants. Molecular Plant-Microbe Interactions, 20(11), 1376–1383. doi:10.1094/MPMI-20-11-1376.
Guan, C. Y., Li, F. Q., Li, X., Chen, S. Y., Wang, G. H., & Liu, Z. S. (2003). Resistance of the double-low rapeseed cultivar Xiangyou 15 (B. napus) to Sclerotinia Sclerotiorum. Acta Agronomica Sinica, 29(5), 715–718.
Guo, X., & Stotz, H. U. (2010). ABA signaling inhibits oxalate-induced production of reactive oxygen species and protects against Sclerotinia sclerotiorum in Arabidopsis thaliana. European Journal of Plant Pathology, 128, 7–19.
Haruta, M., Major, I. T., Christopher, M. E., Patton, J. J., & Constabel, C. P. (2001). A Kunitz trypsin inhibitor gene family from trembling aspen (Populus tremuloides Michx.): cloning, functional expression, and induction by wounding and herbivory. Plant Molecular Biology, 46(3), 347–359.
Hegedus, D. D., Li, R., Buchwaldt, L., Parkin, I., Whitwill, S., Coutu, C., et al. (2008). Brassica napus possesses an expanded set of polygalacturonase inhibitor protein genes that are differentially regulated in response to Sclerotinia sclerotiorum infection, wounding and defense hormone treatment. Planta, 228(2), 241–253. doi:10.1007/s00425-008-0733-1.
Heller, J., & Tudzynski, P. (2011). Reactive oxygen species in phytopathogenic fungi: signaling, development, and disease. Annual Review of Phytopathology, 49, 369–390. doi:10.1146/annurev-phyto-072910-095355.
Himanen, S. J., Nissinen, A., Auriola, S., Poppy, G. M., Stewart, C. N., Jr., Holopainen, J. K., et al. (2008). Constitutive and herbivore-inducible glucosinolate concentrations in oilseed rape (Brassica napus) leaves are not affected by Bt Cry1Ac insertion but change under elevated atmospheric CO2 and O3. Planta, 227(2), 427–437. doi:10.1007/s00425-007-0629-5.
Hu, X., Bidney, D. L., Yalpani, N., Duvick, J. P., Crasta, O., Folkerts, O., et al. (2003). Overexpression of a gene encoding hydrogen peroxide-generating oxalate oxidase evokes defense responses in sunflower. Plant Physiology, 133(1), 170–181.
Huang, H., Qi, S. D., Qi, F., Wu, C. A., Yang, G. D., & Zheng, C. C. (2010). NtKTI1, a Kunitz trypsin inhibitor with antifungal activity from Nicotiana tabacum, plays an important role in tobacco’s defense response. FEBS Journal, 277(19), 4076–4088. doi:10.1111/j.1742-4658.2010.07803.x.
Huang, J., Ge, X., & Sun, M. (2000). Modified CTAB protocol using a silica matrix for isolation of plant genomic DNA. Biotechniques, 28(3), 432–434.
Huerta-Ocampo, J. A., Leon-Galvan, M. F., Ortega-Cruz, L. B., Barrera-Pacheco, A., De Leon-Rodriguez, A., Mendoza-Hernandez, G., et al. (2011). Water stress induces up-regulation of DOF1 and MIF1 transcription factors and down-regulation of proteins involved in secondary metabolism in amaranth roots (Amaranthus hypochondriacus L.). Plant Biology (Stuttgart, Germany), 13(3), 472–482. doi:10.1111/j.1438-8677.2010.00391.x.
Itoh, N., Toda, H., Matsuda, M., Negishi, T., Taniguchi, T., & Ohsawa, N. (2009). Involvement of S-adenosylmethionine-dependent halide/thiol methyltransferase (HTMT) in methyl halide emissions from agricultural plants: isolation and characterization of an HTMT-coding gene from Raphanus sativus (daikon radish). BMC Plant Biology, 9, 116. doi:10.1186/1471-2229-9-116.
Jofre, A., Molinas, M., & Pla, M. (2003). A 10-kDa class-CI sHsp protects E. coli from oxidative and high-temperature stress. Planta, 217(5), 813–819. doi:10.1007/s00425-003-1048-x.
Jones, J. D., & Dangl, J. L. (2006). The plant immune system. Nature, 444(7117), 323–329. doi:10.1038/nature05286.
Karp, N. A., Spencer, M., Lindsay, H., O’Dell, K., & Lilley, K. S. (2005). Impact of replicate types on proteomic expression analysis. Journal of Proteome Research, 4(5), 1867–1871. doi:10.1021/pr050084g.
Kim, J. S., Kim, Y. O., Ryu, H. J., Kwak, Y. S., Lee, J. Y., & Kang, H. (2003). Isolation of stress-related genes of rubber particles and latex in fig tree (Ficus carica) and their expressions by abiotic stress or plant hormone treatments. Plant and Cell Physiology, 44(4), 412–414.
Kim, J. Y., Park, S. J., Jang, B., Jung, C. H., Ahn, S. J., Goh, C. H., et al. (2007). Functional characterization of a glycine-rich RNA-binding protein 2 in Arabidopsis thaliana under abiotic stress conditions. Plant Journal, 50(3), 439–451. doi:10.1111/j.1365-313X.2007.03057.x.
Kim, K. S., Min, J. Y., & Dickman, M. B. (2008). Oxalic acid is an elicitor of plant programmed cell death during Sclerotinia sclerotiorum disease development. Molecular Plant-Microbe Interactions, 21(5), 605–612. doi:10.1094/MPMI-21-5-0605.
Klopfleisch, R., & Gruber, A. D. (2012). Transcriptome and proteome research in veterinary science: what is possible and what questions can be asked? ScientificWorld Journal, 2012, 254962. doi:10.1100/2012/254962.
Kuzniak, E., & Sklodowska, M. (2005). Fungal pathogen-induced changes in the antioxidant systems of leaf peroxisomes from infected tomato plants. Planta, 222(1), 192–200. doi:10.1007/s00425-005-1514-8.
Lee, K., Lee, J., Kim, Y., Bae, D., Kang, K. Y., Yoon, S. C., et al. (2004). Defining the plant disulfide proteome. Electrophoresis, 25(3), 532–541. doi:10.1002/elps.200305677.
Li, R., Rimmer, R., Buchwaldt, L., Sharpe, A. G., Seguin-Swartz, G., & Hegedus, D. D. (2004). Interaction of Sclerotinia sclerotiorum with Brassica napus: cloning and characterization of endo- and exo-polygalacturonases expressed during saprophytic and parasitic modes. Fungal Genetics and Biology, 41(8), 754–765. doi:10.1016/j.fgb.2004.03.002.
Liang, Y., Srivastava, S., Rahman, M. H., Strelkov, S. E., & Kav, N. N. (2008). Proteome changes in leaves of Brassica napus L. as a result of Sclerotinia sclerotiorum challenge. Journal of Agricultural and Food Chemistry, 56(6), 1963–1976. doi:10.1021/jf073012d.
Liang, Y., Strelkov, S. E., & Kav, N. N. (2009). Oxalic acid-mediated stress responses in Brassica napus L. Proteomics, 9(11), 3156–3173. doi:10.1002/pmic.200800966.
Livak, K. J., & Schmittgen, T. D. (2001). Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods, 25(4), 402–408. doi:10.1006/meth.2001.1262.
Lorang, J., Kidarsa, T., Bradford, C. S., Gilbert, B., Curtis, M., Tzeng, S. C., et al. (2012). Tricking the guard: exploiting plant defense for disease susceptibility. Science, 338(6107), 659–662. doi:10.1126/science.1226743.
Luo, M., Ding, L. W., Ge, Z. J., Wang, Z. Y., Hu, B. L., Yang, X. B., et al. (2012). The characterization of SaPIN2b, a plant trichome-localized proteinase inhibitor from Solanum americanum. International Journal of Molecular Sciences, 13(11), 15162–15176. doi:10.3390/ijms131115162.
Mengiste, T. (2012). Plant immunity to necrotrophs. Annual Review of Phytopathology, 50, 267–294. doi:10.1146/annurev-phyto-081211-172955.
Mishra, M., Mahajan, N., Tamhane, V. A., Kulkarni, M. J., Baldwin, I. T., Gupta, V. S., et al. (2012). Stress inducible proteinase inhibitor diversity in Capsicum annuum. BMC Plant Biology, 12, 217. doi:10.1186/1471-2229-12-217.
Mukhtar, M. S., Carvunis, A. R., Dreze, M., Epple, P., Steinbrenner, J., Moore, J., et al. (2011). Independently evolved virulence effectors converge onto hubs in a plant immune system network. Science, 333(6042), 596–601. doi:10.1126/science.1203659.
Park, S. J., Kwak, K. J., Oh, T. R., Kim, Y. O., & Kang, H. (2009). Cold shock domain proteins affect seed germination and growth of Arabidopsis thaliana under abiotic stress conditions. Plant and Cell Physiology, 50(4), 869–878. doi:10.1093/pcp/pcp037.
Parkhey, S., Naithani, S. C., & Keshavkant, S. (2012). ROS production and lipid catabolism in desiccating Shorea robusta seeds during aging. Plant Physiology and Biochemistry, 57, 261–267. doi:10.1016/j.plaphy.2012.06.008.
Pucciariello, C., Parlanti, S., Banti, V., Novi, G., & Perata, P. (2012). Reactive oxygen species-driven transcription in Arabidopsis under oxygen deprivation. Plant Physiology, 159(1), 184–196. doi:10.1104/pp. 111.191122.
Rahmanpour, S., Backhouse, D., & Nonhebel, H. M. (2010). Reaction of glucosinolate-myrosinase defence system in Brassica plants to pathogenicity factor of Sclerotinia sclerotiorum. European Journal of Plant Pathology, 128(4), 429–433.
Rollins, J. A., & Dickman, M. B. (1998). Increase in endogenous and exogenous cyclic AMP levels inhibits Sclerotial development in Sclerotinia sclerotiorum. Applied and Environmental Microbiology, 64(7), 2539–2544.
Sathiyaraj, G., Lee, O. R., Parvin, S., Khorolragchaa, A., Kim, Y. J., & Yang, D. C. (2011). Transcript profiling of antioxidant genes during biotic and abiotic stresses in Panax ginseng C. A. Meyer. Molecular Biology Reports, 38(4), 2761–2769. doi:10.1007/s11033-010-0421-7.
Sattler, S. E., Gilliland, L. U., Magallanes-Lundback, M., Pollard, M., & DellaPenna, D. (2004). Vitamin E is essential for seed longevity and for preventing lipid peroxidation during germination. Plant Cell, 16(6), 1419–1432. doi:10.1105/tpc.021360.
Schmidt, F., Marnef, A., Cheung, M. K., Wilson, I., Hancock, J., Staiger, D., et al. (2010). A proteomic analysis of oligo(dT)-bound mRNP containing oxidative stress-induced Arabidopsis thaliana RNA-binding proteins ATGRP7 and ATGRP8. Molecular Biology Reports, 37(2), 839–845. doi:10.1007/s11033-009-9636-x.
Sexton, A. C., Whitten, A. R., & Howlett, B. J. (2006). Population structure of Sclerotinia sclerotiorum in an Australian canola field at flowering and stem-infection stages of the disease cycle. Genome, 49(11), 1408–1415. doi:10.1139/g06-101.
Srinivasan, T., Kumar, K. R., & Kirti, P. B. (2009). Constitutive expression of a trypsin protease inhibitor confers multiple stress tolerance in transgenic tobacco. Plant and Cell Physiology, 50(3), 541–553. doi:10.1093/pcp/pcp014.
Stotz, H. U., Sawada, Y., Shimada, Y., Hirai, M. Y., Sasaki, E., Krischke, M., et al. (2011). Role of camalexin, indole glucosinolates, and side chain modification of glucosinolate-derived isothiocyanates in defense of Arabidopsis against Sclerotinia sclerotiorum. Plant Journal, 67(1), 81–93. doi:10.1111/j.1365-313X.2011.04578.x.
Swindell, W. R., Huebner, M., & Weber, A. P. (2007). Transcriptional profiling of Arabidopsis heat shock proteins and transcription factors reveals extensive overlap between heat and non-heat stress response pathways. BMC Genomics, 8, 125. doi:10.1186/1471-2164-8-125.
Turoczy, Z., Kis, P., Torok, K., Cserhati, M., Lendvai, A., Dudits, D., et al. (2011). Overproduction of a rice aldo-keto reductase increases oxidative and heat stress tolerance by malondialdehyde and methylglyoxal detoxification. Plant Molecular Biology, 75(4–5), 399–412. doi:10.1007/s11103-011-9735-7.
Verma, S., & Mishra, S. N. (2005). Putrescine alleviation of growth in salt stressed Brassica juncea by inducing antioxidative defense system. Journal of Plant Physiology, 162(6), 669–677.
Wan, C., Li, S., Wen, L., Kong, J., Wang, K., & Zhu, Y. (2007). Damage of oxidative stress on mitochondria during microspores development in Honglian CMS line of rice. Plant Cell Reports, 26(3), 373–382. doi:10.1007/s00299-006-0234-2.
Wang, C., Zhang, D. W., Wang, Y. C., Zheng, L., & Yang, C. P. (2012). A glycine-rich RNA-binding protein can mediate physiological responses in transgenic plants under salt stress. Molecular Biology Reports, 39(2), 1047–1053. doi:10.1007/s11033-011-0830-2.
Wen, L., Liu, G., Li, S. Q., Wan, C. X., Tao, J., Xu, K. Y., et al. (2007). Proteomic analysis of anthers from Honglian cytoplasmic male sterility line rice and its corresponding maintainer and hybrid. Botanical Studies, 48(3), 293–309.
Zhao, J., Buchwaldt, L., Rimmer, S. R., Sharpe, A., McGregor, L., Bekkaoui, D., et al. (2009). Patterns of differential gene expression in Brassica napus cultivars infected with Sclerotinia sclerotiorum. Molecular Plant Pathology, 10(5), 635–649. doi:10.1111/j.1364-3703.2009.00558.x.
Zhao, J., Wang, J., An, L., Doerge, R. W., Chen, Z. J., Grau, C. R., et al. (2007). Analysis of gene expression profiles in response to Sclerotinia sclerotiorum in Brassica napus. Planta, 227(1), 13–24. doi:10.1007/s00425-007-0586-z.
Acknowledgments
We would like to thank Dr. Li Shuimin for providing technical assistance with the MALDI–TOF/TOF MS analysis. We are grateful to Dr. Liu Yisong for assisting us with the bioinformatics analysis. This work was supported by the China Postdoctoral Science Foundation (Grant No. 20110490147) and the Special Financial Grant from China Postdoctoral Science Foundation (Grant No. 2012 T50693). This work was also supported by the Foundation of Science and Technology Project of Hunan Province (Grant No. 2011RS4007) and Project supported by Hunan Provincial Natural Science Foundation of China (13JJ2027).
Author information
Authors and Affiliations
Corresponding authors
Additional information
The two institutions contribute the same.
Rights and permissions
About this article
Cite this article
Wen, L., Tan, TL., Shu, JB. et al. Using proteomic analysis to find the proteins involved in resistance against Sclerotinia sclerotiorum in adult Brassica napus . Eur J Plant Pathol 137, 505–523 (2013). https://doi.org/10.1007/s10658-013-0262-z
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-013-0262-z