Abstract
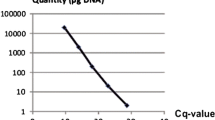
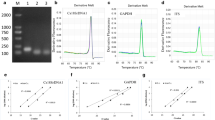
Apple scab, the most important disease of apple worldwide, is caused by Venturia inaequalis. Currently, evaluation of fungal pathogenicity and host resistance is based on a symptomatic disease rating. However, this method does not provide an accurate measurement of the degree of infection and cannot detect early fungal development in symptomless leaves. In this study, a Venturia-specific real-time PCR assay was developed using primers designed around the specific internal transcribed spacer 2 (ITS2) region of the 5.8S rRNA gene. Using SYBR® Green I technology, the assay can accurately quantify Venturia DNA over a concentration range of at least five orders of magnitude. Detection sensitivities were in the order of 100 fg. The method was used to quantify Venturia genomic DNA levels in leaves of three apple cultivars with different levels and types of scab resistance and artificially infected with V. inaequalis. The assay clearly discriminated between Venturia levels in monogenic resistant (‘Topaz’), polygenic resistant (‘Discovery’), and susceptible (‘Golden Delicious’) cultivars, and proved especially useful to quantify pathogen levels during the initial latent stage of infection. The real-time PCR data of ‘Golden Delicious’ were consistent with the observed evolution of the degree of sporulation during a time-course experiment. Although measurements were influenced by the presence of co-extracted PCR-inhibitors, the impact of these compounds was independent of the apple cultivar or the initial amount of fungal DNA present. In conclusion, real-time PCR amplification of the ITS2-5.8S rDNA of Venturia spp. is a faster, more objective and more sensitive method to monitor fungal growth and to evaluate host resistance than phenotypic disease rating scores.





Similar content being viewed by others
References
Bénaouf, G., & Parisi, L. (1998). Characterization of Venturia inaequalis pathogenicity on leaf discs of apple trees. European Journal of Plant Pathology, 104, 785–793.
Borneman, J., & Hartin, R. J. (2000). PCR primers that amplify fungal rRNA genes from environmental samples. Applied and Environmental Microbiology, 66, 4356–4360.
Bowen, J. A., Mesarich, C. H., Bus, V. G. M., Beresford, R. M., Plummer, K. M., & Templeton, M. D. (2011). Venturia inaequalis: the causal agent of apple scab. Molecular Plant Pathology, 12, 105–122.
Brouwer, M., Lievens, B., Van Hemelrijck, W., Van den Ackerveken, G., Cammue, B. P. A., & Thomma, B. P. H. J. (2003). Quantification of disease progression of several microbial pathogens on Arabidopsis thaliana using real-time fluorescence PCR. FEMS Microbiology Letters, 228, 241–248.
Bus, V., Rikkerink, E., Aldwinckle, H. S., Caffier, V., Durel, C. E., Gardiner, S., et al. (2009). A proposal for the nomenclature of Venturia inaequalis races. Acta Horticulturae, 814, 739–746.
Chevalier, M., Lespinasse, Y., & Renaudin, S. (1991). A microscopic study of the different classes of symptoms coded by the Vf gene in apple for resistance to scab (Venturia inaequalis). Plant Pathology, 40, 249–256.
Croxall, H. E., Gwynne, D. C., & Jenkins, J. E. E. (1952). The rapid assessment of apple scab on leaves. Plant Pathology, 1, 39–41.
Debode, J., Van Hemelrijck, W., Baeyen, S., Creemers, P., Heungens, K., & Maes, M. (2009). Quantitative detection and monitoring of Colletotrichum acutatum in strawberry leaves using real-time PCR. Plant Pathology, 58, 504–514.
Develey-Rivière, M. P., & Galiana, E. (2007). Resistance to pathogens and host developmental stage: a multifaceted relationship within the plant kingdom. New Phytologist, 175, 405–416.
Gessler, C., Patocchi, A., Sansavini, S., Tartarini, S., & Gianfranceschi, L. (2006). Venturia inaequalis resistance in apple. Critical Reviews in Plant Sciences, 25, 473–503.
Gu, G., Hu, J., Cevallos-Cevallos, J. M., Richardson, S. M., Bartz, J. A., & van Bruggen, A. H. C. (2011). Internal colonization of Salmonella enterica serovar Typhimurium in tomato plants. PLoS One. doi:10.1371/journal.pone.0027340.
Heid, C. A., Stevens, J., Livak, K. J., & Williams, P. M. (1996). Real time quantitative PCR. Genome Research, 6, 986–994.
Hrazdina, G. (2003). Response of scab-susceptible (McIntosh) and scab-resistant (Liberty) apple tissues to treatment with yeast extract and Venturia inaequalis. Phytochemistry, 64, 485–492.
Hu, X., Nazar, R. N., & Robb, J. (1993). Quantification of Verticillium biomass in wilt disease development. Physiological and Molecular Plant Pathology, 42, 23–36.
Josefsen, L., Droce, A., Sondergaard, T. E., Sorensen, J. L., Bormann, J., Schafer, W., et al. (2012). Autophagy provides nutrients for nonassimilating fungal structures and is necessary for plant colonization but not for infection in the necrotrophic plant pathogen Fusarium graminearum. Autophagy, 8, 326–337.
Köller, W., Parker, D. M., Turechek, W. W., Avila-Adame, C., & Cronshaw, K. (2004). A two-phase resistance response of Venturia inaequalis populations to the QoI fungicides kresoxim-methyl and trifloxystrobin. Plant Disease, 88, 537–544.
Korsman, J., Meisel, B., Kloppers, F. J., Crampton, B. G., & Berger, D. K. (2012). Quantitative phenotyping of grey leaf spot disease in maize using real-time PCR. European Journal of Plant Pathology, 133, 461–471.
Lamar, R. T., Schoenike, B., Vanden Wymelenberg, A., Stewart, P., Dietrich, D. M., & Cullen, D. (1995). Quantitation of fungal mRNAs in complex substrates by reverse transcription PCR and its application to Phanerochaete chrysosporium-colonized soil. Applied and Environmental Microbiology, 61, 2122–2126.
Lee, S. B., & Taylor, J. W. (1992). Phylogeny of five fungus-like protoctistan Phytophthora species, inferred from the internal transcribed spacers of ribosomal DNA. Molecular Biology and Evolution, 9, 636–653.
Lespinasse, Y., Pinet, C., & Parisi, L. (2002). European research for durable resistance to scab on apple: the DARE project. Acta Horticulturae, 595, 17–22.
Lievens, B., Brouwer, M., Vanachter, A. C. R. C., Cammue, B. P. A., & Thomma, B. P. H. J. (2006). Real-time PCR for detection and quantification of fungal and oomycete tomato pathogens in plant and soil samples. Plant Science, 171, 155–165.
MacHardy, W. E. (1996). Apple Scab: biology, epidemiology and management. Saint Paul: The American Phytopathological Society Press.
Mahe, A., Grisvard, J., & Dron, M. (1992). Fungal-specific and plant-specific gene markers to follow the bean anthracnose infection process and normalize a bean chitinase mRNA induction. Molecular Plant-Microbe Interactions, 5, 242–248.
Morrison, T. B., Weis, J. J., & Wittwer, C. T. (1998). Quantification of low-copy transcripts by continuous SYBR Green I monitoring during amplification. BioTechniques, 24, 954–962.
Ostle, B. (1954). Correlation methods. In B. Ostle (Ed.), Statistics in research (pp. 174–201). Ames: Iowa State College Press.
Parisi, L., Lespinasse, Y., Guillaumes, J., & Kruger, J. (1993). A new race of Venturia inaequalis virulent to apples with resistance due to the Vf gene. Phytopathology, 83, 533–537.
Qi, M., & Yang, Y. (2002). Quantification of Magnaporthe grisea during infection of rice plants using real-time polymerase chain reaction and northern blot/phosphoimaging analyses. Phytopathology, 92, 870–876.
Schena, L., Nigro, F., Ippolito, A., & Gallitelli, D. (2004). Real-time quantitative PCR: a new technology to detect and study phytopathogenic and antagonistic fungi. European Journal of Plant Pathology, 110, 893–908.
Schnabel, G., Schnabel, E. L., & Jones, A. L. (1999). Characterization of ribosomal DNA from Venturia inaequalis and its phylogenetic relationship to rDNA from other tree-fruit Venturia species. Phytopathology, 89, 100–108.
Shaw, D. V., Gordon, T. R., Hansen, J., & Kirkpatrick, S. C. (2010). Relationship between the extent of colonization by Verticillium dahliae and symptom expression in the strawberry (Fragaria × ananassa) genotypes resistant to verticillium wilt. Plant Pathology, 59, 376–381.
Stehmann, C., Pennycook, A., & Plummer, K. M. (2001). Molecular identification of a sexual interloper: the pear pathogen, Venturia pyrina, has sex on apple. Phytopathology, 91, 633–641.
Stephens, A. E., Gardiner, D. M., White, R. G., Munn, A. L., & Manners, J. M. (2008). Phases of infection and gene expression of Fusarium graminearum during crown rot disease of wheat. Molecular Plant-Microbe Interactions, 21, 1571–1581.
Thomma, B. P. H. J., Eggermont, K., Tierens, K. F. M., & Broekaert, W. F. (1999). Requirement of functional EIN2 (ethylene insensitive 2) gene for efficient resistance of Arabidopsis thaliana to infection by Botrytis cinerea. Plant Physiology, 121, 1093–1101.
Towsend, G. R., & Heuberger, J. W. (1943). Methods for estimating losses caused by diseases in fungicide experiments. Plant Disease Reports, 24, 340–343.
Turechek, W. W. (2004). Apple diseases and their management. In S. A. M. H. Naqvi (Ed.), Diseases of fruits and vegetables. Diagnosis and management. Volume I (pp. 1–108). Dordrecht: Kluwer Academic Publishers.
Vandemark, G. J., Barker, B. M., & Gritsenko, M. A. (2002). Quantifying Aphanomyces euteiches in alfalfa with a fluorescent polymerase chain reaction assay. Phytopathology, 92, 265–272.
White, T. J., Bruns, T., Lee, S., & Taylor, J. A. (1990). Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In M. A. Innis, D. H. Gelfand, J. J. Shinsky, & T. J. White (Eds.), PCR Protocols: a guide to methods and applications (pp. 315–322). San Diego: Academic.
Wilson, I. G. (1997). Inhibition and facilitation of nucleic acid amplification. Applied and Environmental Microbiology, 63, 3741–3751.
Zubini, P., Baraldi, E., De Santis, A., Bertolini, P., & Mari, M. (2007). Expression of anti-oxidant enzyme genes in scald-resistant ‘Belfort’ and scald-susceptible ‘Granny Smith’ apples during cold storage. The Journal of Horticultural Science and Biotechnology, 82, 149–155.
Acknowledgments
This research was funded by a PhD grant of the Institute for the Promotion of Innovation through Science and Technology in Flanders (IWT-Vlaanderen). The authors thank Valérie Caffier from INRA (Angers, France) for kindly providing V. inaequalis isolates on PDA. Our gratitude also goes to Jane Debode from the Flemish Institute for Agriculture and Fishery Research (ILVO) in Merelbeke (Belgium), and to Bart Lievens from the Scientia Terrae Research Institute (Sint-Katelijne-Waver, Belgium) for their advice and support on protocols for extraction of gDNA from plant tissue.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Daniëls, B., De landtsheer, A., Dreesen, R. et al. Real-time PCR as a promising tool to monitor growth of Venturia spp. in scab-susceptible and -resistant apple leaves. Eur J Plant Pathol 134, 821–833 (2012). https://doi.org/10.1007/s10658-012-0058-6
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-012-0058-6