Abstract
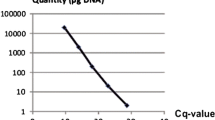
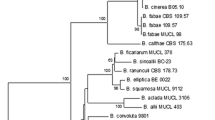
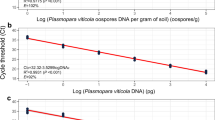
Downy mildew disease caused by Peronospora sparsa, also known as ‘dryberry’ disease, is a serious threat to the cultivation of arctic bramble (Rubus arcticus) and boysenberry (Rubus spp. hybrid). A quantitative and sensitive screening method is necessary for the breeding of downy mildew resistant cultivars and for determining efficient disease control methods. A quantitative real-time PCR method using SYBR® Green I fluorescent dye was developed for the analysis of P. sparsa in arctic bramble, other Rubus species and roses. Primers were designed to amplify a P. sparsa specific 94-bp fragment from the internal transcribed spacer region (ITS1) and a 140-bp fragment from a conserved region of plant 5.8S ribosomal DNA, which served as an internal control in the samples. Linear amplification from genomic DNA and control plasmids was achieved with both primers, and even 37 fg of P. sparsa conidial DNA was detected. In the samples collected from the field, quantities as low as 0.2 ppm of P. sparsa DNA in plant DNA were detected, thus enabling the diagnosis of weak and latent infections. Arctic bramble cvs Pima, Mespi and Mesma, all showing distinct foliar symptoms, were tested to assess the relative amount of downy mildew DNA present. The symptoms and the amount of P. sparsa DNA detected correlated only in cv. Pima, indicating that visual inspection of symptoms is not a reliable method for assessing the extent of tissue infection. The number of conidiophores of P. sparsa on in vitro inoculated leaves of two arctic bramble cultivars correlated with the results obtained by real-time PCR.
Similar content being viewed by others
Abbreviations
- bp:
-
base pair
- Ct:
-
threshold cycle
- ITS:
-
internal transcribed spacer
- rDNA:
-
ribosomal DNA
References
Aegerter BJ, Nuñez JJ, Davis RM (2002) Detection and management of downy mildew in rose rootstock. Plant Disease 86: 1363–1368
Agnola B, Boury S, Monot C, Quillévéré A, Hervé Y, Silué D (2003) Evidence that a leaf-disk test allows assessment of isolate-specific resistance in Brassica oleracea crops against downy mildew (Peronospora parasitica). European Journal of Plant Pathology 109: 471–478
Breese WA, Shattock RC, Williamson B, Hackett C (1994) In vitro spore germination and infection of cultivars of Rubus and Rosa by downy mildews from both hosts. Annals of Applied Biology 125: 73–85
Brown MV, Moore JN, Fenn P, McNew RW (1999) Comparison of leaf disk, greenhouse, and field screening procedures for evaluation of grape seedlings for downy mildew resistance. HortScience 34: 331–333
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12: 13–15
Filion M, St-Arnaud M, Jabaji-Hare SH (2003) Direct quantification of fungal DNA from soil substrate using real-time PCR. Journal of Microbiological Methods 53: 67–76
Gachon C, Mingam A, Charrier B (2004) Real-time PCR: what relevance to plant studies? Journal of Experimental Botany 55: 1445–1454
Gachon C, Saindrenan P (2004) Real-time PCR monitoring of fungal development in Arabidopsis thaliana infected by Alternaria brassicicola and Botrytis cinerea. Plant Physiology and Biochemistry 42: 367–371
Hall G (1989) Peronospora rubi. CMI descriptions of pathogenic fungi and bacteria no. 976. Mycopathologia 106: 195–197
Kokko H, Virtaharju O, Kärenlampi S, Cooke DEL, Williams NA, Williamson B (1999) Downy mildew (Peronospora rubi) in Rubus arcticus: a threat to commercial berry production in Finland. Acta Horticulturae 505: 137–141
Koponen H, Hellqvist S, Lindqvist-Kreuze H, Bång U, Valkonen JPT (2000) Occurrence of Peronospora sparsa (P. rubi) on cultivated and wild Rubus species in Finland and Sweden. Annals of Applied Biology 137: 107–112
Lindqvist H, Koponen H, Valkonen JPT (1998) Peronospora sparsa on cultivated Rubus arcticus and its detection by PCR based on ITS sequences. Plant Disease 82: 1305–1311
Lindqvist-Kreuze H, Koponen H, Valkonen JPT (2002) Variability of Peronospora sparsa (syn. P. rubi) in Finland as measured by amplified fragment length polymorphism. European Journal of Plant Pathology 108: 327–339
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25: 402–408
McKeown B (1988) Downy mildew of boysenberry and tummelberry in the UK. Plant Pathology 37: 281–284
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue culture. Physiologia Plantarum 15: 473–497
O’Neill TM, Pye D, Locke T (2002) The effect of fungicides, irrigation and plant density on the development of Peronospora sparsa, the cause of downy mildew in rose and blackberry. Annals of Applied Biology 140: 207–214
Pezet R, Gindro K, Viret O, Spring JL (2004) Glycosylation and oxidative dimerization of resveratrol are respectively associated to sensitivity and resistance of grapevine cultivars to downy mildew. Physiological and Molecular Plant Pathology 65: 297–303
Reuveni M (1998) Relationships between leaf age, peroxidase and beta−1,3-glucanase activity, and resistance to downy mildew in grapevines. Journal of Phytopathology 146: 525–530
Ririe KM, Rasmussen RP, Wittwer CT (1997) Product differentiation by analysis of DNA melting curves during the polymerase chain reaction. Analytical Biochemistry 245: 154–160
Schena L, Nigro F, Ippolito A, Gallitelli D (2004) Real-time quantitative PCR: a new technology to detect and study phytopathogenic and antagonistic fungi. European Journal of Plant Pathology 110: 893–908
Tate KG (1981) Aetiology of dryberry disease of boysenberry in New Zealand. New Zealand Journal of Experimental Agriculture 9: 371–376
Voglmayr H (2003) Phylogenetic relationships of Peronospora and related genera based on nuclear ribosomal ITS sequences. Mycological Research 107: 1132–1142
Walter M, Harris-Virgin P, Thomas W, Tate G, Waipara NW, Langford G (2004) Agrochemicals suitable for downy mildew control in New Zealand boysenberry production. Crop Protection 23: 327–333
Williamson B, Cooke DEL, Duncan JM, Leifert C, Breese WA, Shattock RC (1998) Fungal infections of micropropagated plants at weaning: a problem exemplified by downy mildews in Rubus and Rosa. Plant Cell, Tissue and Organ Culture 52: 89–96
Winton LM, Stone JK, Watrud LS, Hansen EM (2001) Simultaneous one-tube quantification of host and pathogen DNA with real-time polymerase chain reaction. Phytopathology 91: 112–116
Acknowledgements
This research was funded by the Northern Periphery Programme (INTERREG IIIB, Grant no. 102-12874-02), Tiura Foundation, and Kuopio Naturalists’ Society. We also thank Dr A. Stensvand (Norwegian Crop Research Institute), Dr A. Buchala (University of Fribourg), and Prof. J. Hantula (Finnish Forest Research Institute) for providing us with the fungal samples.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hukkanen, A., Pietikäinen, L., Kärenlampi, S. et al. Quantification of downy mildew (Peronospora sparsa) in Rubus species using real-time PCR. Eur J Plant Pathol 116, 225–235 (2006). https://doi.org/10.1007/s10658-006-9054-z
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-006-9054-z