Summary
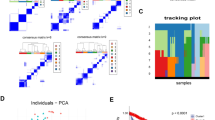
Melanoma is a highly aggressive malignant skin tumor with a high rate of metastasis and mortality. In this study, a comprehensive bioinformatics analysis was used to clarify the hub genes and potential drugs. Download the GSE3189, GSE22301, and GSE35388 microarray datasets from the Gene Expression Omnibus (GEO), which contains a total of 33 normal samples and 67 melanoma samples. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) approach analyze DEGs based on the DAVID. Use STRING to construct protein-protein interaction network, and use MCODE and cytoHubba plug-ins in Cytoscape to perform module analysis and identified hub genes. Use Gene Expression Profile Interactive Analysis (GEPIA) to assess the prognosis of genes in tumors. Finally, use the Drug-Gene Interaction Database (DGIdb) to screen targeted drugs related to hub genes. A total of 140 overlapping DEGs were identified from the three microarray datasets, including 59 up-regulated DEGs and 81 down-regulated DEGs. GO enrichment analysis showed that these DEGs are mainly involved in the biological process such as positive regulation of gene expression, positive regulation of cell proliferation, positive regulation of MAP kinase activity, cell migration, and negative regulation of the apoptotic process. The cellular components are concentrated in the membrane, dendritic spine, the perinuclear region of cytoplasm, extracellular exosome, and membrane raft. Molecular functions include protein homodimerization activity, calmodulin-binding, transcription factor binding, protein binding, and cytoskeletal protein binding. KEGG pathway analysis shows that these DEGs are mainly related to protein digestion and absorption, PPAR signaling pathway, signaling pathways regulating stem cells’ pluripotency, and Retinol metabolism. The 23 most closely related DEGs were identified from the PPI network and combined with the GEPIA prognostic analysis, CDH3, ESRP1, FGF2, GBP2, KCNN4, KIT, SEMA4D, and ZEB1 were selected as hub genes, which are considered to be associated with poor prognosis of melanoma closely related. Besides, ten related drugs that may have therapeutic effects on melanoma were also screened. These newly discovered genes and drugs provide new ideas for further research on melanoma.











Similar content being viewed by others
References
Kandolf Sekulovic L, Peris K, Stratigos A, Hauschild A, Forsea AM, Lebbe C, Lallas A, Grob JJ, Harwood C, Gogas H, Rutkowski P, Olah J, Kelleners-Smeets NWJ, Paoli J, Dummer R, Moreno-Ramirez D, Bastholt L, Putnik K, Karls R, Hoeller C, Vandersleyen V, Vieira R, Arenberger P, Bylaite-Buckinskiene M, Ocvirk J, Situm M, Weinlich G, Banjin M, Todorovic V, Ymeri A, Zhukavets A, Garbe C (2020) Which medical disciplines diagnose and treat melanoma in Europe in 2019? A survey of experts from melanoma centers in 27 European countries. J Eur Acad Dermatol Venereol. https://doi.org/10.1111/jdv.17086
Ferguson NN (2020) Challenges and controversy in determining UV exposure as a risk factor for cutaneous melanoma in skin of color. JAMA Dermatol. https://doi.org/10.1001/jamadermatol.2020.4615
Lopes F, Sleiman MG, Sebastian K, Bogucka R, Jacobs EA, Adamson AS (2020) UV exposure and the risk of cutaneous melanoma in skin of color: a systematic review. JAMA Dermatol. https://doi.org/10.1001/jamadermatol.2020.4616
Jeyakumar A, Chua TC, Lam AK, Gopalan V (2020) The melanoma and breast Cancer association: an overview of their 'Second primary Cancers' and the epidemiological, Genetic and Biological correlations. Crit Rev Oncol Hematol 152:102989
Siegel RL, Miller KD, Jemal A (2020) Cancer statistics, 2020. CA Cancer J Clin 70(1):7–30
Guo J, Qin S, Liang J, Lin T, Si L, Chen X, Chi Z, Cui C, Du N, Fan Y et al (2015) Chinese guidelines on the diagnosis and treatment of melanoma (2015 edition). Ann Transl Med 3(21):322
Balch CM, Gershenwald JE, Soong SJ, Thompson JF, Atkins MB, Byrd DR, Buzaid AC, Cochran AJ, Coit DG, Ding S, Eggermont AM, Flaherty KT, Gimotty PA, Kirkwood JM, McMasters KM, Mihm MC Jr, Morton DL, Ross MI, Sober AJ, Sondak VK (2009) Final version of 2009 AJCC melanoma staging and classification. J Clin Oncol 27(36):6199–6206
Feero WG (2020) Bioinformatics, sequencing accuracy, and the credibility of clinical genomics. JAMA 324(19):1945–1947
Djulbegovic MB, Uversky VN (2020) Expanding the understanding of the heterogeneous nature of melanoma with bioinformatics and disorder-based proteomics. Int J Biol Macromol 150:1281–1293
Barrett T, Troup DB, Wilhite SE, Ledoux P, Rudnev D, Evangelista C, Kim IF, Soboleva A, Tomashevsky M, Edgar R (2007) NCBI GEO: mining tens of millions of expression profiles--database and tools update. Nucleic Acids Res 35:D760–D765
Talantov D, Mazumder A, Yu JX, Briggs T, Jiang Y, Backus J, Atkins D, Wang Y (2005) Novel genes associated with malignant melanoma but not benign melanocytic lesions. Clin Cancer Res 11(20):7234–7242
Rose AE, Poliseno L, Wang J, Clark M, Pearlman A, Wang G, Vega YSEC, Medicherla R, Christos PJ, Shapiro R et al (2011) Integrative genomics identifies molecular alterations that challenge the linear model of melanoma progression. Cancer Res 71(7):2561–2571
Xiao D, Ohlendorf J, Chen Y, Taylor DD, Rai SN, Waigel S, Zacharias W, Hao H, McMasters KM (2012) Identifying mRNA, microRNA and protein profiles of melanoma exosomes. PLoS One 7(10):e46874
Gautier L, Cope L, Bolstad BM, Irizarry RA (2004) Affy--analysis of Affymetrix GeneChip data at the probe level. Bioinformatics 20(3):307–315
Chen B, Ding P, Hua Z, Qin X, Li Z (2020a) Analysis and identification of novel biomarkers involved in neuroblastoma via integrated bioinformatics. Investig New Drugs. https://doi.org/10.1007/s10637-020-00980-9
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK (2015) limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43(7):e47
Chen B, Hua Z, Gong B, Tan X, Zhang S, Li Q, Chen Y, Zhang J, Li Z (2020b) Downregulation of PIF1, a potential new target of MYCN, induces apoptosis and inhibits cell migration in neuroblastoma cells. Life Sci 256:117820
Wang Y, Zhang Y, Huang Q, Li C (2018a) Integrated bioinformatics analysis reveals key candidate genes and pathways in breast cancer. Mol Med Rep 17(6):8091–8100
Niu J, Yan T, Guo W, Wang W, Zhao Z, Ren T, Huang Y, Zhang H, Yu Y, Liang X (2020) Identification of potential therapeutic targets and immune cell infiltration characteristics in osteosarcoma using bioinformatics strategy. Front Oncol 10:1628
Huang da W, Sherman BT, Lempicki RA: Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 2009, 4(1):44–57
Sherman BT, Huang da W, Tan Q, Guo Y, Bour S, Liu D, Stephens R, Baseler MW, Lane HC, Lempicki RA (2007) DAVID Knowledgebase: a gene-centered database integrating heterogeneous gene annotation resources to facilitate high-throughput gene functional analysis. BMC Bioinforma 8:426
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, Hill DP, Issel-Tarver L, Kasarskis A, Lewis S, Matese JC, Richardson JE, Ringwald M, Rubin GM, Sherlock G (2000) Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25(1):25–29
Kanehisa M, Furumichi M, Tanabe M, Sato Y, Morishima K (2017) KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res 45(D1):D353–D361
Szklarczyk D, Gable AL, Lyon D, Junge A, Wyder S, Huerta-Cepas J, Simonovic M, Doncheva NT, Morris JH, Bork P, Jensen LJ, Mering C (2019) STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res 47(D1):D607–D613
Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z (2017) GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res 45(W1):W98–W102
Freshour SL, Kiwala S, Cotto KC, Coffman AC, McMichael, JF, Song JJ, Griffith M, Griffith OL, Wagner AH (2020) Integration of the drug-gene interaction database (DGIdb 4.0) with open crowdsource efforts. Nucleic Acids Res 49(D1):D1144–D1151
Mo R, Chen C, Mi L, Ma Z, Tan Q (2020) Skin melanoma survival is not superior in females in the new stage IIID of the 8th edition of the staging system: an analysis of data from the surveillance, epidemiology, and end results (SEER) database. Ann Transl Med 8(21):1381
Durbec F, Martin L, Derancourt C, Grange F (2012) Melanoma of the hand and foot: epidemiological, prognostic and genetic features. A systematic review. Br J Dermatol 166(4):727–739
Takahashi J, Nagasawa S (2020) Immunostimulatory effects of radiotherapy for local and systemic control of melanoma: a review. Int J Mol Sci 21(23). https://doi.org/10.3390/ijms21239324
Ribas A, Algazi A, Ascierto PA, Butler MO, Chandra S, Gordon M, Hernandez-Aya L, Lawrence D, Lutzky J, Miller WH Jr, Campbell KM, Delafont B, Marshall S, Mueller N, Robert C (2020) PD-L1 blockade in combination with inhibition of MAPK oncogenic signaling in patients with advanced melanoma. Nat Commun 11(1):6262
Wang C, Shi X, Song H, Zhang C, Wang X, Huang P, Dong A, Zhang Y, Kong D, Wang W (2020a) Polymer-lipid hybrid nanovesicle-enabled combination of immunogenic chemotherapy and RNAi-mediated PD-L1 knockdown elicits antitumor immunity against melanoma. Biomaterials 268:120579
Jessen C, Kress JKC, Baluapuri A, Hufnagel A, Schmitz W, Kneitz S, Roth S, Marquardt A, Appenzeller S, Ade CP et al (2020) Correction: the transcription factor NRF2 enhances melanoma malignancy by blocking differentiation and inducing COX2 expression. Oncogene 39(44):6841–6855
Won YS, Kim JH, Lizardo RCM, Min HJ, Cho HD, Hong SM, Seo KI (2020) The flavonol isoquercitrin promotes mitochondrial-dependent apoptosis in SK-Mel-2 melanoma cell via the PI3K/AKT/mTOR pathway. Nutrients 12(12). https://doi.org/10.3390/nu12123683
Wang Z, Xu Q, Zhang N, Du X, Xu G, Yan X (2020b) CD146, from a melanoma cell adhesion molecule to a signaling receptor. Signal Transduct Target Ther 5(1):148
Liu JF, Chen PC, Chang TM, Hou CH (2020a) Monocyte Chemoattractant Protein-1 promotes cancer cell migration via c-Raf/MAPK/AP-1 pathway and MMP-9 production in osteosarcoma. J Exp Clin Cancer Res 39(1):254
Xu M, Wang J, Li H, Zhang Z, Cheng Z (2020) AIM2 inhibits colorectal cancer cell proliferation and migration through suppression of Gli1. Aging (Albany NY) 13(1):1017–1031
Gomes-da-Silva LC, Jimenez AJ, Sauvat A, Xie W, Souquere S, Divoux S, Storch M, Sveinbjornsson B, Rekdal O, Arnaut LG et al (2019) Recruitment of LC3 to damaged Golgi apparatus. Cell Death Differ 26(8):1467–1484
Yang E, Wang X, Gong Z, Yu M, Wu H, Zhang D (2020) Exosome-mediated metabolic reprogramming: the emerging role in tumor microenvironment remodeling and its influence on cancer progression. Signal Transduct Target Ther 5(1):242
Arnold J, Schattschneider J, Blechner C, Krisp C, Schluter H, Schweizer M, Nalaskowski M, Oliveira-Ferrer L, Windhorst S (2020) Tubulin tyrosine ligase like 4 (TTLL4) overexpression in breast cancer cells is associated with brain metastasis and alters exosome biogenesis. J Exp Clin Cancer Res 39(1):205
Meerson NR, Delautier D, Durand-Schneider AM, Moreau A, Schilsky ML, Sternlieb I, Feldmann G, Maurice M (1998) Identification of B10, an alkaline phosphodiesterase of the apical plasma membrane of hepatocytes and biliary cells, in rat serum: increased levels following bile duct ligation and during the development of cholangiocarcinoma. Hepatology 27(2):563–568
Fancy RM, Kim H, Zhou T, Zinn KR, Buchsbaum DJ, Song Y (2017) Calmodulin binding to death receptor 5-mediated death-inducing signaling complex in breast Cancer cells. J Cell Biochem 118(8):2285–2294
Chen Y, Pawar P, Pan G, Ma L, Liu H, McDonald JM (2008) Calmodulin binding to the Fas-mediated death-inducing signaling complex in cholangiocarcinoma cells. J Cell Biochem 103(3):788–799
Karp CM, Shukla MN, Buckley DJ, Buckley AR (2007) HRPAP20: a novel calmodulin-binding protein that increases breast cancer cell invasion. Oncogene 26(12):1780–1788
Mauro JA, Yavorski JM, Blanck G (2017) Stratifying melanoma and breast cancer TCGA datasets on the basis of the CNV of transcription factor binding sites common to proliferation- and apoptosis-effector genes. Gene 614:37–48
Liu X, Qian D, Liu H, Abbruzzese JL, Luo S, Walsh KM, Wei Q (2020b) Genetic variants of the peroxisome proliferator-activated receptor (PPAR) signaling pathway genes and risk of pancreatic cancer. Mol Carcinog 59(8):930–939
Zhang X, Yao J, Shi H, Gao B, Zhang L (2019) LncRNA TINCR/microRNA-107/CD36 regulates cell proliferation and apoptosis in colorectal cancer via PPAR signaling pathway based on bioinformatics analysis. Biol Chem 400(5):663–675
Chen YZ, Xue JY, Chen CM, Yang BL, Xu QH, Wu F, Liu F, Ye X, Meng X, Liu GY, Shen ZZ, Shao ZM, Wu J (2012) PPAR signaling pathway may be an important predictor of breast cancer response to neoadjuvant chemotherapy. Cancer Chemother Pharmacol 70(5):637–644
Huang R, Li Z, Li C, Wang G, Yan P, Peng L, Wang J, Zhu X, Hu P, Zhang J et al (2019) Germ cell-specific gene 1-like protein regulated by splicing factor CUGBP Elav-like family member 5 and primary bile acid biosynthesis are prognostic in glioblastoma Multiforme. Front Genet 10:1380
Jiang Y, Song H, Jiang L, Qiao Y, Yang D, Wang D, Li J (2020) Silybin prevents prostate Cancer by inhibited the ALDH1A1 expression in the retinol metabolism pathway. Front Cell Dev Biol 8:574394
Guo X, Knudsen BS, Peehl DM, Ruiz A, Bok D, Rando RR, Rhim JS, Nanus DM, Gudas LJ (2002) Retinol metabolism and lecithin:retinol acyltransferase levels are reduced in cultured human prostate cancer cells and tissue specimens. Cancer Res 62(6):1654–1661
Chen AC, Guo X, Derguini F, Gudas LJ (1997) Human breast cancer cells and normal mammary epithelial cells: retinol metabolism and growth inhibition by the retinol metabolite 4-oxoretinol. Cancer Res 57(20):4642–4651
Hayden LJ, Satre MA (2002) Alterations in cellular retinol metabolism contribute to differential retinoid responsiveness in normal human mammary epithelial cells versus breast cancer cells. Breast Cancer Res Treat 72(2):95–105
Vieira AF, Dionisio MR, Gomes M, Cameselle-Teijeiro JF, Lacerda M, Amendoeira I, Schmitt F, Paredes J (2017) P-cadherin: a useful biomarker for axillary-based breast cancer decisions in the clinical practice. Mod Pathol 30(5):698–709
Royo F, Zuniga-Garcia P, Torrano V, Loizaga A, Sanchez-Mosquera P, Ugalde-Olano A, Gonzalez E, Cortazar AR, Palomo L, Fernandez-Ruiz S et al (2016) Transcriptomic profiling of urine extracellular vesicles reveals alterations of CDH3 in prostate cancer. Oncotarget 7(6):6835–6846
Liu D, Wu K, Yang Y, Zhu D, Zhang C, Zhao S (2020c) Long noncoding RNA ADAMTS9-AS2 suppresses the progression of esophageal cancer by mediating CDH3 promoter methylation. Mol Carcinog 59(1):32–44
Li L, Yu S, Wu Q, Dou N, Li Y, Gao Y (2019) KLF4-mediated CDH3 Upregulation suppresses human Hepatoma cell growth and migration via GSK-3beta signaling. Int J Biol Sci 15(5):953–961
Taniuchi K, Nakagawa H, Hosokawa M, Nakamura T, Eguchi H, Ohigashi H, Ishikawa O, Katagiri T, Nakamura Y (2005) Overexpressed P-cadherin/CDH3 promotes motility of pancreatic cancer cells by interacting with p120ctn and activating rho-family GTPases. Cancer Res 65(8):3092–3099
Haqq C, Nosrati M, Sudilovsky D, Crothers J, Khodabakhsh D, Pulliam BL, Federman S, Miller JR 3rd, Allen RE, Singer MI et al (2005) The gene expression signatures of melanoma progression. Proc Natl Acad Sci U S A 102(17):6092–6097
Seline PC, Norris DA, Horikawa T, Fujita M, Middleton MH, Morelli JG (1996) Expression of E and P-cadherin by melanoma cells decreases in progressive melanomas and following ultraviolet radiation. J Invest Dermatol 106(6):1320–1324
Ciolczyk-Wierzbicka D, Laidler P (2018) The inhibition of invasion of human melanoma cells through N-cadherin knock-down. Med Oncol 35(4):42
Zhang GM, Zheng L, He H, Song CC, Zhang ZJ, Cao XK, Lei CZ, Lan XY, Qi XL, Chen H, Huang YZ (2018) Associations of GBP2 gene copy number variations with growth traits and transcriptional expression in Chinese cattle. Gene 647:101–106
Yu S, Yu X, Sun L, Zheng Y, Chen L, Xu H, Jin J, Lan Q, Chen CC, Li M (2020) GBP2 enhances glioblastoma invasion through Stat3/fibronectin pathway. Oncogene 39(27):5042–5055
Miao Q, Ge M, Huang L (2017) Up-regulation of GBP2 is associated with neuronal apoptosis in rat brain cortex following traumatic brain injury. Neurochem Res 42(5):1515–1523
Rahvar F, Salimi M, Mozdarani H (2020) Plasma GBP2 promoter methylation is associated with advanced stages in breast cancer. Genet Mol Biol 43(4):e20190230
Guimaraes DP, Oliveira IM, de Moraes E, Paiva GR, Souza DM, Barnas C, Olmedo DB, Pinto CE, Faria PA, De Moura Gallo CV et al (2009) Interferon-inducible guanylate binding protein (GBP)-2: a novel p53-regulated tumor marker in esophageal squamous cell carcinomas. Int J Cancer 124(2):272–279
Wang Q, Wang X, Liang Q, Wang S, Xiwen L, Pan F, Chen H, Li D (2018b) Distinct prognostic value of mRNA expression of guanylate-binding protein genes in skin cutaneous melanoma. Oncol Lett 15(5):7914–7922
Lai W, Chen S, Wu H, Guan Y, Liu L, Zeng Y, Zhao H, Jiang J, Chu Z (2011) PRL-3 promotes the proliferation of LoVo cells via the upregulation of KCNN4 channels. Oncol Rep 26(4):909–917
Jiang SH, Zhu LL, Zhang M, Li RK, Yang Q, Yan JY, Zhang C, Yang JY, Dong FY, Dai M, Hu LP, Li J, Li Q, Wang YH, Yang XM, Zhang YL, Nie HZ, Zhu L, Zhang XL, Tian GA, Zhang XX, Cao XY, Tao LY, Huang S, Jiang YS, Hua R, Qian Luo K, Gu JR, Sun YW, Hou S, Zhang ZG (2019) GABRP regulates chemokine signalling, macrophage recruitment and tumour progression in pancreatic cancer through tuning KCNN4-mediated ca(2+) signalling in a GABA-independent manner. Gut 68(11):1994–2006
Gole HK, Tharp DL, Bowles DK (2014) Upregulation of intermediate-conductance Ca2+−activated K+ channels (KCNN4) in porcine coronary smooth muscle requires NADPH oxidase 5 (NOX5). PLoS One 9(8):e105337
Jiang S, Zhu L, Yang J, Hu L, Gu J, Xing X, Sun Y, Zhang Z (2017) Integrated expression profiling of potassium channels identifys KCNN4 as a prognostic biomarker of pancreatic cancer. Biochem Biophys Res Commun 494(1–2):113–119
Lai W, Liu L, Zeng Y, Wu H, Xu H, Chen S, Chu Z (2013) KCNN4 channels participate in the EMT induced by PRL-3 in colorectal cancer. Med Oncol 30(2):566
Bulk E, Ay AS, Hammadi M, Ouadid-Ahidouch H, Schelhaas S, Hascher A, Rohde C, Thoennissen NH, Wiewrodt R, Schmidt E, Marra A, Hillejan L, Jacobs AH, Klein HU, Dugas M, Berdel WE, Müller-Tidow C, Schwab A (2015) Epigenetic dysregulation of KCa 3.1 channels induces poor prognosis in lung cancer. Int J Cancer 137(6):1306–1317
Lallet-Daher H, Roudbaraki M, Bavencoffe A, Mariot P, Gackiere F, Bidaux G, Urbain R, Gosset P, Delcourt P, Fleurisse L et al (2009) Intermediate-conductance Ca2+−activated K+ channels (IKCa1) regulate human prostate cancer cell proliferation through a close control of calcium entry. Oncogene 28(15):1792–1806
Rabjerg M, Olivan-Viguera A, Hansen LK, Jensen L, Sevelsted-Moller L, Walter S, Jensen BL, Marcussen N, Kohler R (2015) High expression of KCa3.1 in patients with clear cell renal carcinoma predicts high metastatic risk and poor survival. PLoS One 10(4):e0122992
Zhou H, Yang YH, Basile JR (2017) Characterization of the effects of Semaphorin 4D signaling on angiogenesis. Methods Mol Biol 1493:429–441
Mou P, Zeng Z, Li Q, Liu X, Xin X, Wannemacher KM, Ruan C, Li R, Brass LF, Zhu L (2013) Identification of a calmodulin-binding domain in Sema4D that regulates its exodomain shedding in platelets. Blood 121(20):4221–4230
Sierra JR, Corso S, Caione L, Cepero V, Conrotto P, Cignetti A, Piacibello W, Kumanogoh A, Kikutani H, Comoglio PM, Tamagnone L, Giordano S (2008) Tumor angiogenesis and progression are enhanced by Sema4D produced by tumor-associated macrophages. J Exp Med 205(7):1673–1685
Wang Y, Zhao H, Zhi W (2020c) SEMA4D under the posttranscriptional regulation of HuR and miR-4319 boosts cancer progression in esophageal squamous cell carcinoma. Cancer Biol Ther 21(2):122–129
Xia Y, Cai XY, Fan JQ, Zhang LL, Ren JH, Li ZY, Zhang RG, Zhu F, Wu G (2019) The role of sema4D in vasculogenic mimicry formation in non-small cell lung cancer and the underlying mechanisms. Int J Cancer 144(9):2227–2238
Soong J, Chen Y, Shustef EM, Scott GA (2012) Sema4D, the ligand for Plexin B1, suppresses c-met activation and migration and promotes melanocyte survival and growth. J Invest Dermatol 132(4):1230–1238
Warzecha CC, Jiang P, Amirikian K, Dittmar KA, Lu H, Shen S, Guo W, Xing Y, Carstens RP (2010) An ESRP-regulated splicing programme is abrogated during the epithelial-mesenchymal transition. EMBO J 29(19):3286–3300
Ishii H, Saitoh M, Sakamoto K, Kondo T, Katoh R, Tanaka S, Motizuki M, Masuyama K, Miyazawa K (2014) Epithelial splicing regulatory proteins 1 (ESRP1) and 2 (ESRP2) suppress cancer cell motility via different mechanisms. J Biol Chem 289(40):27386–27399
Vadlamudi Y, Dey DK, Kang SC (2020) Emerging multi-cancer regulatory role of ESRP1: orchestration of alternative splicing to control EMT. Curr Cancer Drug Targets 20(9):654–665
Yao J, Caballero OL, Huang Y, Lin C, Rimoldi D, Behren A, Cebon JS, Hung MC, Weinstein JN, Strausberg RL, Zhao Q (2016) Altered expression and splicing of ESRP1 in malignant melanoma correlates with epithelial-mesenchymal status and tumor-associated immune Cytolytic activity. Cancer Immunol Res 4(6):552–561
Ueda J, Matsuda Y, Yamahatsu K, Uchida E, Naito Z, Korc M, Ishiwata T (2014) Epithelial splicing regulatory protein 1 is a favorable prognostic factor in pancreatic cancer that attenuates pancreatic metastases. Oncogene 33(36):4485–4495
Deng G, Zhou X, Chen L, Yao Y, Li J, Zhang Y, Luo C, Sun L, Tang J (2020a) High expression of ESRP1 regulated by circ-0005585 promotes cell colonization in ovarian cancer. Cancer Cell Int 20:174
Chen ZH, Jing YJ, Yu JB, Jin ZS, Li Z, He TT, Su XZ (2019) ESRP1 induces cervical cancer cell G1-phase arrest via regulating Cyclin A2 mRNA stability. Int J Mol Sci 20(15). https://doi.org/10.3390/ijms20153705
Im JH, Buzzelli JN, Jones K, Franchini F, Gordon-Weeks A, Markelc B, Chen J, Kim J, Cao Y, Muschel RJ (2020) FGF2 alters macrophage polarization, tumour immunity and growth and can be targeted during radiotherapy. Nat Commun 11(1):4064
Wehrle-Haller B (2003) The role of kit-ligand in melanocyte development and epidermal homeostasis. Pigment Cell Res 16(3):287–296
Tetu P, Delyon J, Andre J, Reger de Moura C, Sabbah M, Ghanem GE, Battistella M, Mourah S, Lebbe C, Dumaz N (2020) FGF2 induces resistance to nilotinib through MAPK pathway activation in KIT mutated melanoma. Cancers (Basel) 12(5). https://doi.org/10.3390/cancers12051062
Lefevre G, Babchia N, Calipel A, Mouriaux F, Faussat AM, Mrzyk S, Mascarelli F (2009) Activation of the FGF2/FGFR1 autocrine loop for cell proliferation and survival in uveal melanoma cells. Invest Ophthalmol Vis Sci 50(3):1047–1057
Yu Y, Gao S, Li Q, Wang C, Lai X, Chen X, Wang R, Di J, Li T, Wang W et al (2012) The FGF2-binding peptide P7 inhibits melanoma growth in vitro and in vivo. J Cancer Res Clin Oncol 138(8):1321–1328
Baljinnyam E, Umemura M, Chuang C, De Lorenzo MS, Iwatsubo M, Chen S, Goydos JS, Ishikawa Y, Whitelock JM, Iwatsubo K (2014) Epac1 increases migration of endothelial cells and melanoma cells via FGF2-mediated paracrine signaling. Pigment Cell Melanoma Res 27(4):611–620
Higashi Y, Moribe H, Takagi T, Sekido R, Kawakami K, Kikutani H, Kondoh H (1997) Impairment of T cell development in deltaEF1 mutant mice. J Exp Med 185(8):1467–1479
Tan X, Banerjee P, Liu X, Yu J, Gibbons DL, Wu P, Scott KL, Diao L, Zheng X, Wang J, Jalali A, Suraokar M, Fujimoto J, Behrens C, Liu X, Liu CG, Creighton CJ, Wistuba II, Kurie JM (2018) The epithelial-to-mesenchymal transition activator ZEB1 initiates a prometastatic competing endogenous RNA network. J Clin Invest 128(4):1267–1282
Liang L, Zhang Z, Qin X, Gao Y, Zhao P, Liu J, Zeng W (2018) Gambogic acid inhibits melanoma through regulation of miR-199a-3p/ZEB1 Signalling. Basic Clin Pharmacol Toxicol 123(6):692–703
Zhu L, Liu Z, Dong R, Wang X, Zhang M, Guo X, Yu N, Zeng A (2019) MicroRNA-3662 targets ZEB1 and attenuates the invasion of the highly aggressive melanoma cell line A375. Cancer Manag Res 11:5845–5856
Root AR, Cao W, Li B, LaPan P, Meade C, Sanford J, Jin M, O'Sullivan C, Cummins E, Lambert M et al (2016) Development of PF-06671008, a Highly Potent Anti-P-cadherin/Anti-CD3 Bispecific DART molecule with extended half-life for the treatment of cancer. Antibodies (Basel) 5(1). https://doi.org/10.3390/antib5010006
Naujokat C, Fuchs D, Opelz G (2010) Salinomycin in cancer: a new mission for an old agent. Mol Med Rep 3(4):555–559
Gupta PB, Onder TT, Jiang G, Tao K, Kuperwasser C, Weinberg RA, Lander ES (2009) Identification of selective inhibitors of cancer stem cells by high-throughput screening. Cell 138(4):645–659
Liu Y, Hao Y, Li Y, Zheng Y, Dai J, Zhong F, Wei W, Fang Z (2020d) Salinomycin induces autophagic cell death in salinomycin-sensitive melanoma cells through inhibition of autophagic flux. Sci Rep 10(1):18515
Zhou J, Liu S, Wang Y, Dai W, Zou H, Wang S, Zhang J, Pan J (2019) Salinomycin effectively eliminates cancer stem-like cells and obviates hepatic metastasis in uveal melanoma. Mol Cancer 18(1):159
Ataga KI, Stocker J (2009) Senicapoc (ICA-17043): a potential therapy for the prevention and treatment of hemolysis-associated complications in sickle cell anemia. Expert Opin Investig Drugs 18(2):231–239
Paka L, Smith DE, Jung D, McCormack S, Zhou P, Duan B, Li JS, Shi J, Hao YJ, Jiang K, Yamin M, Goldberg ID, Narayan P (2017) Anti-steatotic and anti-fibrotic effects of the KCa3.1 channel inhibitor, Senicapoc, in non-alcoholic liver disease. World J Gastroenterol 23(23):4181–4190
Hohmann N, Blank A, Burhenne J, Suzuki Y, Mikus G, Haefeli WE (2019) Simultaneous phenotyping of CYP2E1 and CYP3A using oral chlorzoxazone and midazolam microdoses. Br J Clin Pharmacol 85(10):2310–2320
Quesnot N, Bucher S, Gade C, Vlach M, Vene E, Valenca S, Gicquel T, Holst H, Robin MA, Loyer P (2018) Production of chlorzoxazone glucuronides via cytochrome P4502E1 dependent and independent pathways in human hepatocytes. Arch Toxicol 92(10):3077–3091
Lu T, Liang WZ, Hao LJ, Kuo CC, Shieh P, Chou CT, Jan CR (2019) Action of chlorzoxazone on ca(2)(+)movement and viability in human oral cancer cells. Chin J Phys 62(3):123–130
Deng L, Li H, Su X, Zhang Y, Xu H, Fan L, Fan J, Han Q, Bai X, Zhao RC (2020b) Chlorzoxazone, a small molecule drug, augments immunosuppressive capacity of mesenchymal stem cells via modulation of FOXO3 phosphorylation. Cell Death Dis 11(3):158
Eamlamnam K, Patumraj S, Visedopas N, Thong-Ngam D (2006) Effects of Aloe vera and sucralfate on gastric microcirculatory changes, cytokine levels and gastric ulcer healing in rats. World J Gastroenterol 12(13):2034–2039
Zur E (2019) Oral viscous Sucralfate gel for post-procedural treatment of Barrett's esophagus. Int J Pharm Compd 23(5):376–381
Falkowski S, Trouillas P, Duroux JL, Bonnetblanc JM, Clavere P (2011) Radiodermatitis prevention with sucralfate in breast cancer: fundamental and clinical studies. Support Care Cancer 19(1):57–65
Kneebone A, Mameghan H, Bolin T, Berry M, Turner S, Kearsley J, Graham P, Fisher R, Delaney G (2004) Effect of oral sucralfate on late rectal injury associated with radiotherapy for prostate cancer: a double-blind, randomized trial. Int J Radiat Oncol Biol Phys 60(4):1088–1097
Etiz D, Erkal HS, Serin M, Kucuk B, Hepari A, Elhan AH, Tulunay O, Cakmak A (2000) Clinical and histopathological evaluation of sucralfate in prevention of oral mucositis induced by radiation therapy in patients with head and neck malignancies. Oral Oncol 36(1):116–120
Evans EK, Gardino AK, Kim JL, Hodous BL, Shutes A, Davis A, Zhu XJ, Schmidt-Kittler O, Wilson D, Wilson K et al (2017) A precision therapy against cancers driven by KIT/PDGFRA mutations. Sci Transl Med 9(414). https://doi.org/10.1126/scitranslmed.aao1690
Cocorocchio E, Pala L, Conforti F, Guerini-Rocco E, De Pas T, Ferrucci PF (2020) Successful treatment with avapritinib in patient with mucosal metastatic melanoma. Ther Adv Med Oncol 12:1758835920946158
Lubke J, Naumann N, Kluger S, Schwaab J, Metzgeroth G, Evans E, Gardino AK, Lengauer C, Hofmann WK, Fabarius A et al (2019) Inhibitory effects of midostaurin and avapritinib on myeloid progenitors derived from patients with KIT D816V positive advanced systemic mastocytosis. Leukemia 33(5):1195–1205
Dhillon S (2020) Avapritinib: first approval. Drugs 80(4):433–439
Tajima K, Hattori T, Takahashi H, Katahira H, Narimatsu A, Kumakura S, Goto H (2018) Rebamipide suppresses TNF-alpha production and macrophage infiltration in the conjunctiva. Vet Ophthalmol 21(4):347–352
Ueno T, Zenda S, Konishi T, Yurikusa T, Shibasaki Y, Nagamoto H, Fujii M (2019) The post hoc analysis comparing the severity grades of chemoradiotherapy-induced oral mucositis scored between the central and local assessors in a multicenter, randomized controlled trial of rebamipide for head and neck cancer. Int J Clin Oncol 24(3):241–247
Fukui K, Yachi K, Yoshida H, Tanji K, Matsumiya T, Hayakari R, Tsuruga K, Tanaka H, Imaizumi T (2017) Rebamipide reduces amyloid-beta 1-42 (Abeta42) production and ameliorates Abeta43-lowered cell viability in cultured SH-SY5Y human neuroblastoma cells. Neurosci Res 124:40–50
Tanigawa T, Pai R, Arakawa T, Tarnawski AS (2007) Rebamipide inhibits gastric cancer cell growth. Dig Dis Sci 52(1):240–247
Weisberg E, Meng C, Case AE, Sattler M, Tiv HL, Gokhale PC, Buhrlage SJ, Liu X, Yang J, Wang J, Gray N, Stone RM, Adamia S, Dubreuil P, Letard S, Griffin JD (2019) Comparison of effects of midostaurin, crenolanib, quizartinib, gilteritinib, sorafenib and BLU-285 on oncogenic mutants of KIT, CBL and FLT3 in haematological malignancies. Br J Haematol 187(4):488–501
Papadopoulos KP, Ben-Ami E, Patnaik A, Trone D, Li J, Demetri GD (2018) Safety and tolerability of quizartinib, a FLT3 inhibitor, in advanced solid tumors: a phase 1 dose-escalation trial. BMC Cancer 18(1):790
Herbert KE, Prince HM, Ritchie DS, Seymour JF (2010) The role of ancestim (recombinant human stem-cell factor, rhSCF) in hematopoietic stem cell mobilization and hematopoietic reconstitution. Expert Opin Biol Ther 10(1):113–125
Motawi TM, Sadik NA, Fahim SA, Shouman SA (2015) Combination of imatinib and clotrimazole enhances cell growth inhibition in T47D breast cancer cells. Chem Biol Interact 233:147–156
Adinolfi B, Carpi S, Romanini A, Da Pozzo E, Castagna M, Costa B, Martini C, Olesen SP, Schmitt N, Breschi MC et al (2015) Analysis of the antitumor activity of Clotrimazole on A375 human melanoma cells. Anticancer Res 35(7):3781–3786
Penso J, Beitner R (1998) Clotrimazole and bifonazole detach hexokinase from mitochondria of melanoma cells. Eur J Pharmacol 342(1):113–117
Acknowledgements
We would like to appreciate the Grammarly (https://www.grammarly. com/upgrade) for editing the English text of a draft of this manuscript.
Availability of data and material
All data is available under reasonable request.
Funding
The authors received no specific funding for this work.
Author information
Authors and Affiliations
Contributions
Bo Chen and Donghong Sun had full access to all of the data in the study and take responsibility for the integrity of the data and the accuracy of the data analysis. Both contributed equally to the study and are co-first authors; Xiuni Qin contributed to collection of data; Xing-Hua Gao contributed to the study design, interpretation of the data, the writing of the manuscript, and the submission of the manuscript for publication.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed consent
For this type of study, formal consent is not required.
Consent for publication
All authors consent to the publication of this study.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Chen, B., Sun, D., Qin, X. et al. Screening and identification of potential biomarkers and therapeutic drugs in melanoma via integrated bioinformatics analysis. Invest New Drugs 39, 928–948 (2021). https://doi.org/10.1007/s10637-021-01072-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10637-021-01072-y