Abstract
Background
We previously reported that the anti-transforming growth factor-beta1 (TGF-β1) ribozymes directed by T7 and CMV promoters could reverse the character of activated hepatic stellate cells (HSCs) in vitro and improve fibrotic pathology in vivo. However, nonspecific elimination of the effects of TGF-β1 without selectivity might have unfavorable consequences, such as overwhelming inflammation, tissue necrosis, etc.
Aims
To establish an activated-HSC-specific gene silencing method and validate its feasibility for antifibrosis in vitro.
Methods
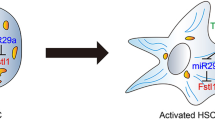
An artificial intronic microRNA (miRNA) expression system was established, containing three parts: (1) a 1,074-bp SM-α actin promoter SMP8, which is a kind of RNA polymerase II promoter and has no activity in normal liver-derived cells but is switched on during the activation of HSCs, (2) intron1 modified by inserting an artificial pre-miRNA sequence against TGF-β1, and (3) report gene enhanced green fluorescent proteins (EGFP). The feasibility of this system for artificial microRNA expression was validated through microRNA detection by real-time polymerase chain reaction (PCR). Alteration of biological characteristics of HSCs with the anti-TGF-β1 miRNAs was preliminarily evaluated by measuring the expression levels of TGF-β1 and its downstream molecules, including collagen I, matrix metalloproteinase 2 (MMP2), tissue inhibitor of metalloproteinase 1 (TIMP-1), etc.
Results
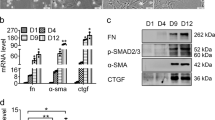
The microRNA expression system could successfully produce mature anti-TGF-β1 miRNAs in an activated-HSC-specific manner. The microRNA-induced inhibition rate of TGF-β1 reached 70% and above. Accompanied by TGF-β1 suppression, its downstream targets such as collagen I, MMP2, TIMP-1, etc. were also significantly downregulated in vitro.
Conclusions
Activated-HSC-cell-specific gene silencing could be induced well by the artificial intronic microRNA expression system to realize antifibrosis in vitro.






Similar content being viewed by others
References
Gressner AM, Weiskirchen R. Modern pathogenetic concepts of liver fibrosis suggest stellate cells and TGF-beta as major players and therapeutic targets. J Cell Mol Med. 2006;10:76–99.
Bataller R, Brenner DA. Liver fibrosis. J Clin Invest. 2005;115:209–218.
Bataller R, Paik YH, Lindquist JN, Lemasters JJ, Brenner DA. Hepatitis C virus core and nonstructural proteins induce fibrogenic effects in hepatic stellate cells. Gastroenterology. 2004;126:529–540.
Bridle KR, Crawford DH, Ramm GA. Identification and characterization of the hepatic stellate cell transferrin receptor. Am J Pathol. 2003;162:1661–1667.
Iredale JP. Models of liver fibrosis: Exploring the dynamic nature of inflammation and repair in a solid organ. J Clin Invest. 2007;117:539–548.
Qi Z, Atsuchi N, Ooshima A, Takeshita A, Ueno H. Blockade of type beta transforming growth factor signaling prevents liver fibrosis and dysfunction in the rat. Proc Natl Acad Sci USA. 1999;96:2345–2349.
Zardi EM, Dobrina A, Ambrosino G, Margiotta D, Polistina F, Afeltra A. New therapeutic approaches to liver fibrosis: A practicable route? Curr Med Chem. 2008;15:1628–1644.
Song YH, Chen XL, Kong XJ, et al. Ribozymes against TGFbeta1 reverse character of activated hepatic stellate cells in vitro and inhibit liver fibrosis in rats. J Gene Med. 2005;7:965–976.
Xu Y, Pasche B. TGF-beta signaling alterations and susceptibility to colorectal cancer. Hum Mol Genet. 2007;16(1):R14–20.
Schaffert D, Wagner E. Gene therapy progress and prospects: Synthetic polymer-based systems. Gene Ther. 2008;15:1131–1138.
Zhang M, Smith EP, Kuroda H, Banach W, Chernausek SD, Fagin JA. Targeted expression of a protease-resistant IGFBP-4 mutant in smooth muscle of transgenic mice results in IGFBP-4 stabilization and smooth muscle hypotrophy. J Biol Chem. 2002;277:21285–21290.
George J, Roulot D, Koteliansky VE, Bissell DM. In vivo inhibition of rat stellate cell activation by soluble transforming growth factor beta type II receptor: A potential new therapy for hepatic fibrosis. Proc Natl Acad Sci USA. 1999;96:12719–12724.
Shi R, Chiang VL. Facile means for quantifying microRNA expression by real-time PCR. Biotechniques. 2005;39:519–525.
Zeng Y, Wagner EJ, Cullen BR. Both natural and designed micro RNAs can inhibit the expression of cognate mRNAs when expressed in human cells. Mol Cell. 2002;9:1327–1333.
Zeng Y, Cullen BR. Sequence requirements for micro RNA processing and function in human cells. RNA. 2003;9:112–123.
Zhou H, Xia XG, Xu Z. An RNA polymerase II construct synthesizes short-hairpin RNA with a quantitative indicator and mediates highly efficient RNAi. Nucleic Acids Res. 2005;33:e62.
Buck AH, Santoyo-Lopez J, Robertson KA, Kumar DS, Reczko M, Ghazal P. Discrete clusters of virus-encoded micrornas are associated with complementary strands of the genome and the 7.2-kilobase stable intron in murine cytomegalovirus. J Virol. 2007;81:13761–13770.
Castoldi M, Schmidt S, Benes V, et al. A sensitive array for microRNA expression profiling (miChip) based on locked nucleic acids (LNA). RNA. 2006;12:913–920.
Rojkind M, Giambrone MA, Biempica L. Collagen types in normal and cirrhotic liver. Gastroenterology. 1979;76:710–719.
Rayburn ER, Zhang R. Antisense, RNAi, and gene silencing strategies for therapy: Mission possible or impossible? Drug Discov Today. 2008;13:513–521.
Kim D, Rossi J. RNAi mechanisms and applications. Biotechniques. 2008;44:613–616.
Huang C, Li M, Chen C, Yao Q. Small interfering RNA therapy in cancer: mechanism, potential targets, and clinical applications. Expert Opin Ther Targets. 2008;12:637–645.
Rockey DC. Gene therapy for hepatic fibrosis-bringing treatment into the new millennium. Hepatology. 1999;30:816–818.
Spankuch B, Strebhardt K. RNA interference-based gene silencing in mice: The development of a novel therapeutical strategy. Curr Pharm Des. 2005;11:3405–3419.
Xia XG, Zhou H, Xu Z. Transgenic RNAi: Accelerating and expanding reverse genetics in mammals. Transgenic Res. 2006;15:271–275.
Cai X, Zhou J, Chang Y, Sun X, Li P, Lin J. Targeting gene therapy for hepatocarcinoma cells with the E. coli purine nucleoside phosphorylase suicide gene system directed by a chimeric alpha-fetoprotein promoter. Cancer Lett. 2008;264:71–82.
Wang J, Niu W, Nikiforov Y, et al. Targeted overexpression of IGF-I evokes distinct patterns of organ remodeling in smooth muscle cell tissue beds of transgenic mice. J Clin Invest. 1997;100:1425–1439.
Moreira RK. Hepatic stellate cells and liver fibrosis. Arch Pathol Lab Med. 2007;131:1728–1734.
Vogel S, Piantedosi R, Frank J, et al. An immortalized rat liver stellate cell line (HSC-T6): A new cell model for the study of retinoid metabolism in vitro. J Lipid Res. 2000;41:882–893.
Kim Y, Ratziu V, Choi SG, et al. Transcriptional activation of transforming growth factor beta1 and its receptors by the Kruppel-like factor Zf9/core promoter-binding protein and Sp1. Potential mechanisms for autocrine fibrogenesis in response to injury. J Biol Chem. 1998;273:33750–33758.
Zhou XM, Lin JS, Shi Y, et al. Establishment of a screening system for selection of siRNA target sites directed against hepatitis B virus surface gene. Acta Biochim Biophys Sin (Shanghai). 2005;37:310–316.
Cramer P, Armache KJ, Baumli S, et al. Structure of eukaryotic RNA polymerases. Annu Rev Biophys. 2008;37:337–352.
Lee Y, Kim M, Han J, et al. MicroRNA genes are transcribed by RNA polymerase II. EMBO J. 2004;23:4051–4060.
Lin SL, Miller JD, Ying SY. Intronic microRNA (miRNA). J Biomed Biotechnol. 2006;2006:26818.
Rajewsky N. microRNA target predictions in animals. Nat Genet. 2006;38(Suppl):S8–13.
Krek A, Grun D, Poy MN, et al. Combinatorial microRNA target predictions. Nat Genet. 2005;37:495–500.
Lewis BP, Shih IH, Jones-Rhoades MW, Bartel DP, Burge CB. Prediction of mammalian microRNA targets. Cell. 2003;115:787–798.
Zhang B, Wang Q, Pan X. MicroRNAs and their regulatory roles in animals and plants. J Cell Physiol. 2007;210:279–289.
Jones-Rhoades MW, Bartel DP, Bartel B. MicroRNAS and their regulatory roles in plants. Annu Rev Plant Biol. 2006;57:19–53.
Rhoades MW, Reinhart BJ, Lim LP, Burge CB, Bartel B, Bartel DP. Prediction of plant microRNA targets. Cell. 2002;110:513–520.
Cuellar TL, McManus MT. MicroRNAs and endocrine biology. J Endocrinol. 2005;187:327–332.
Bartel DP. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–297.
Cheng K, Yang N, Mahato RI. TGF-beta1 gene silencing for treating liver fibrosis. Mol Pharm. 2009;6:772–779.
Kim KH, Kim HC, Hwang MY, et al. The antifibrotic effect of TGF-beta1 siRNAs in murine model of liver cirrhosis. Biochem Biophys Res Commun. 2006;343:1072–1078.
Li G, Xie Q, Shi Y, et al. Inhibition of connective tissue growth factor by siRNA prevents liver fibrosis in rats. J Gene Med. 2006;8:889–900.
Xu W, Wang LW, Shi JZ, Gong ZJ. Effects of RNA interference targeting transforming growth factor-beta 1 on immune hepatic fibrosis induced by Concanavalin A in mice. Hepatobiliary Pancreat Dis Int. 2009;8:300–308.
Schnabl B, Kweon YO, Frederick JP, Wang XF, Rippe RA, Brenner DA. The role of Smad3 in mediating mouse hepatic stellate cell activation. Hepatology. 2001;34:89–100.
Acknowledgments
We thank Mr. James A Fagin (University of Cincinnati Medical Center, USA) for the plasmid pSMP8. Supported by grants from the National Natural Science Foundation of China (No. 30600277) and the National Basic Research Program of China (No. 2007CB512903).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Ying Chang and Hua-jun Jiang contributed equally to this work.
Rights and permissions
About this article
Cite this article
Chang, Y., Jiang, Hj., Sun, Xm. et al. Hepatic Stellate Cell-Specific Gene Silencing Induced by an Artificial MicroRNA for Antifibrosis In Vitro. Dig Dis Sci 55, 642–653 (2010). https://doi.org/10.1007/s10620-009-1021-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10620-009-1021-z