Abstract
Chronic wasting disease (CWD) is a transmissible spongiform encephalopathy caused by prions that has spread across cervid species in North America since the 1960s and has recently been detected in Eurasian cervids. The Association of Zoos and Aquariums (AZA) considers CWD to be of major concern for cervids in AZA-accredited facilities because of the indirect transmission risk of the disease and the impact of CWD regulatory protocols on captive breeding programs. Vulnerability to CWD is affected by variation in the PRNP gene that encodes the prion protein. We therefore sequenced PRNP in Pere David’s deer (Elaphurus davidianus), a species that was extinct in the wild for more than a century and descends from ca. 11 founders. In 27 individuals, we detected two PRNP haplotypes, designated Elad1 (51 of 54 sequences) and Elad2 (3 of 54 sequences). The two haplotypes are separated by four single nucleotide polymorphisms (SNPs), three of which are non-synonymous. Both Elad1 and Elad2 have polymorphisms that in other cervid taxa are associated with reduced vulnerability to CWD. The two haplotypes are more similar in sequence to PRNP in other cervids than to each other. This suggests that PRNP in cervids may have been under long-term balancing selection, as has been shown for PRNP in non-cervid taxa, and which could account for the presence of multiple haplotypes among founders. There may be a fitness benefit in maintaining both PRNP haplotypes in the species because variation in the prion protein amino acid sequence can limit transmission of CWD.

Similar content being viewed by others
Data availability
PRNP haplotype sequences for Pere David’s deer have been deposited in GenBank under accession numbers MW804582 and MW804583. Sample information is listed in Table 1.
Code availability
Not applicable.
References
Almberg ES, Cross PC, Johnson CJ, Heisey DM, Richards BJ (2011) Modeling routes of chronic wasting disease transmission: environmental prion persistence promotes deer population decline and extinction. PLoS ONE 6:e19896
Angers R, Kang HE, Napier D, Browning S, Seward T, Mathiason C, Balachandran A, McKenzie D, Castilla J, Soto C, Jewell J, Graham C, Hoover EA, Telling GC (2010) Prion strain mutation determined by prion protein conformational compatibility and primary structure. Science 328:1154–1158
Arifin MI, Staskevicius A, Shim SY, Huang Y-H, Fenton H, McLoughlin PD, Mitchell G, Cullingham CI, Gilch S (2020) Large-scale prion protein genotyping in Canadian caribou populations and potential impact on chronic wasting disease susceptibility. Mol Ecol 29:3830–3840
Arifin MI, Hannaoui S, Chang SC, Thapa S, Schatzl HM, Gilch S (2021) Cervid prion protein polymorphisms: role in chronic wasting disease pathogenesis. Int J Mol Sci 22:2271
AZA Board of Directors A (2003) Guidelines for chronic wasting disease surveillance. https://www.aza.org/guidelines-for-chronic-wasting-disease-surveillance
Balachandran A, Harrington NP, Algire J, Soutyrine A, Spraker TR, Jeffrey M, González L, O’Rourke KI (2010) Experimental oral transmission of chronic wasting disease to red deer (Cervus elaphus elaphus): early detection and late stage distribution of protease-resistant prion protein. Can Vet J 51:169–178
Bandelt HJ, Forster P, Röhl A (1999) Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol 16:37–48
Bartelt-Hunt SL, Bartz JC (2013) Behavior of prions in the environment: implications for prion biology. PLOS Pathogens 9:e1003113
Belay ED, Maddox RA, Williams ES, Miller MW, Gambetti P, Schonberger LB (2004) Chronic wasting disease and potential transmission to humans. Emerg Infect Dis 10:977–984
Benestad SL, Mitchell G, Simmons M, Ytrehus B, Vikøren T (2016) First case of chronic wasting disease in Europe in a Norwegian free-ranging reindeer. Vet Res 47:88
Béringue V, Vilotte J-L, Laude H (2008) Prion agent diversity and species barrier. Vet Res 39:47
Betts MJ, Russell RB (2003) Amino acid properties and consequences of substitutions. Bioinformatics for geneticists. Wiley, Chichester
Brandt AL, Kelly AC, Green ML, Shelton P, Novakofski J, Mateus-Pinilla NE (2015) Prion protein gene sequence and chronic wasting disease susceptibility in white-tailed deer (Odocoileus virginianus). Prion 9:449–462
Brandt AL, Green ML, Ishida Y, Roca AL, Novakofski J, Mateus-Pinilla NE (2018) Influence of the geographic distribution of prion protein gene sequence variation on patterns of chronic wasting disease spread in white-tailed deer (Odocoileus virginianus). Prion 12:204–215
CDC (2020) Chronic wasting disease (CWD): occurrence. Centers for Disease Control and Prevention. https://www.cdc.gov/prions/cwd/occurrence.html
Charlesworth D (2006) Balancing selection and its effects on sequences in nearby genome regions. PLOS Genet 2:e64
Cheng YC, Musiani M, Cavedon M, Gilch S (2017) High prevalence of prion protein genotype associated with resistance to chronic wasting disease in one Alberta woodland caribou population. Prion 11:136–142
Collinge J, Clarke AR (2007) A general model of prion strains and their pathogenicity. Science 318:930–936
Cullingham CI, Peery RM, Dao A, McKenzie DI, Coltman DW (2020) Predicting the spread-risk potential of chronic wasting disease to sympatric ungulate species. Prion 14:56–66
Cunningham AA, Daszak P, Wood JLN (2017) One Health, emerging infectious diseases and wildlife: two decades of progress? Philos Trans R Soc Lond B Biol Sci 372:20160167
Edgeworth JA, Jackson GS, Clarke AR, Weissmann C, Collinge J (2009) Highly sensitive, quantitative cell-based assay for prions adsorbed to solid surfaces. PNAS 106:3479–3483
Edmunds DR, Kauffman MJ, Schumaker BA, Lindzey FG, Cook WE, Kreeger TJ, Grogan RG, Cornish TE (2016) Chronic wasting disease drives population decline of white-tailed deer. PLoS ONE 11:e0161127
Fernández MH, Vrba ES (2005) A complete estimate of the phylogenetic relationships in Ruminantia: a dated species-level supertree of the extant ruminants. Biol Rev 80:269–302
Geremia C, Hoeting JA, Wolfe LL, Galloway NL, Antolin MF, Spraker TR, Miller MW, Hobbs NT (2015) Age and repeated biopsy influence antemortem PrP (CWD) testing in mule deer (Odocoileus hemionus) in Colorado, USA. J Wildl Dis 51:801–810
Haley NJ, Seelig DM, Zabel MD, Telling GC, Hoover EA (2009) Detection of CWD prions in urine and saliva of deer by transgenic mouse bioassay. PLoS ONE 4:e4848–e4848
Haley NJ, Mathiason CK, Carver S, Zabel M, Telling GC, Hoover EA (2011) Detection of chronic wasting disease prions in salivary, urinary, and intestinal tissues of deer: potential mechanisms of prion shedding and transmission. J Virol 85:6309–6318
Haley NJ, Rielinger R, Davenport KA, Rourke K, Mitchell G, Richt JA (2017) Estimating chronic wasting disease susceptibility in cervids using real-time quaking-induced conversion. J Gen Virol 98:2882–2892
Hanke M, Wink M (1994) Direct DNA sequencing of PCR-amplified vector inserts following enzymatic degradation of primer and dNTPs. Biotechniques 17:858–860
Happ G, Huson H, Beckmen K, Kennedy L (2007) Prion protein genes in caribou from Alaska. J Wildl Dis 43:224–228
Harrathi C, Fernández-Borges N, Eraña H, Elezgarai SR, Venegas V, Charco JM, Castilla J (2019) Insights into the bidirectional properties of the sheep-deer prion transmission barrier. Mol Neurobiol 56:5287–5303
Hazards EPanel oB, Ricci A, Allende A, Bolton D, Chemaly M, Davies R, Fernández Escámez PS, Gironés R, Herman L, Koutsoumanis K, Lindqvist R, Nørrung B, Robertson L, Sanaa M, Skandamis P, Snary E, Speybroeck N, Ter Kuile B, Threlfall J, Wahlström H, Benestad S, Gavier-Widen D, Miller MW, Ru G, Telling GC, Tryland M, Ortiz Pelaez A, Simmons M (2017) Chronic wasting disease (CWD) in cervids. EFSA J 15:4667
Hazra A (2017) Using the confidence interval confidently. J Thorac Dis 9:4125–4130
Heckeberg NS (2020) The systematics of the Cervidae: a total evidence approach. PeerJ 8:e8114
Henderson DM, Denkers ND, Hoover CE, Garbino N, Mathiason CK, Hoover EA (2015) Longitudinal detection of prion shedding in saliva and urine by chronic wasting disease-infected deer by real-time quaking-induced conversion. J Virol 89:9338–9347
Hu H, Jiang Z (2002) Trial release of Père David’s deer Elaphurus davidianus in the Dafeng Reserve, China. Oryx 36:196–199
Ishida Y, Tian T, Brandt AL, Kelly AC, Shelton P, Roca AL, Novakofski J, Mateus-Pinilla NE (2020) Association of chronic wasting disease susceptibility with prion protein variation in white-tailed deer (Odocoileus virginianus). Prion 14:214–225
Jewell JE, Conner MM, Wolfe LL, Miller MW, Williams ES (2005) Low frequency of PrP genotype 225SF among free-ranging mule deer (Odocoileus hemionus) with chronic wasting disease. J Gen Virol 86:2127–2134
Jiang Z, Yu C, Feng Z, Zhang L, Xia J, Ding Y, Lindsay N (2000) Reintroduction and recovery of Père David’s deer in China. Wildl Soc Bull 28:681–687
Johnson C, Johnson J, Clayton M, McKenzie D, Aiken J (2003) Prion protein gene heterogeneity in free-ranging white-tailed deer within the chronic wasting disease affected region of Wisconsin. J Wildl Dis 39:576–581
Jones FW (1951) A contribution to the history and anatomy of Père David’s Deer (Elaphurus davidianus). Proc Zool Soc Lond 121:319–370
Kelly AC, Mateus-Pinilla NE, Diffendorfer J, Jewell E, Ruiz MO, Killefer J, Shelton P, Beissel T, Novakofski J (2008) Prion sequence polymorphisms and chronic wasting disease resistance in Illinois white-tailed deer (Odocoileus virginianus). Prion 2:28–36
Klein J, Sato A, Nagl S, O’hUigín C (1998) Molecular trans-species polymorphism. Annu Rev Ecol Evol Syst 29:1–21
Koenig D, Hagmann J, Li R, Bemm F, Slotte T, Neuffer B, Wright SI, Weigel D (2019) Long-term balancing selection drives evolution of immunity genes in Capsella. eLife 8:e43606
Kramm C, Gomez-Gutierrez R, Soto C, Telling G, Nichols T, Morales R (2020) In vitro detection of chronic wasting disease (CWD) prions in semen and reproductive tissues of white tailed deer bucks (Odocoileus virginianus). PLoS ONE 14:e0226560
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549
Kurt TD, Sigurdson CJ (2016) Cross-species transmission of CWD prions. Prion 10:83–91
Kurt TD, Perrott MR, Wilusz CJ, Wilusz J, Supattapone S, Telling GC, Zabel MD, Hoover EA (2007) Efficient in vitro amplification of chronic wasting disease PrPRES. J Virol 81:9605–9608
Kuznetsova A, McKenzie D, Banser P, Siddique T, Aiken JM (2014) Potential role of soil properties in the spread of CWD in western Canada. Prion 8:92–99
Kuznetsova A, Cullingham C, McKenzie D, Aiken JM (2018) Soil humic acids degrade CWD prions and reduce infectivity. PLoS Pathog 14:e1007414–e1007414
LaFauci G, Carp RI, Meeker HC, Ye X, Kim JI, Natelli M, Cedeno M, Petersen RB, Kascsak R, Rubenstein R (2006) Passage of chronic wasting disease prion into transgenic mice expressing Rocky Mountain elk (Cervus elaphus nelsoni) PrPC. J Gen Virol 87:3773–3780
Leigh J, Bryant D (2015) PopART: full-feature software for haplotype network construction. Methods Ecol Evol 6:1110–1116
Li C, Yang X, Ding Y, Zhang L, Fang H, Tang S, Jiang Z (2011) Do Père David’s deer lose memories of their ancestral predators? PLoS ONE 6:e23623
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452
Manjerovic MB, Green ML, Mateus-Pinilla N, Novakofski J (2014) The importance of localized culling in stabilizing chronic wasting disease prevalence in white-tailed deer populations. Prev Vet Med 113:139–145
Mateus-Pinilla N, Weng H-Y, Ruiz MO, Shelton P, Novakofski J (2013) Evaluation of a wild white-tailed deer population management program for controlling chronic wasting disease in Illinois, 2003–2008. Prev Vet Med 110:541–548
Mathiason CK, Powers JG, Dahmes SJ, Osborn DA, Miller KV, Warren RJ, Mason GL, Hays SA, Hayes-Klug J, Seelig DM, Wild MA, Wolfe LL, Spraker TR, Miller MW, Sigurdson CJ, Telling GC, Hoover EA (2006) Infectious prions in the saliva and blood of deer with chronic wasting disease. Science 314:133–136
Mathiason CK, Hays SA, Powers J, Hayes-Klug J, Langenberg J, Dahmes SJ, Osborn DA, Miller KV, Warren RJ, Mason GL, Hoover EA (2009) Infectious prions in pre-clinical deer and transmission of chronic wasting disease solely by environmental exposure. PLoS ONE 4:e5916–e5916
Mead S, Stumpf MPH, Whitfield J, Beck JA, Poulter M, Campbell T, Uphill JB, Goldstein D, Alpers M, Fisher EMC, Collinge J (2003) Balancing selection at the prion protein gene consistent with prehistoric kurulike epidemics. Science 300:640–643
Miller MW, Williams ES, Hobbs NT, Wolfe LL (2004) Environmental sources of prion transmission in mule deer. Emerg Infect Dis 10:1003–1006
Mitchell GB, Sigurdson CJ, O’Rourke KI, Algire J, Harrington NP, Walther I, Spraker TR, Balachandran A (2012) Experimental oral transmission of chronic wasting disease to reindeer (Rangifer tarandus tarandus). PLoS ONE 7:e39055
Monello RJ, Galloway NL, Powers JG, Madsen-Bouterse SA, Edwards WH, Wood ME, O’Rourke KI, Wild MA (2017) Pathogen-mediated selection in free-ranging elk populations infected by chronic wasting disease. PNAS 114:12208
Moore SJ, Kunkle R, Greenlee MHW, Nicholson E, Richt J, Hamir A, Waters WR, Greenlee J (2016) Horizontal transmission of chronic wasting disease in reindeer. Emerg Infect Dis 22:2142–2145
Nichols TA, Pulford B, Wyckoff AC, Meyerett C, Michel B, Gertig K, Hoover EA, Jewell JE, Telling GC, Zabel MD (2009) Detection of protease-resistant cervid prion protein in water from a CWD-endemic area. Prion 3:171–183
O’Rourke KI, Spraker TR, Hamburg LK, Besser TE, Brayton KA, Knowles DP (2004) Polymorphisms in the prion precursor functional gene but not the pseudogene are associated with susceptibility to chronic wasting disease in white-tailed deer. J Gen Virol 85:1339–1346
Ohmura T, Ueda T, Hashimoto Y, Imoto T (2001) Tolerance of point substitution of methionine for isoleucine in hen egg white lysozyme. Protein Eng Des Sel 14:421–425
Pitra C, Fickel J, Meijaard E, Groves C (2004) Evolution and phylogeny of old world deer. Mol Phylogenet Evol 33:880–895
Pritzkow S, Morales R, Lyon A, Concha-Marambio L, Urayama A, Soto C (2018) Efficient prion disease transmission through common environmental materials. J Biol Chem 293:3363–3373
Prusiner SB (1982) Novel proteinaceous infectious particles cause scrapie. Science 216:136–144
Prusiner S (1991) Molecular biology of prion diseases. Science 252:1515–1522
Raymond GJ, Bossers A, Raymond LD, O’Rourke KI, McHolland LE, Bryant PK 3rd, Miller MW, Williams ES, Smits M, Caughey B (2000) Evidence of a molecular barrier limiting susceptibility of humans, cattle and sheep to chronic wasting disease. EMBO J 19:4425–4430
Rhyan J, Miller M, Spraker T, McCollum M, Nol P, Wolfe L, Davis T, Creekmore L, O’Rourke K (2011) Failure of fallow deer (Dama dama) to develop chronic wasting disease when exposed to a contaminated environment and infected mule deer (Odocoileus hemionus). J Wildl Dis 47:739–744
Richards B (2021) Expanding distribution of chronic wasting disease. U.S. Geological Survey. https://www.usgs.gov/centers/nwhc/science/expanding-distribution-chronic-wasting-disease?qt-science_center_objects=0#qt-science_center_objects
Rivera NA, Brandt AL, Novakofski JE, Mateus-Pinilla NE (2019) Chronic wasting disease in cervids: prevalence, impact and management strategies. Vet Med (auckl) 10:123–139
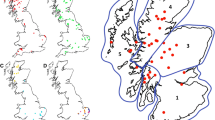
Robinson AL, Williamson H, Güere ME, Tharaldsen H, Baker K, Smith SL, Pérez-Espona S, Krojerová-Prokešová J, Pemberton JM, Goldmann W, Houston F (2019) Variation in the prion protein gene (PRNP) sequence of wild deer in Great Britain and mainland Europe. Vet Res 50:1
Saborio GP, Permanne B, Soto C (2001) Sensitive detection of pathological prion protein by cyclic amplification of protein misfolding. Nature 411:810–813
Saunders SE, Bartelt-Hunt SL, Bartz JC (2012) Occurrence, transmission, and zoonotic potential of chronic wasting disease. Emerg Infect Dis 18:369–376
Schafer EH (1968) Hunting parks and animal enclosures in ancient China. JESHO 11:318
Slate J (2005) Molecular evolution of the sheep prion protein gene. Proc Biol Sci 272:2371–2377
Stephens M, Smith NJ, Donnelly P (2001) A new statistical method for haplotype reconstruction from population data. Am J Hum Genet 68:978–989
Turvey ST, Barnes I, Marr M, Brace S (2017) Imperial trophy or island relict? A new extinction paradigm for Pere David’s deer: a Chinese conservation icon. R Soc Open Sci 4:171096
Viggers K, Lindenmayer D, Spratt D (1993) The importance of disease in reintroduction programmes. Wildl Res 20:687–698
Wildt D, Miller P, Koepfli K-P, Pukazhenthi B, Palfrey K, Livingston G, Beetem D, Shurter S, Gregory J, Takács M, Snodgrass K (2019) Breeding centers, private ranches, and genomics for creating sustainable wildlife populations. Bioscience 69:928–943
Williams ES, Young S (1980) Chronic wasting disease of captive mule deer: a spongiform encephalopathy. J Wildl Dis 16:89–98
Williams ES, Young S (1982) Spongiform encephalopathy of Rocky Mountain elk. J Wildl Dis 18:465–471
Yuan B, Xie S, Liu B, Xue D, Sun D (2019) Differential movement pattern of Père David’s deer associated with the temporal rhythm using GPS collar fix. Glob Ecol Conserv 18:e00641
Zeng Y, Jiang Z, Li C (2007) Genetic variability in relocated Père David’s deer (Elaphurus davidianus) populations—implications to reintroduction program. Conserv Genet 8:1051–1059
Zhang X, Deng C, Ding J, Ren Y, Zhao X, Qin S, Zhu S, Wang Z, Chai X, Huang H, Ding Y, Lu G, Zhu L (2016) Comparative genomics and metagenomics analyses of endangered Père David’s deer (Elaphurus davidianus) provide insights into population recovery. bioRxiv, 073528.
Zhang C, Chen L, Zhou Y, Wang K, Chemnick LG, Ryder OA, Wang W, Zhang G, Qiu Q (2017) Draft genome of the milu (Elaphurus davidianus). GigaScience 7
Zhang Y, Bai J, Zhu A, Chen R, Xue D, Zhong Z, Cheng Z (2021) Reversing extinction in China’s Père David’s deer. Science 371:685–685
Zhu L, Deng C, Zhao X, Ding J, Huang H, Zhu S, Wang Z, Qin S, Ding Y, Lu G, Yang Z (2018) Endangered Père David’s deer genome provides insights into population recovering. Evol Appl 11:2040–2053
Acknowledgements
For providing Pere David’s deer samples, we thank The Wilds in Cumberland, Ohio; the San Diego Zoo Institute for Conservation Research in San Diego, California; and the Wildlife Conservation Society and the Bronx Zoo in Bronx, New York. We would like to thank Dr. Dee McAloose, Steve Metzler, Dr. Jean Paré, Dr. Cynthia Steiner and anyone else who was involved in providing sample information and assisting in sample preparation and collection. We thank Alida de Flamingh for assistance in designing Supplementary Fig. 1.
Funding
This work was supported by the Cooperative State Research, Education, and Extension Service, U.S. Department of Agriculture, under Project Number ILLU 875-952 and ILLU 538-966. This work was also supported by the Francis M. and Harlie M. Clark Research Support Grant.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interests
The authors declare no conflict of interest in presenting this publication.
Ethics approval
Research was conducted under the Illinois Institutional Animal Care and Use Committee protocol 18212. Individual material transfer agreements are in place for each institution that contributed samples, and all terms were adhered to.
Consent to participate (include appropriate statements)
Not applicable.
Consent for publication (include appropriate statements)
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Perrin-Stowe, T.I.N., Ishida, Y., Terrill, E.E. et al. Variation in the PRNP gene of Pere David’s deer (Elaphurus davidianus) may impact genetic vulnerability to chronic wasting disease. Conserv Genet 23, 313–323 (2022). https://doi.org/10.1007/s10592-021-01419-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-021-01419-1