Abstract
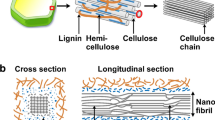
The recalcitrance of plant cell wall lignocellulosic biomass to deconstruction is a major hurdle to sustainable biofuel/bioproduct economy. A multitude of interactions stabilize lignocellulosic biomass structure. Among these, tight packing of hemicellulose-cellulose is partly responsible for biomass recalcitrance. Here, unrestrained molecular dynamics simulations are employed to understand the influence of the nature and pattern of naturally-occuring acetyl decorations of the xylan backbone on interactions with the (100) hydrophobic cellulose surface. Periodically O2-acetylated xylan (2AcX) assume twofold helical screw conformations that are stabilized by a combination of multiple hydrophobic contacts and hydrogen bonds with the hydrophobic cellulose surface. In contrast, acetylation at the O3 position in xylan obstructs interactions, thereby adopting threefold helical screw conformations that potentially preferably interact with lignin rather than cellulose. Fully acetylated xylan desorbs from the surface implying a minimum number of unsubstituted residues on the xylan backbone is required for interaction with the surface. The substituted residues must form ~ 20% fewer contacts than the unsubstituted residues to sustain stable twofold helical screw xylan conformations on the cellulose surface. Thus, specific roles of macromolecular conformations of cellulose and hemicellulose in influencing the supramolecular interactions and function of plant cell walls have been determined.







Similar content being viewed by others
Data availability
Not applicable.
References
Abraham MJ, Murtola T, Schulz R et al (2015) Gromacs: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2:19–25. https://doi.org/10.1016/j.softx.2015.06.001
Bergenstråhle M, Wohlert J, Larsson PT et al (2008) Dynamics of cellulose−water interfaces: NMR spin−lattice relaxation times calculated from atomistic computer simulations. J Phys Chem B 112:2590–2595
Berglund J, Angles d’Ortoli T, Vilaplana F et al (2016) A molecular dynamics study of the effect of glycosidic linkage type in the hemicellulose backbone on the molecular chain flexibility. Plant J 88:56–70. https://doi.org/10.1111/tpj.13259
Berglund J, Kishani S, Morais de Carvalho D et al (2020) Acetylation and sugar composition influence the (in)solubility of plant β-mannans and their interaction with cellulose surfaces. ACS Sustain Chem Eng 8:10027–10040. https://doi.org/10.1021/acssuschemeng.0c01716
Bootten TJ, Harris PJ, Melton LD, Newman RH (2004) Solid-state 13C-NMR spectroscopy shows that the xyloglucans in the primary cell walls of mung bean (Vigna radiata L.) occur in different domains: a new model for xyloglucan-cellulose interactions in the cell wall. J Exp Bot 55:571–583. https://doi.org/10.1093/jxb/erh065
Bosmans TJ, Stépán AM, Toriz G et al (2014) Assembly of debranched xylan from solution and on nanocellulosic surfaces. Biomacromol 15:924–930. https://doi.org/10.1021/bm4017868
Bromley JR, Busse-Wicher M, Tryfona T et al (2013) GUX1 and GUX2 glucuronyltransferases decorate distinct domains of glucuronoxylan with different substitution patterns. Plant J 74:423–434. https://doi.org/10.1111/tpj.12135
Brown DM, Goubet F, Wong VW et al (2007) Comparison of five xylan synthesis mutants reveals new insight into the mechanisms of xylan synthesis. Plant J 52:1154–1168. https://doi.org/10.1111/j.1365-313X.2007.03307.x
Busse-Wicher M, Gomes TCF, Tryfona T et al (2014) The pattern of xylan acetylation suggests xylan may interact with cellulose microfibrils as a twofold helical screw in the secondary plant cell wall of Arabidopsis thaliana. Plant J 79:492–506. https://doi.org/10.1111/tpj.12575
Busse-Wicher M, Li A, Silveira RL et al (2016) Evolution of xylan substitution patterns in gymnosperms and angiosperms: implications for xylan interaction with cellulose. Plant Physiol 171:2418–2431. https://doi.org/10.1104/pp.16.00539
Bussi G, Donadio D, Parrinello M (2007) Canonical sampling through velocity rescaling. J Chem Phys 126:014101. https://doi.org/10.1063/1.2408420
Celińska E, Nicaud JM, Białas W (2021) Hydrolytic secretome engineering in Yarrowia lipolytica for consolidated bioprocessing on polysaccharide resources: review on starch, cellulose, xylan, and inulin. Appl Microbiol Technol 105:975–989. https://doi.org/10.1007/s00253-021-11097-1
Chundawat SPS, Beckham GT, Himmel ME, Dale BE (2011) Deconstruction of lignocellulosic biomass to fuels and chemicals. Annu Rev Chem Biomol Eng 2:121–145. https://doi.org/10.1146/annurev-chembioeng-061010-114205
Cosgrove D (2014) Re-constructing our models of cellulose and primary cell wall assembly. Curr Opin Plant Biol 22:122–131
Cosgrove DJ, Jarvis MC (2012) Comparative structure and biomechanics of plant primary and secondary cell walls. Front Plant Sci 3:1–6. https://doi.org/10.3389/fpls.2012.00204
Crowe JD, Hao P, Pattathil S et al (2021) Xylan is critical for proper bundling and alignment of cellulose microfibrils in plant secondary cell walls. Front Plant Sci 12:1–19. https://doi.org/10.3389/fpls.2021.737690
Darden T, York D, Pedersen L (1993) Particle mesh Ewald: an N⋅log(N) method for Ewald sums in large systems. J Chem Phys 98:10089–10092. https://doi.org/10.1063/1.464397
de Carvalho DM, Berglund J, Marchand C et al (2019) Improving the thermal stability of different types of xylan by acetylation. Carbohydr Polym 220:132–140. https://doi.org/10.1016/j.carbpol.2019.05.063
Ebringerova A, Heinze T (2000) Xylan and xylan derivatives–biopolymers with valuable properties, 1. Naturally occurring xylans structures, isolation procedures and properties. Macromol Rapid Commun 21:869–883. https://doi.org/10.1002/1521-3927
Falcoz-Vigne L, Ogawa Y, Molina-Boisseau S et al (2017) Quantification of a tightly adsorbed monolayer of xylan on cellulose surface. Cellulose 24:3725–3739. https://doi.org/10.1007/s10570-017-1401-z
Fernandes AN, Thomas LH, Altaner CM et al (2011) Nanostructure of cellulose microfibrils in spruce wood. Proc Nat Acad Sci USA 108:E1195–E1203. https://doi.org/10.1073/pnas.1108942108
French AD, Johnson GP (2009) Cellulose and the twofold screw axis: modeling and experimental arguments. Cellulose 16:959–973. https://doi.org/10.1007/s10570-009-9347-4
Gomes TCF, Skaf MS (2012) Cellulose-builder: a toolkit for building crystalline structures of cellulose. J Comput Chem 33:1338–1346. https://doi.org/10.1002/jcc.22959
Grantham NJ, Wurman-Rodrich J, Terrett OM et al (2017) An even pattern of xylan substitution is critical for interaction with cellulose in plant cell walls. Nat Plants 3:859–865. https://doi.org/10.1038/s41477-017-0030-8
Gupta M, Rawal TB, Dupree P et al (2021) Spontaneous rearrangement of acetylated xylan on hydrophilic cellulose surfaces. Cellulose 28:3327. https://doi.org/10.1007/s10570-021-03706-z
Guvench O, Hatcher E, Venable RM et al (2009) CHARMM additive all-atom force field for glycosidic linkages between hexopyranoses. J Chem Theory Comput 5:2353–2370. https://doi.org/10.1021/ct900242e
Hess B (2008) P-LINCS: a parallel linear constraint solver for molecular simulation. J Chem Theory Comput 4:116–122. https://doi.org/10.1021/ct700200b
Hess B, Bekker H, Berendsen HJC, Fraaije JGEM (1997) LINCS: a linear constraint solver for molecular simulations. J Comput Chem 18:1463–1472. https://doi.org/10.1002/(SICI)1096-987X(199709)18:12%3c1463::AID-JCC4%3e3.0.CO;2-H
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14:33–38. https://doi.org/10.1016/0263-7855(96)00018-5
Jo S, Song KC, Desaire H et al (2011) Glycan reader: Automated sugar identification and simulation preparation for carbohydrates and glycoproteins. J Comput Chem 32:3135–3141. https://doi.org/10.1002/jcc.21886
Johnson AM, Kim H, Ralph J, Mansfield SD (2017) Natural acetylation impacts carbohydrate recovery during deconstruction of Populus trichocarpa wood. Biotechnol Biofuels 10:1–12. https://doi.org/10.1186/s13068-017-0734-z
Jorgensen WL, Chandrasekhar J, Madura JD et al (1983) Comparison of simple potential functions for simulating liquid water Comparison of simple potential functions for simulating liquid water. J Chem Phys 79:926. https://doi.org/10.1063/1.445869
Kabel MA, van den Borne H, Vincken JP et al (2007) Structural differences of xylans affect their interaction with cellulose. Carbohydr Polym 69:94–105. https://doi.org/10.1016/j.carbpol.2006.09.006
Kang X, Kirui A, Dickwella Widanage MC et al (2019) Lignin-polysaccharide interactions in plant secondary cell walls revealed by solid-state NMR. Nat Commun 10:347. https://doi.org/10.1038/s41467-018-08252-0
Köhnke T, Östlund Å, Brelid H (2011) Adsorption of arabinoxylan on cellulosic surfaces: influence of degree of substitution and substitution pattern on adsorption characteristics. Biomacromol 12:2633–2641. https://doi.org/10.1021/bm200437m
Kong Y, Li L, Fu S (2022) Insights from molecular dynamics simulations for interaction between cellulose microfibrils and hemicellulose. J Mater Chem A. https://doi.org/10.1039/d2ta03164g
Larsson PT, Hult EL, Wickholm K et al (1999) CP/MAS 13C-NMR spectroscopy applied to structure and interaction studies on cellulose I. Solid State Nucl Magn Reson 15:31–40. https://doi.org/10.1016/S0926-2040(99)00044-2
Liitiä T, Maunu SL, Hortling B et al (2003) Cellulose crystallinity and ordering of hemicelluloses in pine and birch pulps as revealed by solid-state NMR spectroscopic methods. Cellulose 10:307–316. https://doi.org/10.1023/A:1027302526861
Linder A, Bergman R, Bodin A, Gatenholm P (2003) Mechanism of assembly of xylan onto cellulose surfaces. Langmuir 19:5072–5077. https://doi.org/10.1021/la0341355
Lindman B, Medronho B, Alves L et al (2021) Hydrophobic interactions control the self-assembly of DNA and cellulose. Q Rev Biophys 54:1–22. https://doi.org/10.1017/S0033583521000019
Ling Z, Edwards JV, Nam S et al (2020) Conformational analysis of xylobiose by DFT quantum mechanics. Cellulose 27:1207–1224. https://doi.org/10.1007/s10570-019-02874-3
Luzar A (2000) Resolving the hydrogen bond dynamics conundrum. J Chem Phys 113:10663–10675. https://doi.org/10.1063/1.1320826
Martínez-Abad A, Berglund J, Toriz G et al (2017) Regular motifs in xylan modulate molecular flexibility and interactions with cellulose surfaces. Plant Physiol 175:1579–1592. https://doi.org/10.1104/pp.17.01184
Martínez-abad A, Jiménez-quero A (2020) Influence of the molecular motifs of mannan and xylan populations on their recalcitrance and organization in spruce softwoods. Green Chem 22:3956–3970. https://doi.org/10.1039/d0gc01207f
Martínez-abad A, Giummarella N, Vilaplana F, Lawoko M (2018) Differences in extractability under subcritical water reveal interconnected hemicellulose and lignin recalcitrance in birch hardwoods. Green Chem 20:2534–2546. https://doi.org/10.1039/c8gc00385h
Mazeau K, Moine C, Krausz P, Gloaguen V (2005) Conformational analysis of xylan chains. Carbohydr Res 340:2752–2760. https://doi.org/10.1016/j.carres.2005.09.023
Nishiyama Y, Langan P, Chanzy H (2002) Crystal structure and hydrogen-bonding system in cellulose Iβ from synchrotron X-ray and neutron fiber diffraction. J Am Chem Soc 124:9074–9082. https://doi.org/10.1021/ja0257319
Park SJ, Lee J, Qi Y et al (2019) CHARMM-GUI Glycan Modeler for modeling and simulation of carbohydrates and glycoconjugates. Glycobiology 29:320–331. https://doi.org/10.1093/glycob/cwz003
Parrinello M, Rahman A (1981) Polymorphic transitions in single crystals: a new molecular dynamics method. J Appl Phys 52:7182–7190. https://doi.org/10.1063/1.328693
Pawar PMA, Koutaniemi S, Tenkanen M, Mellerowicz EJ (2013) Acetylation of woody lignocellulose: significance and regulation. Front Plant Sci 4:1–8. https://doi.org/10.3389/fpls.2013.00118
Penttilä PA, Várnai A, Pere J et al (2013) Xylan as limiting factor in enzymatic hydrolysis of nanocellulose. Bioresour Technol 129:135–141. https://doi.org/10.1016/j.biortech.2012.11.017
Pereira CS, Silveira RL, Dupree P, Skaf MS (2017) Effects of xylan side-chain substitutions on xylan-cellulose interactions and implications for thermal pretreatment of cellulosic biomass. Biomacromol 18:1311–1321. https://doi.org/10.1021/acs.biomac.7b00067
Scheller HV, Ulvskov P (2010) Hemicelluloses. Annu Rev Plant Biol 61:263–289. https://doi.org/10.1146/annurev-arplant-042809-112315
Simmons TJ, Mortimer JC, Bernardinelli OD et al (2016) Folding of xylan onto cellulose fibrils in plant cell walls revealed by solid-state NMR. Nat Commun 7:13902. https://doi.org/10.1038/ncomms13902
Teleman A, Tenkanen M, Jacobs A, Dahlman O (2002) Characterization of O-acetyl-(4-O-methylglucurono)xylan isolated from birch and beech. Carbohydr Res 337:373–377. https://doi.org/10.1016/S0008-6215(01)00327-5
Torshin IY, Weber IT, Harrison RW (2002) Geometric criteria of hydrogen bonds in proteins and identification of “bifurcated” hydrogen bonds. Protein Eng 15:359–363. https://doi.org/10.1093/protein/15.5.359
Tryfona T, Bourdon M, Delgado Marques R et al (2023) Grass xylan structural variation suggests functional specialization and distinctive interaction with cellulose and lignin. Plant J 113:1004–1020. https://doi.org/10.1111/tpj.16096
Wierzbicki MP, Maloney V, Mizrachi E, Myburg AA (2019) Xylan in the middle: understanding xylan biosynthesis and its metabolic dependencies toward improving wood fiber for industrial processing. Front Plant Sci 10:1–29. https://doi.org/10.3389/fpls.2019.00176
Wohlert M, Benselfelt T, Wågberg L et al (2022) Cellulose and the role of hydrogen bonds: not in charge of everything. Cellulose 29:1–23. https://doi.org/10.1007/s10570-021-04325-4
Xiong G, Dama M, Pauly M (2015) Glucuronic acid moieties on xylan are functionally equivalent to O-acetyl-substituents. Mol Plant 8:1119–1121. https://doi.org/10.1016/j.molp.2015.02.013
Zhao Z, Crespi VH, Kubicki JD et al (2014) Molecular dynamics simulation study of xyloglucan adsorption on cellulose surfaces: effects of surface hydrophobicity and side-chain variation. Cellulose 21:1025–1039. https://doi.org/10.1007/s10570-013-0041-1
Acknowledgments
This study was supported by SERB-POWER grant received from Science and Engineering Research Board (SERB), under the Department of Science and Technology (DST) Government of India (Reference No. SPG/2021/002224). This research was supported by the Center for Lignocellulose Structure and Formation, an Energy Frontier Research Center funded by the U.S. Department of Energy, Office of Science, Basic Energy Sciences under Award DE- SC0001090 and by the Genomic Science Program, Office of Biological and Environmental Research, U. S. Department of Energy (DOE) under Contract FWP ERKP752. This research used resources of the National Energy Research Scientific Computing Center, supported under Contract No. DE-AC02- 05CH11231. Oak Ridge National Laboratory, which is supported by the Office of Science of the U.S. Department of Energy under Contract No. DE-AC05-00OR22725. This work used resources of the Compute and Data Environment for Science (CADES) at ORNL.
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
MG performed all the simulations and analysis and wrote the main manuscript along with preparation of figures and tables. PD, LP and JS provided important inputs to improve the manuscript and reviewed the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
There are no conflicts to declare.
Ethical approval and consent to participate
Not applicable.
Consent for publication
The authors give their consent for publishing the manuscript after peer review and its acceptance.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gupta, M., Dupree, P., Petridis, L. et al. Patterns in interactions of variably acetylated xylans with hydrophobic cellulose surfaces. Cellulose 30, 11323–11340 (2023). https://doi.org/10.1007/s10570-023-05584-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10570-023-05584-z