Abstract
Purpose
Promoter mutations may affect transcription and can be associated with human diseases. However, the promoters of the breast cancer (BC) genes are not regularly screened. Our goal was to investigate the BRCA2 promoter in order to study a possible correlation between impaired transcription and disease.
Methods
The proximal and core promoter of the BRCA2 gene was sequenced in 95 high-risk BC patients. A BRCA2-promoter insert [− 938 to + 312 from the transcription start site (TSS)] was generated and cloned into the firefly luciferase vector pGL4.10. Promoter variants and deletions were introduced by site-directed mutagenesis and quantified by Dual-Luciferase assays and semi-quantitative RT-PCR.
Results
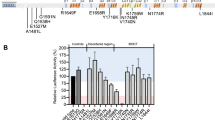
Three different variants were detected in high-risk BC patients: rs3092989, rs206118, and rs563971900. Functional mapping of 13 overlapping deletions revealed four down-regulating segments (TSS positions): −59_−10del/µdel3 (16% of activity of the wild-type construct), −104_−55del/µdel4 (62%), −239_−190del/µdel7 (39%), −464_−415/µdel12 (78%), suggesting the presence therein of putative transcriptional activator motifs. Additionally, six microdeletions rendered luciferase overexpression: +32_+81del/µdel1 (356%), −14_+36del/µdel2 (180%), −194_−145del/µdel6 (154%), −284_−235del/µdel8 (168%), −329_−280del/µdel9 (111%), and −509_−460del/µdel13 (139%), which is indicative of repressor elements. Functional assays of 15 promoter variants (including those detected in patients) showed that ten of them significantly altered expression with seven up-regulating (113–163%) and three down-regulating (rs551887850_G, rs570548398_T, rs55880202_T; 72–83%) SNPs. Eight of them were located in an ENCODE-DNase Hypersensitive Cluster (TSS − 185 to + 105) where most active transcriptional motifs are known to be placed.
Conclusions
BRCA2 expression is highly sensitive to promoter variations as most of them induced relevant changes. Moreover, we mapped critical regions of the BRCA2 promoter that may constitute potential targets for regulatory variants. Three SNPs moderately decreased luciferase activity, but confirmation of its potential pathogenicity requires further analysis. These data reinforce the need to screen the promoter regions of breast cancer genes with a view to discovering novel deleterious mutations.



Similar content being viewed by others
References
Roy R, Chun J, Powell SN (2012) BRCA1 and BRCA2: different roles in a common pathway of genome protection. Nat Rev Cancer 12:68–78. https://doi.org/10.1038/nrc3181
Nielsen FC, van Overeem Hansen T, Sørensen CS (2016) Hereditary breast and ovarian cancer: new genes in confined pathways. Nat Rev Cancer 16:599–612. https://doi.org/10.1038/nrc.2016.72
Miki Y, Swensen J, Shattuck-Eidens D et al (1994) A strong candidate for the breast and ovarian cancer susceptibility gene BRCA1. Science 266:66–71
Wooster R, Bignell G, Lancaster J et al (1995) Identification of the breast-cancer susceptibility gene Brca2. Nature 378:789–792
Foulkes WD (2008) Inherited susceptibility to common cancers. N Engl J Med 359:2143–2153. https://doi.org/10.1056/NEJMra0802968
Stratton MR, Rahman N (2008) The emerging landscape of breast cancer susceptibility. Nat Genet 40:17–22. https://doi.org/10.1038/ng.2007.53
Balmaña J, Domchek SM, Tutt A, Garber JE (2011) Stumbling blocks on the path to personalized medicine in breast cancer: the case of PARP inhibitors for BRCA1/2-associated cancers. Cancer Discov 1:29–34. https://doi.org/10.1158/2159-8274.CD-11-0048
Turner NC, Ashworth A (2011) Biomarkers of PARP inhibitor sensitivity. Breast Cancer Res Treat 127:283–286. https://doi.org/10.1007/s10549-011-1375-8
De Vooght KMK, Wijk R, Van, Van Solinge WW (2009) Management of gene promoter mutations in molecular diagnostics. Clin Chem 55:698–708. https://doi.org/10.1373/clinchem.2008.120931
Halvorsen M, Martin JS, Broadaway S, Laederach A (2010) Disease-associated mutations that alter the RNA structural ensemble. PLoS Genet 6:1–11. https://doi.org/10.1371/journal.pgen.1001074
Brewster BL, Rossiello F, French JD et al (2012) Identification of fifteen novel germline variants in the BRCA1 3′UTR reveals a variant in a breast cancer case that introduces a functional miR-103 target site. Hum Mutat 33:1665–1675. https://doi.org/10.1002/humu.22159
Fraile-Bethencourt E, Díez-Gómez B, Velásquez-Zapata V et al (2017) Functional classification of DNA variants by hybrid minigenes: identification of 30 spliceogenic variants of BRCA2 exons 17 and 18. PLoS Genet 13:e1006691. https://doi.org/10.1371/journal.pgen.1006691
Sanz DJ, Acedo A, Infante M et al (2010) A high proportion of DNA variants of BRCA1 and BRCA2 is associated with aberrant splicing in breast/ovarian cancer patients. Clin Cancer Res 16:1957–1967. https://doi.org/10.1158/1078-0432.CCR-09-2564
Quiles F, Menéndez M, Tornero E et al (2016) Investigating the effect of 28 BRCA1 and BRCA2 mutations on their related transcribed mRNA. Breast Cancer Res Treat. https://doi.org/10.1007/s10549-015-3676-9
Théry JC, Krieger S, Gaildrat P et al (2011) Contribution of bioinformatics predictions and functional splicing assays to the interpretation of unclassified variants of the BRCA genes. Eur J Hum Genet 19:1052–1058. https://doi.org/10.1038/ejhg.2011.100
Lheureux S, Lambert B, Krieger S et al (2011) Two novel variants in the 3′UTR of the BRCA1 gene in familial breast and/or ovarian cancer. Breast Cancer Res Treat 125:885–891. https://doi.org/10.1007/s10549-010-1165-8
Hindorff LA, Sethupathy P, Junkins HA et al (2009) Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc Natl Acad Sci USA 106:9362–9367. https://doi.org/10.1073/pnas.0903103106
Shim S-Y, Jeong HJ, Park HJ et al (2017) Functional variation of SHP-2 promoter is associated with preterm birth and delayed myelination and motor development in preterm infants. Sci Rep 7:6052. https://doi.org/10.1038/s41598-017-06401-x
Haberle V, Lenhard B (2015) Promoter architectures and developmental gene regulation. Semin Cell Dev Biol. https://doi.org/10.1016/j.semcdb.2016.01.014
Cooper DN (2002) Human gene mutation in pathology and evolution. J Inherit Metab Dis 25:157–182. https://doi.org/10.1023/A:1015621710660
Poncz M, Ballantine M, Solowiejczyk D et al (1982) β-Thalassemia in a Kurdish Jew: single base changes in the T-A-T-A box. J Biol Chem 257:5994–5996
Davis PL, Miron A, Andersen LM et al (1999) Isolation and initial characterization of the BRCA2 promoter. Oncogene 18:6000–6012
Velasco E, Infante M, Durán M et al (2007) Heteroduplex analysis by capillary array electrophoresis for rapid mutation detection in large multiexon genes. Nat Protoc 2:237–246. https://doi.org/10.1038/nprot.2006.482
de la Hoya M, Gutierrez-Enriquez S, Velasco E et al (2006) Genomic rearrangements at the BRCA1 locus in Spanish families with breast/ovarian cancer. Clin Chem 52:1480–1485. https://doi.org/10.1373/clinchem.2006.070110
Li W, Cowley A, Uludag M et al (2015) The EMBL-EBI bioinformatics web and programmatic tools framework. Nucleic Acids Res 43:W580–W584. https://doi.org/10.1093/nar/gkv279
Bryksin AV, Matsumura I (2010) Overlap extension PCR cloning: a simple and reliable way to create recombinant plasmids. Biotechniques 48:463–465. https://doi.org/10.2144/000113418
Dos Santos ES, Caputo SM, Castera L et al (2017) Assessment of the functional impact of germline BRCA1/2 variants located in non-coding regions in families with breast and/or ovarian cancer predisposition. Breast Cancer Res Treat. https://doi.org/10.1007/s10549-017-4602-0
Dunham I, Kundaje A, Aldred SF et al (2012) An integrated encyclopedia of DNA elements in the human genome. Nature 489:57–74. https://doi.org/10.1038/nature11247
Farré D, Roset R, Huerta M et al (2003) Identification of patterns in biological sequences at the ALGGEN server: PROMO and MALGEN. Nucleic Acids Res 31:3651–3653
Sandelin A, Alkema W, Engström P et al (2004) JASPAR: an open-access database for eukaryotic transcription factor binding profiles. Nucleic Acids Res 32:D91–D94. https://doi.org/10.1093/nar/gkh012
Acedo A, Sanz DJ, Durán M et al (2012) Comprehensive splicing functional analysis of DNA variants of the BRCA2 gene by hybrid minigenes. Breast Cancer Res 14:R87. https://doi.org/10.1186/bcr3202
Acedo A, Hernández-Moro C, Curiel-García Á et al (2015) Functional classification of BRCA2 DNA variants by splicing assays in a large minigene with 9 exons. Hum Mutat 36:210–221. https://doi.org/10.1002/humu.22725
Karimi M, Goldie LC, Cruickshank MN et al (2009) A critical assessment of the factors affecting reporter gene assays for promoter SNP function: a reassessment of -308 TNF polymorphism function using a novel integrated reporter system. Eur J Hum Genet 17:1454–1462. https://doi.org/10.1038/ejhg.2009.80
Liu Q, Thompson BA, Ward RL et al (2016) Understanding the pathogenicity of noncoding mismatch repair gene promoter variants in Lynch syndrome. Hum Mutat 37:417–426. https://doi.org/10.1002/humu.22971
de la Hoya M, Soukarieh O, López-Perolio I et al (2016) Combined genetic and splicing analysis of BRCA1 c.[594-2A> C; 641A> G] highlights the relevance of naturally occurring in-frame transcripts for developing disease gene variant classification algorithms. Hum Mol Genet 25:2256–2268. https://doi.org/10.1093/hmg/ddw094
Michailidou K, Lindström S, Dennis J et al (2017) Association analysis identifies 65 new breast cancer risk loci. Nature 551:92–94. https://doi.org/10.1038/nature24284
GTEx Consortium (2013) The genotype-tissue expression (GTEx) project. Nat Genet 45:580–585. https://doi.org/10.1038/ng.2653
Harper AR, Nayee S, Topol EJ (2015) Protective alleles and modifier variants in human health and disease. Nat Rev Genet 16:689–701. https://doi.org/10.1038/nrg4017
Capasso M, Diskin S, Cimmino F et al (2014) Common genetic variants in NEFL influence gene expression and neuroblastoma risk. Cancer Res 74:6913–6924. https://doi.org/10.1158/0008-5472.CAN-14-0431
Gochhait S, Bukhari SIA, Bairwa N et al (2007) Implication of BRCA2 -26G> A 5′ untranslated region polymorphism in susceptibility to sporadic breast cancer and its modulation by p53 codon 72 Arg> Pro polymorphism. Breast Cancer Res 9:R71. https://doi.org/10.1186/bcr1780
Maia A-T, Antoniou AC, O’Reilly M et al (2012) Effects of BRCA2 cis-regulation in normal breast and cancer risk amongst BRCA2 mutation carriers. Breast Cancer Res 14:R63. https://doi.org/10.1186/bcr3169
Kauffmann A, Rosselli F, Lazar V et al (2007) High expression of DNA repair pathways is associated with metastasis in melanoma patients. Oncogene 27:565–573. https://doi.org/10.1038/sj.onc.1210700
Sarasin A, Dessen P (2010) DNA repair pathways and human metastatic malignant melanoma. Curr Mol Med 10:413–418
Klein HL (2008) The consequences of Rad51 overexpression for normal and tumor cells. DNA Repair 7:686–693. https://doi.org/10.1016/j.dnarep.2007.12.008
Bièche I, Noguès C, Lidereau R (1999) Overexpression of BRCA2 gene in sporadic breast tumours. Oncogene 18:5232–5238. https://doi.org/10.1038/sj.onc.1202903
Acknowledgements
We are grateful to the breast/ovarian cancer patients and clinicians who participated in this study. EAV’s lab was supported by grants from the Spanish Ministry of Economy and Competitivity, Plan Nacional de I + D + I 2013–2016, ISCIII (Fis: PI13/01749) co-funded by FEDER from Regional Development European Funds (European Union), and Grant CSI090U14 from the Consejería de Educación (ORDEN EDU/122/2014), Junta de Castilla y León. EFB was supported by a predoctoral fellowship from the University of Valladolid and Banco de Santander (2015–2019).
Data availability
The datasets generated and/or analyzed during the current study are available in the Figshare repository, https://figshare.com/s/e7c982e6afb9907d56a0.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interests.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Fraile-Bethencourt, E., Valenzuela-Palomo, A., Díez-Gómez, B. et al. Genetic dissection of the BRCA2 promoter and transcriptional impact of DNA variants. Breast Cancer Res Treat 171, 53–63 (2018). https://doi.org/10.1007/s10549-018-4826-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-018-4826-7