Abstract
Six cadmium (Cd)-resistant microbial strains were isolated and their ability to immobilise Cd2+ in soil investigated. Cd-1, Cd-2, Cd-5, and Cd-6 were identified as Stenotrophomonas sp., Cd-3 as Achromobacter sp., and Cd-7 as Staphylococcus sp. The six strains showed a wide adaptation range for salinity and a strong tolerance to Cd2+. The effects of the initial Cd2+ concentration (1–100 mg/L), duration (18–72 h), temperature (10–40 °C), and pH (5.0–9.0) on the efficiency of Cd2+ removal were analysed. The results revealed that the Cd2+ removal rate was higher at an initial Cd2+ concentration of 5–100 mg/L than at 1 mg/L. The maximum Cd2+ removal effect was at a culture duration of 36 h, temperature of 10–35 °C, and pH of 5.0–7.0. X-ray diffraction (XRD) analysis revealed that the Cd2+ was immobilised by Stenotrophomonas sp. Cd-2 and Staphylococcus sp. Cd-7 through bio-precipitation. X-ray photoelectron spectroscopy (XPS) revealed that the Cd2+ was adsorbed by Stenotrophomonas sp. Cd-2, Achromobacter sp. Cd-3, and Staphylococcus sp. Cd-7. Fourier transform infrared spectroscopy (FTIR) analysis revealed that the isolates reacted with the Cd2+ mainly through the O–H, protein N–H, C–N, lipid C–H, fatty acid COO, polysaccharide C–O, P–O, and other functional groups, as well as with lipid molecules on the cell wall surfaces. Scanning electron microscopy (SEM) analysis revealed that there was little difference in the cells after Cd2+ treatment. The results of the soil remediation experiments indicated that the toxicity of Cd in soil could be effectively reduced using certain strains of microbe.
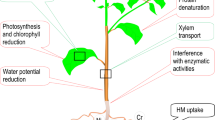
Graphical abstract







Similar content being viewed by others
Data availability
All data generated or analyzed are included in this paper.
References
Agarwal M, Rathore RS, Jagoe C, Chauhan A (2019) Multiple lines of evidences reveal mechanisms underpinning mercury resistance and volatilization by Stenotrophomonas sp. MA5 isolated from the Savannah River site (SRS), USA. Cells 8(4):309–323. https://doi.org/10.3390/cells8040309
Arivalagan P, Singaraj D, Haridass V, Kaliannan T (2014) Removal of cadmium from aqueous solution by batch studies using Bacillus cereus. Ecol Eng 71:728–735. https://doi.org/10.1016/j.ecoleng.2014.08.005
Bravo D, Braissant O (2022) Cadmium-tolerant bacteria: current trends and applications in agriculture. Lett Appl Microbiol 74:311–333. https://doi.org/10.1111/lam.13594
Chakravarty R, Banerjee PC (2012) Mechanism of cadmium binding on the cell wall of an acidophilic bacterium. Bioresour Technol 108:176–183. https://doi.org/10.1016/j.biortech.2011.12.100
Chien CC, Hung CW, Han CT (2010) Removal of cadmium ions during stationary growth phase by an extremely cadmium-resistant strain of Stenotrophomonas sp. Environ Toxicol Chem 26(4):664–668. https://doi.org/10.1897/06-280r.1
Choppala G, Bolan N, Kunhikrishnan A, Skinner W, Seshadri B (2015) Concomitant reduction and immobilization of chromium in relation to its bioavailability in soils. Environ Sci Pollut Res 22(12):8969–8978. https://doi.org/10.1007/s11356-013-1653-6
da Rocha Junior RB, Meira HM, Almeida DG, Rufino RD, Luna JM, Santos VA, Sarubbo LA (2019) Application of a low-cost biosurfactant in heavy metal remediation processes. Biodegradation 30(4):215–233. https://doi.org/10.1007/s10532-018-9833-1
Dong XZ, Cai MY, Lu YY, Xie JY, Liu XL (2001) Identification methods of common bacteria. In: Dong XZ, Cai MY (eds) Handbook of common bacteria systematic identify. Beijing Science press, Beijing, pp. 349–398
Faramarzi MA, Brandl H (2010) Formation of water-soluble metal cyanide complexes from solid minerals by Pseudomonas plecoglossicida. FEMS Microbiol Lett 259(1):47–52. https://doi.org/10.1111/j.1574-6968.2006.00245.x
Fozia A, Azra Y, Torsten T (2018) Essential gene clusters identified in Stenotrophomonas MB339 for multiple metal/antibiotic resistance and xenobiotic degradation. Curr Microbiol 75:1484–1492. https://doi.org/10.1007/s00284-018-1549-2
Green-Ruiz C, Rodriguez-Tirado V, Gomez-Gil B (2008) Cadmium and zinc removal from aqueous solutions by Bacillus jeotgali: pH, salinity and temperature effects. Bioresour Technol 99(9):3864–3870. https://doi.org/10.1016/j.biortech.2007.06.047
Gupta V, Rastogi A (2008) Equilibrium and kinetic modelling of cadmium (II) biosorption by nonliving algal biomass Oedogonium sp. from aqueous phase. J Hazard Mater 153(1–2):759–766. https://doi.org/10.1016/j.jhazmat.2007.09.021
Hlihor RM, Diaconu M, Leon F, Curteanu S, Tavares T, Gavrilescu M (2015) Experimental analysis and mathematical prediction of Cd (II) removal by biosorption using support vector machines and genetic algorithms. New Biotechnol 32(3):358–368. https://doi.org/10.1016/j.nbt.2014.08.003
Huang F, Guo CL, Lu GN, Yi XY, Zhu LD, Dang Z (2014) Bioaccumulation characterization of cadmium by growing Bacillus cereus RC-1 and its mechanism. Chemosphere 109:134–142. https://doi.org/10.1016/j.chemosphere.2014.01.066
Jiang Y, Zhang J, Wen Q, Zheng J, Zhang Y, Wei Q, Qin Y, Zhang X (2022) Up-flow anaerobic column reactor for sulfate-rich cadmium-bearing wastewater purification: system performance, removal mechanism and microbial community structure. Biodegradation 33(3):239–253. https://doi.org/10.1007/s10532-022-09983-0
Kartal S, Aydin Z, Tokalioglu S (2006) Fractionation of metals in street sediment samples by using the BCR sequential extraction procedure and multivariate statistical elucidation of the data. J Hazard Mater 132(1):80–89. https://doi.org/10.1016/j.jhazmat.2005.11.091
Khadivinia E, Sharafi H, Hadi F, Zahiri HS, Modiri S, Tohidi A, Mousavi A, Salmanian AH, Noghabi KA (2014) Cadmium biosorption by a glyphosate-degrading bacterium, a novel biosorbent isolated from pesticide-contaminated agricultural soils. J Ind Eng Chem 20(6):4304–4310. https://doi.org/10.1016/j.jiec.2014.01.037
Khan S, Lv J, Iqbal A, Fu P (2018) Morphophysiological and transcriptome analysis reveals a multiline defense system enabling cyanobacterium Leptolyngbya strain JSC-1 to withstand iron induced oxidative stress. Chemosphere 200:93–105. https://doi.org/10.1016/j.chemosphere.2018.02.100
Kim RY, Yoon JK, Kim TS, Yang JE, Owens G, Kim KR (2015a) Bioavailability of heavy metals in soils: definitions and practical implementation — a critical review. Environ Geochem Health 37(6):1041–1061. https://doi.org/10.1007/s10653-015-9695-y
Kim SY, Jin MR, Chung CH, Yun YS, Jahng KY, Yu KY (2015b) Biosorption of cationic basic dye and cadmium by the novel biosorbent Bacillus catenulatus JB-022 strain. J Biosci Bioeng 119(4):433–439. https://doi.org/10.1016/j.jbiosc.2014.09.022
Leong YK, Chang J-S (2020) Bioremediation of heavy metals using microalgae: recent advances and mechanisms. Bioresour Technol 303:122886. https://doi.org/10.1016/j.biortech.2020.122886
Li Y, Li A-M, Xu J, Li W-W, Yu H-Q (2013) Formation of soluble microbial products (SMP) by activated sludge at various salinities. Biodegradation 24(1):69–78. https://doi.org/10.1007/s10532-012-9558-5
Li W, Chen Y, Wang T (2021) Cadmium biosorption by lactic acid bacteria Weissella viridescens ZY-6. Food Control 123:107747. https://doi.org/10.1016/j.foodcont.2020.107747
Limcharoensuk T, Sooksawat N, Sumarnrote A, Awutpet T, Kruatrachue M, Pokethitiyook P, Auesukaree C (2015) Bioaccumulation and biosorption of Cd2+ and Zn2+ by bacteria isolated from a zinc mine in Thailand. Ecotoxicol Environ Saf 122:322–330. https://doi.org/10.1016/j.ecoenv.2015.08.013
Long J, Yu M, Xu H, Huang S, Wang Z, Zhang X-X (2021) Characterization of cadmium biosorption by inactive biomass of two cadmium-tolerant endophytic bacteria Microbacterium sp. D2–2 and Bacillus sp. C9–3. Ecotoxicology 30(7):1419–1428. https://doi.org/10.1007/s10646-021-02363-z
Luo H, Gu R, Ouyang H, Wang L, Shi S, Ji Y, Bao B, Liao G, Xu B (2021) Cadmium exposure induces osteoporosis through cellular senescence, associated with activation of NF-κB pathway and mitochondrial dysfunction. Environ Pollut 290:118043. https://doi.org/10.1016/j.envpol.2021.118043
Madmanang R, Sriwiriyarat T (2021) Effects of salinity on the respirometric activities of mixed culture bacteria from activated sludge process. J Phys: Conf Ser. https://doi.org/10.1088/1742-6596/1835/1/012104
Manasi RV, Santhana A, Kumar K, Rajesh N (2014) Biosorption of cadmium using a novel bacterium isolated from an electronic industry effluent. Chem Eng J 235(1):176–185. https://doi.org/10.1016/j.cej.2013.09.016
Masoudzadeh N, Zakeri F, Bagheri Lotfabad T, Sharafi H, Masoomi F, Zahiri HS, Ahmadian G, Noghabi KA (2011) Biosorption of cadmium by Brevundimonas sp. ZF12 strain, a novel biosorbent isolated from hot-spring waters in high background radiation areas. J Hazard Mater 197:190–198. https://doi.org/10.1016/j.jhazmat.2011.09.075
Masoumi F, Khadivinia E, Alidoust L, Mansourinejad Z, Shahryari S, Safaei M, Mousavi A, Salmanian A-H, Zahiri HS, Vali H (2016) Nickel and lead biosorption by Curtobacterium sp FM01, an indigenous bacterium isolated from farmland soils of northeast Iran. J Environ Chem Eng 4(1):950–957. https://doi.org/10.1016/j.jece.2015.12.025
Özdemir S, Kilinc E, Poli A, Nicolaus B, Güven K (2009) Biosorption of Cd, Cu, Ni, Mn and Zn from aqueous solutions by thermophilic bacteria, Geobacillus toebii sub. Sp. decanicus and Geobacillus thermoleovorans sub. sp. stromboliensis: equilibrium, kinetic and thermodynamic studies. Chem Eng J 152(1):195–206. https://doi.org/10.1016/j.cej.2009.04.041
Park JH, Chon H-T (2016) Characterization of cadmium biosorption by Exiguobacterium sp isolated from farmland soil near Cu-Pb-Zn mine. Environ Sci Pollut Res 23(12):11814–11822. https://doi.org/10.1007/s11356-016-6335-8
Rahman Z (2020) An overview on heavy metal resistant microorganisms for simultaneous treatment of multiple chemical pollutants at co-contaminated sites, and their multipurpose application. J Hazard Mater 396:122682. https://doi.org/10.1016/j.jhazmat.2020.122682
Ren LY, Hong ZN, Qian W, Li JY, Xu RK (2018) Adsorption mechanism of extracellular polymeric substances from two bacteria on Ultisol and Alfisol. Environ Pollut 237:39–49. https://doi.org/10.1016/j.envpol.2018.01.075
Shao W, Zhang K, Huo Y, Zhu J, Min L (2019) Pb biosorption by Leclercia adecarboxylata: Protective and immobilized mechanisms of extracellular polymeric substances. Chem Eng J 375:122113. https://doi.org/10.1016/j.cej.2019.122113
Subudhi S, Bisht V, Batta N, Pathak M, Devi A (2015) Purification and characterization of exopolysaccharide bioflocculant 1 produced by heavy metal 2 resistant Achromobacter xylosoxidans. Carbohydr Polym 137:441–451. https://doi.org/10.1016/j.carbpol.2015.10.066
Sun L, Zhang X, Ouyang W, Yang E, Sun R (2021) Lowered Cd toxicity, uptake and expression of metal transporter genes in maize plant by ACC deaminase-producing bacteria Achromobacter sp. J Hazard Mater 423:127036. https://doi.org/10.1016/j.jhazmat.2021.127036
Wang L, Li Y, Wang L, Zhu M, Zhu X, Qian C, Li W (2018) Responses of biofilm microorganisms from moving bed biofilm reactor to antibiotics exposure: protective role of extracellular polymeric substances. Bioresour Technol 254:268–277. https://doi.org/10.1016/j.biortech.2018.01.063
Wang X, Hu K, Xu Q, Lu L, Wang G (2020) Immobilization of Cd using mixed Enterobacter and Comamonas bacterial reagents in pot experiments with Brassica rapa L. Environ Sci Technol 54(24):15731–15741. https://doi.org/10.1021/acs.est.0c03114
Wu P, Rane NR, Xing C, Patil SM, Roh HS, Jeon BH, Li XF (2022) Integrative chemical and omics analyses reveal copper biosorption and tolerance mechanisms of Bacillus cereus strain T6. J Hazard Mater 435:129002. https://doi.org/10.1016/j.jhazmat.2022.129002
Yang X, Liu L, Tan W, Liu C, Qiu G (2020) Remediation of heavy metal contaminated soils by organic acid extraction and electrochemical adsorption. Environ Pollut 264:114745. https://doi.org/10.1016/j.envpol.2020.114745
Zakeri F, Noghabi KA, Sadeghizadeh M, Kardan MR, Masoomi F, Farshidpour MR, Atarilar A (2010) Serratia sp. ZF03: an efficient radium biosorbent isolated from hot-spring waters in high background radiation areas. Bioresour Technol 101(23):9163–9170. https://doi.org/10.1016/j.biortech.2010.07.032
Zeeshanur R, Thomas L, Singh VP (2019) Biosorption of heavy metals by a lead (Pb) resistant bacterium, Staphylococcus hominis strain AMB-2. J Basic Microbiol 59(5):477–486. https://doi.org/10.1002/jobm.201900024
Zouboulis A, Loukidou M, Matis K (2004) Biosorption of toxic metals from aqueous solutions by bacteria strains isolated from metal-polluted soils. Process Biochem 39(8):909–916. https://doi.org/10.1016/S0032-9592(03)00200-0
Funding
This work was financially supported by the Ningxia Natural Science Foundation (Excellent Youth Project) (No. 2021AAC05012), the National Natural Science Foundation of China (No. 21966001), and the Collaborative Heavy Metal Detection Project in Soil of Producing Area and Agricultural Products (2021–87).
Author information
Authors and Affiliations
Contributions
RF: conceptualization, methodology, resources, writing original–draft, writing–review & editing, supervision, project administration. WX: investigation, formal analysis, data curation. HM: validation, data curation. MZ: validation, visualization. KM: resources, writing–review & editing. XY: funding acquisition, writing–review & editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
Not applicable.
Consent for publication
Not applicable.
Consent to participate
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fan, R., Xie, W., Ma, H. et al. Isolation of cadmium-resistant microbial strains and their immobilisation of cadmium in soil. Biodegradation 34, 445–459 (2023). https://doi.org/10.1007/s10532-023-10026-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10532-023-10026-5