Abstract
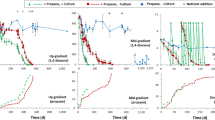
The objective of this research was to evaluate the potential for two gases, methane and ethane, to stimulate the biological degradation of 1,4-dioxane (1,4-D) in groundwater aquifers via aerobic cometabolism. Experiments with aquifer microcosms, enrichment cultures from aquifers, mesophilic pure cultures, and purified enzyme (soluble methane monooxygenase; sMMO) were conducted. During an aquifer microcosm study, ethane was observed to stimulate the aerobic biodegradation of 1,4-D. An ethane-oxidizing enrichment culture from these samples, and a pure culture capable of growing on ethane (Mycobacterium sphagni ENV482) that was isolated from a different aquifer also biodegraded 1,4-D. Unlike ethane, methane was not observed to appreciably stimulate the biodegradation of 1,4-D in aquifer microcosms or in methane-oxidizing mixed cultures enriched from two different aquifers. Three different pure cultures of mesophilic methanotrophs also did not degrade 1,4-D, although each rapidly oxidized 1,1,2-trichloroethene (TCE). Subsequent studies showed that 1,4-D is not a substrate for purified sMMO enzyme from Methylosinus trichosporium OB3b, at least not at the concentrations evaluated, which significantly exceeded those typically observed at contaminated sites. Thus, our data indicate that ethane, which is a common daughter product of the biotic or abiotic reductive dechlorination of chlorinated ethanes and ethenes, may serve as a substrate to enhance 1,4-D degradation in aquifers, particularly in zones where these products mix with aerobic groundwater. It may also be possible to stimulate 1,4-D biodegradation in an aerobic aquifer through addition of ethane gas. Conversely, our results suggest that methane may have limited importance in natural attenuation or for enhancing biodegradation of 1,4-D in groundwater environments.








Similar content being viewed by others
References
Adamson DT, Anderson RH, Mahendra S, Newell CW (2015) Evidence of 1,4-dioxane attenuation at groundwater sites contaminated with chlorinated solvents and 1,4-dioxane. Environ Sci Technol 49:6510–6518
Anderson RH, Anderson JK, Bower PA (2012) Co-occurrence of 1,4-dioxane with trichloroethylene in chlorinated solvent plumes at US Air Force installations: fact or fiction. Integr Environ Assess Manag 8:731–737
ATCC (2016) ATCC medium: 2157 Methylocella medium. https://atcc.org/~/media/01D07065549946AC863C486728B33349.ashx
Banerjee R (2013) Kinetic and spectroscopic characterization of intermediates in the soluble methane monooxygenase catalytic cycle: the old and the new, in biochemistry, molecular biology, and biophysics. University of Minnesota, Minneapolis
Banerjee R, Meier KK, Münck E, Lipscomb JD (2013) Intermediate P* from soluble methane monooxygenase contains a diferrous cluster. Biochemistry 52:4331–4342
Banerjee R, Proshlyakov Y, Lipscomb JD, Proshlyakov DA (2015) Structure of the key species in the enzymatic oxidation of methane to methanol. Nature 518:431–434
Beauvais LG, Lippard SJ (2005) Reactions of the peroxo intermediate of soluble methane monooxygenase hydroxylase with ethers. J Am Chem Soc 127:7370–7378
Bernhardt D, Diekmann H (1991) Degradation of dioxane, tetrahydrofuran and other cyclic ethers by an environmental Rhodococcus strain. Appl Microbiol Biotechnol 36:120–123
Brockman FJ, Payne W, Workman DJ, Soong A, Manley S, Hazen TC (1995) Effect of gaseous nitrogen and phosphorus injection on in istu bioremediation of a trichloroethylene-contaminated site. J Hazard Mat 41:287–298
Chiang S-Y, Mora DR, Diguiseppi WH, Davis G, Sublette K, Gedalanga P, Mahendra S (2012) Characterizing the intrinsic bioremediation potential of 1,4-dioxane and trichloroethene using innovative environmental diagnostic tools. J Environ Monit 14:2317–2326
Chu KH, Alvarez-Cohen L (1999) Evaluation of toxic effects of aeration and trichloroethylene oxidation on methanotrophic bacteria grown with different nitrogen sources. Appl Environ Microbiol 65:766–772
Crombie AT, Murrell JC (2014) Trace-gas metabolic versatility of the facultative methanotroph Methylocella silvestris. Nature 510:148–151
De Bruin WP, Koterman MJ, Posthumus MA, Schraa G, Zehnder AJ (1992) Complete biological reductive transformation of tetrachloroethene to ethane. Appl Environ Microbiol 58:1996–2000
Dedysh SN, Panikov NS, Tiedje JM (1998a) Acidophilic methanotrophic communities from Sphagnum peat bogs. Appl Environ Microbiol 64:922–929
Dedysh SN, Panikov NS, Liesack W, Großkopf R, Zhou J, Tiedje JM (1998b) Isolation of acidophilic methane-oxidizing bacteria from northern peat wetlands. Science 282:281–284
Dedysh SN, Liesack W, Khmelenina VN, Suzina NE, Trotsenko YA, Semrau JD, Bares AM, Panikov NS, Tiedje JM (2000) Methylocella palustris gen nov., a new methane-oxidizing acidophilic bacterium from peat bogs, representing a novel subtype of serine pathway methanotrophs. Int J Syst Evol Microbiol 50:955–969
DiStefano TD, Gossett JM, Zinder SH (1991) Reductive dechlorination of high concentrations of tetrachloroethene to ethene by an anaerobic enrichment culture in the absence of methanogenesis. Appl Environ Microbiol 57:2287–2292
EPA; US Environmental Protection Agency (2009) Drinking Water Contaminant Candidate List 3: Final. Federal Register Notice. https://www.federalregister.gov/documents/2009/10/08/E9-24287/drinking-water-contaminant-candidate-list-3-final
EPA; US Environmental Protection Agency (2014) Technical Fact Sheet: 1,4-Dioxane. United States Environmental Protection Agency,Office of Solid Waste and Emergency Response. EPA 505-F-14-011. https://www.epa.gov/sites/production/files/2014-03/documents/ffrro_factsheet_contaminant_14-dioxane_january2014_final.pdf
Foster JW, Davis RH (1966) A methane-dependent coccus, with notes on classification and nomenclature or obligate, methane-utilizing bacteria. J Bacteriol 91:1924–1931
Fox BG, Borneman JG, Wackett LP, Lipscomb JD (1990a) Haloalkene oxidation by the soluble methane monooxygenase from Methylosinus trichosporium OB3b: mechanistic and environmental implications. Biochemistry 29:6419–6427
Fox BG, Froland WA, Jollie DR, Lipscomb JD (1990b) Methane monooxygenase from Methylosinus trichosporium OB3b. Methods Enzymol 188:191–202
Freedman DL, Gossett JM (1989) Biological reductive dechlorination of tetrachloroethylene and trichloroethylene to ethylene under methanogenic conditions. Appl Environ Microbiol 55:2144–2151
Freedman DL, Herz SD (1996) Use of ethylene and ethane as primary substrates for aerobic cometabolism of vinyl chloride. Water Environ Res 68:320–328
Gedalanga PB, Pornwongthong P, Mora R, Chiang S-YD, Baldwin B, Ogles D, Mahendra S (2014) Identification of biomarker genes to predict biodegradation of 1,4-dioxane. Appl Environ Microbiol 80:3209–3218
Gedalanga P, Madison A, Miao Y, Richards T, Hatton J, DiGuiseppi WH, Wilson J, Mahendra S (2016) A multiple lines of evidence framework to evaluate intrinsic biodegradation of 1,4-dioxane. Remediat J 27(1):93–114
Green J, Dalton H (1989) Substrate specificity of soluble methane monooxygenase. J Biol Chem 264:17698–17703
Grostern A, Alvarez-Cohen L (2013) RubisCO-based CO2 fixation and C1 metabolism in the actinobacterium Pseudonocardia dioxanivorans CB1190. Environ Microbiol 15:3040–3053
Hanson RS, Hanson TE (1996) Methanotrophic bacteria. Microbiol Rev 60:439–471
Hareland WA, Crawford RL, Chapman PJ, Dagley S (1975) Metabolic function and properties of 4-hydroxyphenylacetic acid 1-hydroxylase from Pseudomonas acidovorans. J Bacteriol 121:272–285
Hatzinger PB, Streger SH, Begley JF (2015) Enhancing aerobic biodegradation of 1,2-dibromoethane in groundwater using ethane or propane and inorganic nutrients. J Contam Hydrol 172:61–70
Kim Y-M, Jeon J-R, Murugesan K, Kim E-J, Chang Y-S (2009) Biodegradation of 1,4-dioxane and transformation of related cyclic compounds by a newly isolated Mycobacterium sp. PH-06. Biodegradation 20:511–519
Koh S-C, Bowman JP, Sayler GS (1993) Soluble methane monooxygenase production and trichloroethylene degradation by a type I methanotroph, Methylomonas methanica 68-1. Appl Environ Microbiol 59:960–967
Lee S-W, Keeney DR, Lim D-H, DiSpirito AA, Semrau JD (2006) Mixed pollutant degradation by Methylosinus trichosporium OB3b expressing either soluble or particulate methane monooxygenase: can the tortoise beat the hare? Appl Environ Microbiol 72:7503–7509
Li M, Mathieu J, Yang Y, Fiorenza S, Deng Y, He Z, Zhou J, Alvarez PJJ (2013) Widespread distribution of soluble di-iron monooxygenase (SDIMO) genes in arctic groundwater impacted by 1,4-dioxane. Environ Sci Technol 47:9950–9958
Li M, Liu Y, Mathieu J, Alvarez PJJ (2017) Comparison of 1,4-dioxane cometabolism with the amendment of different alkane gases. Presentation at the fourth International Symposium on Bioremediation and Sustainable Environmental Technologies. Miami, FL, May 22–25
Lippincott D, Streger SH, Schaefer CE, Hinkle J, Stormo J, Steffan RJ (2015) Bioaugmentation and propane biosparging for in situ biodegradation of 1,4-dioxane. Groundw Monit Remediat 35:81–92
Lontoh S, Semrau JD (1998) Methane and trichloroethylene degradation by Methylosinus trichosporium OB3b expressing particulate methane monooxygenase. Appl Environ Microbiol 64:1106–1114
Mahendra S, Alvarez-Cohen L (2005) Pseudonocardia dioxanivorans sp. nov., a novel actinomycete that grows on 1,4-dioxane. Int J Syst Evol Microbiol 55:593–598
Mahendra S, Alvarez-Cohen L (2006) Kinetics of 1,4-dioxane biodegradation by monooxygenase-expressing bacteria. Environ Sci Technol 40:5435–5442
Mahendra S, Petzold CJ, Baidoo EE, Keasling JD, Alvarez-Cohen L (2007) Identification of the intermediates of in vivo oxidation of 1,4-dioxane by monooxygenase-containing bacteria. Environ Sci Technol 41:7330–7336
Mahendra S, Grostern A, Alvarez-Cohen L (2013) The impact of chlorinated solvent co-contaminants on the biodegradation kinetics of 1,4-dioxane. Chemosphere 91:88–92
Masuda H, McClay K, Steffan RJ, Zylstra GL (2012) Biodegradation of tetrahydrofuran and 1,4-dioxane by soluble diiron monooxygenase in Pseudonocardia sp. Strain ENV478. J Mol Microbiol Biotechnol 22:312–316
Maymó-Gatell X, Chien Y-T, Gossett JM, Zinder SH (1997) Isolation of a bacterium that reductively dechlorinates tetrachloroethene to ethene. Science 276:1568–1571
Mohr T, Stickney J, DiGuiseppi W (2010) Environmental investigation and remediation: 1,4-dioxane and other solvent stabilizers. CRC Press, Boca Raton, p 552
Nakamiya K, Hashimoto S, Ito H, Edmonds JS, Morita M (2010) Degradation of 1,4-dioxane and cyclic ethers by an isolated fungus. Appl Environ Microbiol 71:1254–1258
Orth WS, Gilham RW (1995) Dechlorination of trichloroethene in aqueous solution using Fe0. Environ Sci Technol 30:66–71
Parales RE, Adamus JE, White N, May HD (1994) Degradation of 1,4-dioxane by an actinomycete in pure culture. Appl Environ Microbiol 60:4527–4530
Puls RW, Barcelona MJ (1995) Low-flow (minimal drawdown) ground-water sampling procedures. United States Environmental Protection Agency EPA/540/S-95/504. https://www.researchgate.net/profile/Michael_Barcelona/publication/237506331_United_States_Environmental_Protection_Agency_Office_of_Research_and_Development_Office_of_Solid_Waste_and_Emergency_Response_EPA540S-95504_December_1995_EPA_Ground_Water_Issue_LOW-FLOW_MINIMAL_DRAWDO/links/0a85e52ef9573672af000000.pdf
Sadeghi V, Mora R, Jacob P, Chiang S-H (2016) Characterizing in situ methane-enhanced biostimulation potential for 1,4-dioxane biodegradation in groundwater. Remediat J 27(1):115–132
Sales CM, Mahendra S, Grostern A, Parales RE, Goodwin LA, Woyke T, Nolan M, Lapidus A, Chertkov O, Ovchinnikova G, Sczyrba A, Alvarez-Cohen L (2011) Genome sequence of the 1,4-dioxane-degrading Pseudonocardia dioxanivorans Strain CB1190. J Bacteriol 193:4549–4550
Sei K, Miyagaki K, Kakinoki T, Fukugasako K, Inoue D, Michihiko I (2013) Isolation and characterization of bacterial strains that have high ability to degrade 1,4-dioxane as a sole carbon and energy source. Biodegradation 24:665–674
Semrau J (2011) Bioremediation via methanotrophy: overview of recent findings and suggestions for future research. Front Microbiol 2:209–219
Stroo HF, Ward CH (2008) In Situ Bioremediatiion of Perchlorate in Groundwater. Springer Science/Business Media, LLC, New York
Tinberg CE, Lippard SJ (2010) Oxidation reactions performed by soluble methane monooxygenase hydroxylase intermediates Hperoxo and Q proceed by distinct mechanisms. Biochemistry 49:7902–7912
Vainberg S, McClay K, Masuda H, Root D, Condee C, Zylstra GL, Steffan RJ (2006) Biodegradation of ether pollutants by Pseudonocardia sp. strain ENV478. Appl Environ Microbiol 72:5218–5224
Verce MF, Freedman DL (2000) Modeling the biodegradation kinetics of vinyl chloride cometabolism by an ethane-grown Pseudomonas sp. Biotechnol Bioeng 71:274–285
Wymore RA, Lee MH, Keener WK, Miller AR, Colwell FS, Watwood ME, Sorenson KS Jr (2007) Field evidence for intrinsic aerobic chlorinated ethene cometabolism by methanotrophs expressing soluble methane monooxygenase. Biorem J 11:125–139
Zenker MJ, Borden RC, Barlaz MA (2000) Mineralization of 1,4-dioxane in the presence of a structural analog. Biodegradation 11:239–246
Zhang J, Lipscomb JD (2006) Role of the C-terminal region of the B component of Methylosinus trichosporium OB3b methane monooxygenase in the regulation of oxygen activation. Biochemistry 45:1459–1469
Acknowledgements
We wish to acknowledge the Strategic Environmental Research and Development Program (SERDP; Project ER-2306) and the Air Force Civil Engineer Center (AFCEC; BAA Project 769) for supporting some of the studies described herein. Dr Michael Hyman at North Carolina State University provided helpful comments and insights. We also wish to thank Dr. Randi Rothmel and Anthony Soto of CB&I for their analytical support. The results and interpretations presented herein are solely the opinion of the authors and not of SERDP or AFCEC unless otherwise stated in official documentation.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hatzinger, P.B., Banerjee, R., Rezes, R. et al. Potential for cometabolic biodegradation of 1,4-dioxane in aquifers with methane or ethane as primary substrates. Biodegradation 28, 453–468 (2017). https://doi.org/10.1007/s10532-017-9808-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10532-017-9808-7