Abstract
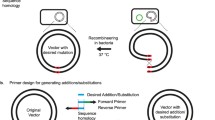
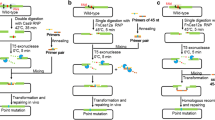
Site-directed and saturation mutagenesis are critical DNA methodologies for studying protein structure and function. For plasmid-based gene mutation, PCR and overlap-extension PCR involve tedious cloning steps. When the plasmid size is large, PCR yield may be too low for cloning; and for saturation mutagenesis of a single codon, one experiment may not enough to generate all twenty coding variants. Oligo-mediated recombineering sidesteps the complicated cloning process by homologous recombination between a mutagenic oligo and its target site. However, the low recombineering efficiency and inability to select for the recombinant makes it necessary to screen a large number of clones. Herein, we describe two plasmid-based mutagenic strategies: CRISPR/Cas9-assisted ssDNA recombineering for site-directed mutagenesis (CRM) and saturation mutagenesis (CRSM). CRM and CRSM involve co-electroporation of target plasmid, sgRNA expression plasmid and mutagenic oligonucleotide into Escherichia coli cells with induced expression of λ-Red recombinase and Cas9, followed by plasmid extraction and characterization. We established CRM and CRSM via ampicillin resistance gene repair and mutagenesis of N-acetyl‑D‑neuraminic acid aldolase. The mutational efficiency was between 80 and 100% and all twenty amino acid coding variants were obtained at a target site via a single CRSM strategy. CRM and CRSM have the potential to be general plasmid-based gene mutagenesis tools.


Similar content being viewed by others
References
Campeotto I, Bolt AH, Harman TA, Dennis C, Trinh CH, Phillips SE, Nelson A, Pearson AR, Berry A (2010) Structural insights into substrate specificity in variants of N-acetylneuraminic acid lyase produced by directed evolution. J Mol Biol 404:56–69
Datsenko KA, Wanner BL (2000) One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci U S A 97:6640–6645
Dong M, Wang F, Li Q, Han R, Li A, Zhai C, Ma L (2020) A single digestion, single-stranded oligonucleotide mediated PCR-independent site-directed mutagenesis method. Appl Microbiol Biotechnol 104:3993–4003
Ellis HM, Yu D, DiTizio T, Court DL (2001) High efficiency mutagenesis, repair, and engineering of chromosomal DNA using single-stranded oligonucleotides. Proc Natl Acad Sci U S A 98:6742–6746
Gao X, Zhang F, Wu M, Wu Z, Shang G (2019) Production of N-acetyl-D-neuraminic acid by whole cells expressing Bacteroides thetaiotaomicron N-acetyl-D-glucosamine 2-epimerase and Escherichia coli N-acetyl-D-neuraminic acid aldolase. J Agric Food Chem 67:6285–6291
Garst AD, Bassalo MC, Pines G, Lynch SA et al (2017) Genome-wide mapping of mutations at single-nucleotide resolution for protein, metabolic and genome engineering. Nat Biotechnol 35:48–55
Graf N, Altenbuchner J (2013) Functional characterization and application of a tightly regulated MekR/PmekA expression system in Escherichia coli and Pseudomonas putida. Appl Microbiol Biotechnol 97:8239–8251
Halperin SO, Tou CJ, Wong EB, Modavi C, Schaffer DV, Dueber JE (2018) CRISPR-guided DNA polymerases enable diversification of all nucleotides in a tunable window. Nature 560:248–252
Horton RM, Hunt HD, Ho SN, Pullen JK, Pease LR (1989) Engineering hybrid genes without the use of restriction enzymes: gene splicing by overlap extension. Gene 77:61–68
Huffman MA, Fryszkowska A, Alvizo O, Borra-Garske M et al (2019) Design of an in vitro biocatalytic cascade for the manufacture of islatravir. Science 366:1255–1259
Jiang W, Bikard D, Cox D, Zhang F, Marraffini LA (2013) RNA-guided editing of bacterial genomes using CRISPR-Cas systems. Nat Biotechnol 31:233–239
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E (2012) A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337:816–821
Li J, Sun J, Gao X, Wu Z, Shang G (2019) Coupling ssDNA recombineering with CRISPR-Cas9 for Escherichia coli DnaG mutations. Appl Microbiol Biotechnol 103:3559–3570
Miyazaki K, Arnold FH (1999) Exploring nonnatural evolutionary pathways by saturation mutagenesis: rapid improvement of protein function. J Mol Evol 49:716–720
Murphy KC (2016) λ Recombination and Recombineering. EcoSal Plus. https://doi.org/10.1128/ecosalplus.ESP-0011-2015
Oh JH, van Pijkeren JP (2014) CRISPR-Cas9-assisted recombineering in Lactobacillus reuteri. Nucleic Acids Res 42:e131
Posfai G, Kolisnychenko V, Bereczki Z, Blattner FR (1999) Markerless gene replacement in Escherichia coli stimulated by a double-strand break in the chromosome. Nucleic Acids Res 27:4409–4415
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory, Cold Spring Harbor
Sawitzke JA, Costantino N, Li XT, Thomason LC, Bubunenko M, Court C, Court DL (2011) Probing cellular processes with oligo-mediated recombination and using the knowledge gained to optimize recombineering. J Mol Biol 407:45–59
She W, Ni J, Shui K, Wang F, He R, Xue J, Reetz MT, Li A, Ma L (2018) Rapid and error-free site-directed mutagenesis by a PCR-free in vitro CRISPR/Cas9-mediated mutagenic system. ACS Synth Biol 7:2236–2244
Slaymaker IM, Gao L, Zetsche B, Scott DA, Yan WX, Zhang F (2016) Rationally engineered Cas9 nucleases with improved specificity. Science 351:84–88
Song J, Dong H, Ma C, Zhao B, Shang G (2010) Construction and functional characterization of an integrative form λ Red recombineering Escherichia coli strain. FEMS Microbiol Lett 309:178–183
Thomason LC, Costantino N, Shaw DV, Court DL (2007) Multicopy plasmid modification with phage λ Red recombineering. Plasmid 58:148–158
von Itzstein M (2007) The war against influenza: discovery and development of sialidase inhibitors. Nat Rev Drug Discov 6:967–974
Zhang Q, Yan Z, Xu Y, Sun J, Shang G (2017) Characterization of inducible ccdB gene as a counterselectable marker in Escherichia coli recombineering. Curr Microbiol 74:961–964
Zhang JJ, Tang X, Huan T, Ross AC, Moore BS (2020) Pass-back chain extension expands multimodular assembly line biosynthesis. Nat Chem Biol 16:42–49
Acknowledgements
We thank for Dr. Frederick Blattner, Dr. Josef Altenbuchner, Dr. John Cronan, and Dr. Barry Wanner for providing the plasmids used in this research. We thank the two anonymous reviewers for their critical comment and insightful contributions to the article.
Funding
Financial support was provided by National Natural Science Foundation of China (NSFC 82073742).
Author information
Authors and Affiliations
Contributions
GS conceptualized and designed the study, supervised the research and finalized the manuscript. GZ, JW and YL carried out the experiments, participated in data analysis and manuscript preparation. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors report no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, G., Wang, J., Li, Y. et al. CRISPR/Cas9-assisted ssDNA recombineering for site-directed mutagenesis and saturation mutagenesis of plasmid-encoded genes. Biotechnol Lett 45, 629–637 (2023). https://doi.org/10.1007/s10529-023-03363-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-023-03363-1