Abstract
Objective
To select potential ligands of ALS3 for drug development with anti-adhesion and/or anti-biofilm activities.
Methodology
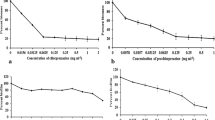
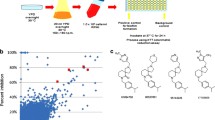
ALS3 model was considered stable by DM. The main features of protein flexibility were represented by two conformers which were used in the virtual screening. Twenty-four small molecules were selected for in vitro assays. Five of them presented the best biological activity with ability to inhibit the adhesion and C. albicans biofilm formation on abiotic surface.
Results
To select potential ligands of ALS3 for drug development with anti-adhesion and/or anti-biofilm activities.
Conclusion
In silico tools application was able to select promising compounds with anti-adhesion activity, opening a new perspective of medical device treatment.




Similar content being viewed by others
References
Abadio AK, Kioshima ES, Teixeira MM, Martins NF, Maigret B, Felipe MS (2011) Comparative genomics allowed the identification of drug targets against human fungal pathogens. BMC Genom 27(12):75. https://doi.org/10.1186/1471-2164-12-75
Abadio AKR, Kioshima ES, Leroux V, Martins NF, Maigret B, Felipe MS (2015) Identification of new antifungal compounds targeting thioredoxin reductase of Paracoccidioides genus. PLoS ONE 10(11):e0142926. https://doi.org/10.1371/journal.pone.0142926
Aoki W, Kitahara N, Miura N, Morisaka H, Kuroda K, Ueda M (2012) Profiling of adhesive properties of the agglutinin-like sequence (ALS) protein family, a virulent attribute of Candida albicans. FEMS Immunol Med Microbiol 65(1):121–124. https://doi.org/10.1111/j.1574-695X.2012.00941.x
Basmaciyan L, Bon F, Paradis T, Lapaquette P, Dalle F (2019) Candida albicans interactions with the host: crossing the intestinal epithelial barrier. Tissue Barriers. https://doi.org/10.1080/21688370.2019.1612661
Beautrait A, Leroux V, Chavent M, Ghemtio L, Devignes MD, Smaïl-Tabbone M, Cai W, Shao X, Moreau G, Bladon P, Yao J, Maigret B (2008) Multiple-step virtual screening using VSM-G: overview and validation of fast geometrical matching enrichment. J Mol Model 14(2):135–148. https://doi.org/10.1007/s00894-007-0257-9
Chang YL, Yu SJ, Heitman J, Wellington M, Chen YL (2017) New facets of antifungal therapy. Virulence 8(2):222–236. https://doi.org/10.1080/21505594.2016.1257457
de Amorim AL, de Lima AVM, Rosário ACAD, Souza ÉTDS, Ferreira JV, Hage-Melim LIDS (2018) Molecular modeling of inhibitors against fructose bisphosphate aldolase from Candida albicans. Silico Pharmacol 6(1):2. https://doi.org/10.1007/s40203-018-0040-x
Di Pasquale N, Gowers RJ, Carbone P (2014) A multiple time step molecular dynamics algorithm for macromolecules. J Comput Chem 35(16):1199–1207. https://doi.org/10.1002/jcc.23594
Dong G, Liu Y, Wu Y, Tu J, Chen S, Liu N, Sheng C (2018) Novel non-peptidic small molecule inhibitors of secreted aspartic protease 2 (SAP2) for the treatment of resistant fungal infections. Chem Commun 54(96):13535–13538. https://doi.org/10.1039/c8cc07810f
Hosseini SS, Ghaemi E, Noroozi A, Niknejad F (2019) Zinc oxide nanoparticles inhibition of initial adhesion and ALS1 and ALS3 gene expression in Candida albicans strains from urinary tract infections. Mycopathologia 184(2):261–271. https://doi.org/10.1007/s11046-019-00327-w
Hoyer LL, Cota E (2016) Candida albicans agglutinin-like sequence (ALS) family vignettes: a review of als protein structure and function. Front Microbiol 7:280. https://doi.org/10.3389/fmicb.2016.00280
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14(1):33–38
Jiang W, Phillips JC, Huang L, Fajer M, Meng Y, Gumbart JC, Luo Y, Schulten K, Roux B (2014) Generalized scalable multiple copy algorithms for molecular dynamics simulations in NAMD. Comput Phys Commun 185(3):908–916. https://doi.org/10.1016/j.cpc.2013.12.014
Jones G, Willett P, Glen RC, Leach AR, Taylor R (1997) Development and validation of a genetic algorithm for flexible docking. J Mol Biol 267:727–748
Khan I, Kanugala S, Shareef MA, Ganapathi T, Shaik AB, Shekar KC, Kamal A, Kumar CG (2019) Synthesis of new bis-pyrazole linked hydrazides and their in vitro evaluation as antimicrobial and anti-biofilm agents: a mechanistic role on ergosterol biosynthesis inhibition in Candida albicans. Chem Biol Drug Des. https://doi.org/10.1111/cbdd.13509
Kullberg BJ, Arendrup MC (2015) Invasive candidiasis. N Engl J Med 373(15):1445–1456. https://doi.org/10.1056/NEJMra1315399
Lamoth F, Lockhart SR, Berkow EL, Calandra T (2018) Changes in the epidemiological landscape of invasive candidiasis. J Antimicrob Chemother 73(Suppl 1):i4–i13. https://doi.org/10.1093/jac/dkx444
Li WS, Chen YC, Kuo SF, Chen FJ, Lee CH (2018) The impact of biofilm formation on the persistence of candidemia. Front Microbiol. 9:1196. https://doi.org/10.3389/fmicb.2018.01196
Lin J, Oh SH, Jones R, Garnett JA, Salgado PS, Rusnakova S, Matthews SJ, Hoyer LL, Cota E (2014) The peptide-binding cavity is essential for Als3-mediated adhesion of Candida albicans to human cells. J Biol Chem 289(26):18401–18412. https://doi.org/10.1074/jbc.M114.547877
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (2001) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 46(1–3):3–26
Liu Y, Filler SG (2011) Candida albicans Als3, a multifunctional adhesin and invasin. Eukaryot Cell 10(2):168–173. https://doi.org/10.1128/EC.00279-10
Marc G, Araniciu C, Oniga S, Vlase L, Pîrnău A, Duma M, Măruescu L, Chifiriuc MC, Oniga O (2018) New N-(oxazolylmethyl)-thiazolidinedione active against Candida albicans biofilm: potential Als proteins inhibitors. Molecules 23(10):2522. https://doi.org/10.3390/molecules23102522
Nobile CJ, Fox EP, Nett JE, Sorrells TR, Mitrovich QM, Hernday AD, Tuch BB, Andes DR, Johnson AD (2012) A recently evolved transcriptional network controls biofilm development in Candida albicans. Cell 148(1–2):126–138. https://doi.org/10.1016/j.cell.2011.10.048
Perlin DS, Rautemaa-Richardson R, Alastruey-Izquierdo A (2017) The global problem of antifungal resistance: prevalence, mechanisms, and management. Lancet Infect Dis 17:e383–e392. https://doi.org/10.1016/S1473-3099(17)30316-X
Pfaller MA, Diekema DJ, Turnidge JD, Castanheira M, Jones RN (2019) Twenty years of the SENTRY antifungal surveillance program: results for Candida species from 1997–2016. Open Forum Infect Dis 6(Suppl 1):S79–S94. https://doi.org/10.1093/ofid/ofy358
Phua AI, Hon KY, Holt A, O'Callaghan M, Bihari S (2019) Candida catheter-related bloodstream infection in patients on home parenteral nutrition: rates, risk factors, outcomes, and management. Clin Nutr ESPEN 31:1–9. https://doi.org/10.1016/j.clnesp.2019.03.007
Quindós G, Marcos-Arias C, San-Millán R, Mateo E, Eraso E (2018) The continuous changes in the aetiology and epidemiology of invasive candidiasis: from familiar Candida albicans to multi resistant Candida auris. Int Microbiol 21(3):107–119. https://doi.org/10.1007/s10123-018-0014-1
Rodrigues-Vendramini FAV, Faria DR, Arita GS, Capoci IRG, Sakita KM, Caparroz-Assef SM, Becker TCA, Bonfim-Mendonça PS, Felipe M, Svidzinski TIE, Maigret B, Kioshima ES (2019) Antifungal activity of two oxadiazole compounds for the paracoccidioidomycosis treatment. PLOS Negl Trop Dis 13(6):e0007441. https://doi.org/10.1371/journal.pntd.0007441
Rozada AM, Rodrigues FA, Sampiron EG, Seixas FA, Basso EA, Scodro RB, Kioshima ES, Gauze GF (2019) Novel 4-methoxynaphthalene-N-acylhydrazones as potential for paracoccidioidomycosis and tuberculosis co-infection. Future Microbiol 14:587–598. https://doi.org/10.2217/fmb-2018-0357
Salci TP, Negri M, Abadio AKR, Bonfim-Mendonça P, Capoci I, Caparroz-Assef SM, Donatti L, Felipe MS, Kioshima ES, Svidzinski TI (2017) A new small-molecule KRE2 inhibitor against invasive Candida parapsilosis infection. Future Microbiol 12:1283–1295. https://doi.org/10.2217/fmb-2017-0065
Salgado PS, Yan R, Taylor JD, Jones R, Hoyer LL, Matthews SJ, Simpson PJ, Cota E (2011) Structural basis for the broad specificity to host-cell ligands by the pathogenic fungus Candida albicans. Proc Natl Acad Sci USA 108(38):15775–15779. https://doi.org/10.1073/pnas.1103496108
Semenyuta IV, Kobzar OL, Hodyna DM, Brovarets VS, Metelytsia LO (2019) In silico study of 4-phosphorylated derivatives of 1,3-oxazole as inhibitors of Candida albicans fructose-1,6-bisphosphate aldolase II. Heliyon 5(4):e01462. https://doi.org/10.1016/j.heliyon.2019.e01462
Shao J, Tanner SW, Thompson N, Cheatham TE (2007) Clustering molecular dynamics trajectories: 1. Characterizing the performance of different clustering algorithms. J Chem Theory Comput. 3(6):2312–2334. https://doi.org/10.1021/ct700119m
Silva DR, Sardi JCO, Freires IA, Silva ACB, Rosalen PL (2019) In silico approaches for screening molecular targets in Candida albicans: a proteomic insight into drug discovery and development. Eur J Pharmacol 5(842):64–69. https://doi.org/10.1016/j.ejphar.2018.10.016
Stierand K, Maass PC, Rarey M (2006) Molecular complexes at a glance: automated generation of two-dimensional complex diagrams. Bioinformatics 22(14):1710–1716. https://doi.org/10.1093/bioinformatics/btl150
Tascini C, Sozio E, Corte L, Sbrana F, Scarparo C, Ripoli A, Bertolino G, Merelli M, Tagliaferri E, Corcione A, Bassetti M, Cardinali G, Menichetti F (2018) The role of biofilm forming on mortality in patients with candidemia: a study derived from real world data. Infect Dis 50(3):214–219. https://doi.org/10.1080/23744235.2017.1384956
Wall G, Montelongo-Jauregui D, Vidal Bonifacio B, Lopez-Ribot JL, Uppuluri P (2019) Candida albicans biofilm growth and dispersal: contributions to pathogenesis. Curr Opin Microbiol 11(52):1–6. https://doi.org/10.1016/j.mib.2019.04.001
Webb BJ, Ferraro JP, Rea S, Kaufusi S, Goodman BE, Spalding J (2018) Epidemiology and clinical features of invasive fungal infection in a US Health Care Network. Open Forum Infect Dis 5(8):187. https://doi.org/10.1093/ofid/ofy187
Wilson D, Naglik JR, Hube B (2016) The missing link between Candida albicans hyphal morphogenesis and host cell damage. PLoS Pathog 12(10):e1005867. https://doi.org/10.1371/journal.ppat.1005867
Zarnowski R, Sanchez H, Covelli AS, Dominguez E, Jaromin A, Bernhardt J, Mitchell KF, Heiss C, Azadi P, Mitchell A, Andes DR (2018) Candida albicans biofilm-induced vesicles confer drug resistance through matrix biogenesis. PLoS Biol 16(10):e2006872. https://doi.org/10.1371/journal.pbio.2006872
Zhao X, Daniels KJ, Oh SH, Green CB, Yeater KM, Soll DR, Hoyer LL (2006) Candida albicans Als3p is required for wild-type biofilm formation on silicone elastomer surfaces. Microbiology 152(Pt 8):2287–2299. https://doi.org/10.1099/mic.0.28959-0
Supporting information
Supplementary Fig. 1
Funding
This study was supported by Conselho Nacional de Desenvolvimento Científico e Tecnológico [Grant: 236764/2012-8] and Coordenação de Aperfeiçoamento de Pessoal de Nível Superior, according to CNPq/casadinho/PROCAD [Grant: 552276/2011-1].
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors have no other relevant affiliations or financial involvement with any organization or entity with a financial interest in or financial conflict with the subject matter or materials discussed in the manuscript apart from those disclosed.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kioshima, E.S., Shinobu-Mesquita, C.S., Abadio, A.K.R. et al. Selection of potential anti-adhesion drugs by in silico approaches targeted to ALS3 from Candida albicans. Biotechnol Lett 41, 1391–1401 (2019). https://doi.org/10.1007/s10529-019-02747-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-019-02747-6