Abstract
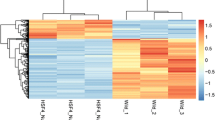
Glucocorticoid-induced cataract (GIC)-associated biomarkers were screened by ceRNA network construction. The GIC samples’ GSE3040 were obtained from the NCBI-GEO database. R’s Limma package was used to identify differentially expressed RNAs (DERs) between the normal and GIC samples group (4- and 16-h). The Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways enrichment analysis for the mRNAs in the constructed GIC lncRNA–miRNA–mRNA ceRNA regulation network was implemented. A total of 1665 and 1443 DERs were obtained in the 4- and 16-h group, respectively. At two time points, 256 overlapping DERs were identified, of which 210 (17 lncRNAs and 203 mRNAs) had significant differential expression (4 down- and 206 up-regulated). A total of 534 co-expressed ligation pairs (all up-regulated) were obtained. A ceRNA regulation network was constructed. RPS6KA5, GAB1, CCR7, CCL2, COL4A4, and PPARG were obtained and significantly enriched in the 4 KEGG signaling pathways and were featured as GIC target molecules.





Similar content being viewed by others
Availability of Data and Material
The datasets generated during and/or analyzed during the current study are available in the [GEO] repository, [https://www.ncbi.nlm.nih.gov/)].
Abbreviations
- GIC:
-
Glucocorticoid-induced cataract
- DERs:
-
Differentially expressed RNAs
- PCC:
-
Pearson correlation coefficient
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- ncRNAs:
-
Non-coding RNAs
- lncRNAs:
-
Long non-coding RNAs
- DEX:
-
Dexamethasone
- HGNC:
-
HUGO Gene Nomenclature Committee
- GO:
-
Gene Ontology
- ceRNA:
-
Competitive endogenous RNA
- GC:
-
Glucocorticoids
References
Badia A, Salas A, Duarri A, Ferreira-de-Souza B, Zapata M, Fontrodona L, García-Arumí J (2021) Transcriptomics analysis of Ccl2/Cx3cr1/Crb1(rd8) deficient mice provides new insights into the pathophysiology of progressive retinal degeneration. Exp Eye Res 203:108424. https://doi.org/10.1016/j.exer.2020.108424
Blanchette G, O’Keefe R, Benuskova L (2012) Inference of a phylogenetic tree: hierarchical clustering versus genetic Algorithm. In: Thielscher M, Zhang D (eds) Australasian Joint Conference on Artificial Intelligence. Springer, Berlin, Heidelberg, pp 300–312
Braschi B et al (2019) Genenames.org: the HGNC and VGNC resources in 2019. Nucleic Acids Res 47:d786–d792. https://doi.org/10.1093/nar/gky930
Bucala R, Callati M, Manabe S, Cotlier E, Cerami A (1985) Glucocorticoid-lens protein adducts in experimentally induced steroid cataracts. Exp Eye Res 40(6):853–863
Caban M (2019) Glucocorticosteroid induced cataract - drug-induced damage of vision organ which is worth paying attention in the treatment of inflammatory bowel diseases . Postepy Biochem 65:227–230. https://doi.org/10.18388/pb.2019_277
da Huang W, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4(1):44–57. https://doi.org/10.1038/nprot.2008.211
Eberly LEJR (2003) Correlation and simple linear regression. Radiology 227:617–622
Fan CN, Ma L, Liu N (2018) Systematic analysis of lncRNA-miRNA-mRNA competing endogenous RNA network identifies four-lncRNA signature as a prognostic biomarker for breast cancer. J Transl Med 16:264. https://doi.org/10.1186/s12967-018-1640-2
Fatica A, Bozzoni I (2014) Long non-coding RNAs: new players in cell differentiation and development. Nat Rev Genet 15(1):7–21
Gupta V, Wagner BJ (2003) Expression of the functional glucocorticoid receptor in mouse and human lens epithelial cells. Invest Ophthalmol vis Sci 44:2041–2046. https://doi.org/10.1167/iovs.02-1091
Gupta V, Galante A, Soteropoulos P, Guo S, Wagner BJJMV (2005) Global gene profiling reveals novel glucocorticoid induced changes in gene expression of human lens epithelial cells. Mol vis 11(1018):1040
Hamamichi S, Kosano H, Nakai S, Ogihara-Umeda I, Nishigori H (2003) Involvement of hepatic glucocorticoid receptor-mediated functions in steroid-induced cataract formation. Exp Eye Res 77(5):575–580
Hanxiao R et al (2018) MiR-326 antagomir delays the progression of age-related cataract by upregulating FGF1-mediated expression of betaB2-crystallin. Biochem Biophys Res Commun 505(2):505–510
Huang DW, Sherman BT, Lempicki RA (2009) Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res 37(1):1–13
Hung T, Chang HY (2010) Long noncoding RNA in genome regulation: prospects and mechanisms. RNA Biol 7:582–585. https://doi.org/10.4161/rna.7.5.13216
Li JH, Liu S, Zhou H, Qu LH, Yang JH (2014) starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res 42:D92–D97. https://doi.org/10.1093/nar/gkt1248
Li Y, Liu S, Zhang F, Jiang P, Wu X, Liang Y (2015) Expression of the microRNAs hsa-miR-15a and hsa-miR-16-1 in lens epithelial cells of patients with age-related cataract. Int J Clin Exp Med 8:2405–2410
Liu Z, Li M, Hua Q, Li Y, Wang G (2019) Identification of an eight-lncRNA prognostic model for breast cancer using WGCNA network analysis and a Coxproportional hazards model based on L1-penalized estimation. Int J Mol Med 44:1333–1343. https://doi.org/10.3892/ijmm.2019.4303
Lucas SEM et al (2018) Rare, potentially pathogenic variants in 21 keratoconus candidate genes are not enriched in cases in a large Australian cohort of European descent. PLoS ONE 13:e0199178. https://doi.org/10.1371/journal.pone.0199178
Mccarty CA, Nanjan MB, Taylor HR (2000) Attributable risk estimates for cataract to prioritize medical and public health action. Invest Ophthalmol vis Sci 41:3720
Mercer TR, Dinger ME, Mattick JS (2009) Long non-coding RNAs: insights into functions. Nat Rev Genet 10(3):155–159
Nishigori H, Hayashi R, Lee JW, Maruyama K, Iwatsuru M (1985) Preventive effect of ascorbic acid against glucocorticoid-induced cataract formation of developing chick embryos. Exp Eye Res 40(3):445–451
Paraskevopoulou MD, Georgakilas G, Kostoulas N, Reczko M, Maragkakis M, Dalamagas TM, Hatzigeorgiou AG (2013) DIANA-LncBase: experimentally verified and computationally predicted microRNA targets on long non-coding RNAs. Nucleic Acids Res 41:D239-245. https://doi.org/10.1093/nar/gks1246
Pawlak-Adamska E, Daroszewski J, Bolanowski M, Oficjalska J, Janusz P, Szalinski M, Frydecka I (2013) PPARg2 Ala12 variant protects against Graves’ orbitopathy and modulates the course of the disease. Immunogenetics 65:493–500. https://doi.org/10.1007/s00251-013-0702-0
Ritchie ME, Phipson B, Wu DI, Hu Y, Law CW, Shi W, Smyth GK (2015) Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43(7):e47–e47
Sargazi S, Moudi M, Heidari Nia M, Saravani R, Malek Raisi H (2019) Association of KIF26B and COL4A4 gene polymorphisms with the risk of keratoconus in a sample of Iranian population. Int Ophthalmol 39:2621–2628. https://doi.org/10.1007/s10792-019-01111-x
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13(11):2498–2504
Silva A, Bullock M, Calin G (2015) The clinical relevance of long non-coding RNAS in cancer. Cancers 7(4):2169–2182
Szekely GJ, Rizzo ML (2005) Hierarchical clustering via joint between-within distances: extending Ward’s minimum variance method. J Classif 22(2):151–184
Tang H, Wu Z, Zhang Y, Xia T, Liu D, Cai J, Ye Q (2019) Identification and function analysis of a five-long noncoding rna prognostic signature for endometrial cancer patients. DNA Cell Biol. https://doi.org/10.1089/dna.2019.4944
Barrett T, Troup DB, Wilhite SE, Ledoux P, Rudnev D, Evangelista C, Kim IF, Soboleva A, Tomashevsky M, Edgar R (2007) NCBI GEO: mining tens of millions of expression profiles--database and tools update. Nucleic Acids Res 35(Database issue):D760–D765
van Ipenburg JA, de Waard NE, Naus NC, Jager MJ, Paridaens D, Verdijk RM (2019) Chemokine receptor expression pattern correlates to progression of conjunctival melanocytic lesions . Invest Ophthalmol vis Sci 60:2950–2957. https://doi.org/10.1167/iovs.19-27162
Wang C, Dawes LJ, Liu Y, Wen L, Lovicu FJ, McAvoy JW (2013) Dexamethasone influences FGF-induced responses in lens epithelial explants and promotes the posterior capsule coverage that is a feature of glucocorticoid-induced cataract. Exp Eye Res 111:79–87. https://doi.org/10.1016/j.exer.2013.03.006
Wang T, Li W, Cheng H, Zhong L, Deng J, Ling S (2019) The important role of the chemokine Axis CCR7-CCL19 and CCR7-CCL21 in the pathophysiology of the immuno-inflammatory response in dry eye disease. Ocul Immunol Inflamm. https://doi.org/10.1080/09273948.2019.1674891
Xiang W et al (2016) miR34a suppresses proliferation and induces apoptosis of human lens epithelial cells by targeting E2F3. Mol Med Rep 14:5049–5056. https://doi.org/10.3892/mmr.2016.5901
Zhang XP et al (2018) Ginsenoside Rh2 inhibits vascular endothelial growth factor-induced corneal neovascularization. FASEB J 32:3782–3791. https://doi.org/10.1096/fj.201701074RR
Zhao Y et al (2020) Expression profiles of inflammatory cytokines in the aqueous humor of children after congenital cataract extraction. Translational Vision Science & Technology 9:3. https://doi.org/10.1167/tvst.9.8.3
Zhou H et al (2019) HMDD v3. 0: a database for experimentally supported human microRNA–disease associations. Nucleic Acids Res 47:D1013–D1017
Acknowledgements
None.
Funding
This work was supported by the Heilongjiang Province Postdoctoral Funds (grant number LBH-Z18194).
Author information
Authors and Affiliations
Contributions
PL and ZS were responsible for the conception and design of the research, and drafting the manuscript. X-M Z and H-Y G performed the data acquisition. Y S and CW performed the data analysis and interpretation. ZS and H-YG participated in the design of the study and performed the statistical analysis. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare that are relevant to the content of this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Shi, Z., Zhao, X., Su, Y. et al. Screening of Biological Target Molecules Related to Glucocorticoid-Induced Cataract (GIC) on the Basis of Constructing ceRNA Network. Biochem Genet 60, 24–38 (2022). https://doi.org/10.1007/s10528-021-10078-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-021-10078-3