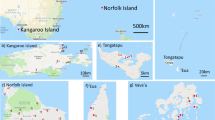
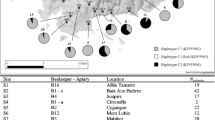
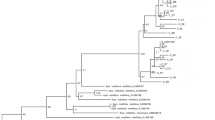
Genetic diversity and population differentiation of the giant honey bee (Apis dorsata) in Thailand were examined. Six PCR-RFLP mitotypes were generated from digestion of the COI-COII, Cytb-tRNAser, ATPase6-8, and lrRNA genes with Dra I and Hin fI. Low genetic diversity (h=0.074, π=0.032%) and a lack of genetic population differentiation between A. dorsata originating from geographically different regions were observed from mtDNA polymorphisms (P > 0.05). In contrast, microsatellite (A14, A24, and A88) polymorphisms revealed a relatively high level of genetic diversity in A. dorsata (H o=0.68–0.74, average number of alleles per locus=6.0–9.0). Both A24 and A88 indicated significant population differentiation between bees from the north-to-central region (north, northeast, and central regions), peninsular Thailand, and Samui Island.




Similar content being viewed by others
REFERENCES
Avise, J. C. (1994). Molecular Markers, Natural History and Evolution. Chapman and Hall, New York. p. 511.
Birky C. W., Jr., Furest, P., and Maruyama, T. (1989). Organelle gene diversity under migration, mutation and drift: equilibrium expectations, approach to equilibrium, effects of heteroplasmic cells, and comparison to nuclear genes. Genetics 121:613–627.
Cavalli-Sforza, L. L., and Edwards, A. W. F. (1967). Phylogenetic analysis: Models and estimation procedures. Amer. J. Hum. Genet. 19:233–257.
Crow, J. F., and Kimura, M. (1965). Evolution in sexual and asexual populations. Am. Nat. 99:439–450.
Crozier, R. H., and Crozier, Y. C. (1993). The mitochondrial genome of the honeybee Apis mellifera: Complete sequence and genome organization. Genetics 133:97–117.
Delarua, P., Serrano, J., and Galian, J. (1998). Mitochondrial DNA variability in the Canary Islands honeybee (Apis mellifera L.). Mol. Ecol. 7:1543–1547.
Deowanish, S., Nakamura, J., Matsuka, M., and Kimura, K. (1996). mtDNA variation among subspecies of Apis cerana using restriction fragment length polymorphism. Apidologie 27:407–413.
Dyer, F. C., and Seeley, T. D. (1994). Colony migration in the tropical honey bee Apis dorsata F. (Hymenoptera: Apidae). Insectes Soc. 41:129–140.
Estoup, A., Solignac, M., Harry, H., and Cornuet, J.-M. (1993). Characterization of (GT)n and (CT)n microsatellites in two insect species: Apis mellifera and Bombus terrestris. Nucleic Acids Res. 21:1427–1431.
Estoup, A., Solignac, M., and Cornuet, J.-M. (1994). Precise assessment of the number of patriline and of genetic relatedness in honey bee colonies. Proc. R.l Soc. London, series B 258:1–7.
Estoup, A., Garnery, L., Solignac, M., and Cornuet, J.-M. (1995). Microsatellite variation in honey bee (Apis mellifera L.) populations: Hierarchical genetic structure and test of the infinite allele and stepwise mutation models. Genetics 140:679–695.
Felsenstein, J. (1993). Phylip (Phylogenetic Inference Package) Version 3.56c. Department of Genetics, University of Washington, Seattle.
Franck, P., Garnery, L., Solignac M., and Cornuet, J.-M. (1998). The origin of west European subspecies of honeybee (Apis mellifera): New insights from microsatellite and mitochondrial data. Evolution 52:1119–1134.
Guo, S. W., and Thompson, E. A. (1992). Performing the exact test of Hardy-Weinberg proportion of multiple alleles. Biometrics 48:361–372.
Koeniger, N., and Koeniger, G. (1980). Observations and experiments on migration and dance communication of Apis dorsata in Sri Lanka. J. Apic. Res. 19:21–34.
Kraus, F. B., Koeniger, N., Tingek, S., and Moritz, R. F. A. (2005). Temporal genetic structure of a drone congregation area of the giant Asian honeybee (Apis dorsata). Naturwissenschaften 92:578–581.
McElroy, D., Moran, P., Birmingham, E., and Kornfield, I. (1991). REAP (Restriction Enzyme Analysis Package) Version 4.0. University of Maine, Orono, Maine.
Nei, M. (1978). Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590.
Nei, M. (1987). Molecular Evolutionary Genetics. Columbia University Press, New York. p. 512.
Nei, M., and Li, W. H. (1979). Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc. Natl. Acad. Sci. USA 76:5269–5273.
Neumann, P., Koeniger, N., Koeniger, G., Tingek, S., Krygerll, P., and Moritz, R. F. A. (2000). Home-site fidelity in migratory honeybees. Nature 406:474–475.
O’Connell, M., Dillon, M. C., Wright, J. M., Bentzen, P., Merkouris, S., and Seeb, J. (1998). Genetic structuring among Alaskan Pacific herring populations identified using microsatellite variation. J. Fish. Biol. 53:150–163.
Parr, J., Oldroyd, B. P., and Kastberger, G. (2000). Giant honeybees return to their nest sites. Nature 406:475.
Parr, J., Oldroyd, B. P., Huettinger, E., and Kastberger, G. (2004). Genetic structure of an Apis dorsata population: The significance of migration and colony aggregation. J. Heredity 95:119– 126.
Raymond, M., and Rousset, F. (1995). GenePop (version 1.2): Population genetics software for exact tests and ecumenicism. J. Heredity 86:248–250.
Roff, D. A., and Bentzen, P. (1989). The statistical analysis of mitochondrial DNA polymorphisms: χ2 and the problems of small samples. Mol. Biol. Evol. 6:539–545.
Saitou, N., and Nei, M. (1987). The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4:406–425.
Sambrook, J., and Russell, D. W. (2001). Molecular Cloning: A Laboratory Manual, 3rd edn. Cold Spring Harbor Laboratory Press, New York.
Sakagami, S. F., Matsumura, T., and Ito, K. (1980). Apis laboriosa in Himalaya, the little known world largest honeybee (Hymenoptera: Apidae). Insecta Matsumrana 19:47–77.
Sihanuntavong, D., Sittipraneed. S., and Klinbunga, S. (1999). Mitochondrial DNA diversity and population structure of the honey bee (Apis cerana) in Thailand. J. Apic. Res. 38:211– 219.
Sittipraneed, S., Laoaroon, S., Klinbunga, S., and Wongsiri, S. (2001a). Genetic differentiation of the honey bee (Apis cerana) in Thailand: Evidence from microsatellite polymorphism. J. Apic. Res. 40:9–16.
Sittipraneed, S., Sihanunthavong, D., and Klinbunga, S. (2001b). Genetic differentiation of the honey bee (Apis cerana) in Thailand revealed by polymorphism of a large subunit of mitochondrial ribosomal DNA. Insectes. Soc. 48:266–272.
Smith, D. R., and Hagen, R. H. (1997). The biogeography of Apis cerana as revealed by mitochondrial DNA sequence data. J. Kansas Entomol. Soc. 69:294–310.
Sylvester, H. A., Limbipichai, K., Wongsiri, S., Rinderer, T. E., and Mardan, M. (1998). Morphometric studies of Apis cerana in Thailand and the Malaysian Pennisula. J. Apic. Res. 37:137–145.
Takezaki, N., and Nei, M. (1996). Genetic distance and reconstruction of phylogenetic trees from microsatellite DNA. Genetics 144:389–399.
Thapa, R., and Wongsiri, S. (1997). Eupatorium odoratum: A honey plant for beekeepers in Thailand. Bee World 78:175–178.
Weir, B. S., and Cockerham, C. C. (1984). Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370.
Wongsiri, S., Thapa, T., Oldroyd, B., and Burgett, D. M. (1996). A magic bee tree: Home to Apis dorsata. American Bee J. 136:796–799.
Wongsiri, S., Chanchao, C., Lekprayoon, C., Wattanasermkit, K., Deowanish, S., and Leepitakrat, S. (2001). Honeybee diversity and management in the new millennium in Thailand. Proc. Seventh International Conference on Tropical Bees: Management and Diversity, 19–25 March 2000. Chiang Mai, Thailand. pp. 9–14.
ACKNOWLEDGMENTS
The authors would like to thank the Departments of Biochemistry and Biology, Faculty of Science, Chulalongkorn University, for providing facilities. This work was supported by the Thailand Research Funds (TRF) Senior Scholar award to Prof. Dr. S. Wongsiri. This research was conducted in cooperation with Louisiana State University.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Insuan, S., Deowanish, S., Klinbunga, S. et al. Genetic Differentiation of the Giant Honey Bee (Apis dorsata) in Thailand Analyzed by Mitochondrial Genes and Microsatellites. Biochem Genet 45, 345–361 (2007). https://doi.org/10.1007/s10528-007-9079-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-007-9079-9