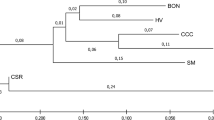
Microsatellite loci were isolated using five repetitive probes for Korean native cattle. Eleven microsatellite loci were developed based on a biotin hybrid capture method, and enrichment of the genomic libraries (AAAT, TG, AG, T, and TGC repeats) was performed using Sau3AI adapters. The isolated markers were tested in two half-sib Korean cattle families and four imported breeds (Angus, Limousine, Holstein, and Shorthorn). Nine informative microsatellite loci were observed, and two microsatellite loci were revealed as monomorphic in Korean cattle. In the imported breeds, however, all of the markers were informative. In total, 213 alleles were obtained at the 11 loci across five breeds, and the average number of alleles found per locus, considering all populations, was 4.26. Heterozygosity was 0.71 (expected) and 0.57 (observed). The range of the polymorphic information content for the markers in all cattle populations was 0.43–0.69. Eleven percent of genetic variation was attributed to differentiation between populations as determined by the mean F ST values. The remaining 89% corresponded to differences among individuals. The isolated markers may be used to identify and classify the local breeds on a molecular basis.
Similar content being viewed by others
REFERENCES
Barker, J. S. F. (1994). A global protocol for determining genetic distances among domestic livestock breeds. Proc. 5th World Congr. Genet. Appl. Livest. Prod. 21:501–508.
Bjornstad, G., Gunby, E., and Roed, K. H. (2000). Genetic structure of Norwegian horse breeds. J. Anim. Breed Genet. 117:307–317.
Botstein, D., White, R. L., Skolnick, M., and Davis, R. W. (1980). Construction of genetic linkage map in man using restriction fragment length polymorphisms. Am. J. Hum. Genet. 32:314–331.
Canon, J., Checa, M. L., Carleos, C., Vega-Pla, J. L., Vallejo, M., and Dunner, S. (2000). The genetic structure of Spanish Celtic horse breeds inferred from microsatellite data. Anim. Genet. 31:39–48.
Chung, H. Y. (2001). Studies on the Effects of the Calpain Family of Genes on Meat Tenderness, Carcass Traits, and Growth Traits in Beef Cattle, PhD Thesis, Ohio State University, Columbus, OH.
Cobb, B. D., and Clarkson, J. M. (1994). A simple procedure for optimizing the polymerase chain reaction (PCR) using modified Taguchi methods. Nucleic Acids Res. 22:3801–3805.
Cooper, S., Bull, C. M., and Gardner, M. G. (1997). Characterization of microsatellite loci from the socially monogamous lizard Tiliqua rugosa using a PCR-based isolation technique. Mol. Ecol. 6:793–795.
Edwards, A., Hammond, H. A., Jin, L., Caskey, C. T., and Chakraborty, R. (1992). Genetic variation at five trimeric and tetrameric tandem repeat loci in four human population groups. Genomics 12:241–253.
Hammond, H. A., Jin, L., Zhong, Y., Caskey, C. T., and Chakraborty, R. (1994). Evaluation of 13 short tandem repeat loci for use in personal identification applications. Am. J. Hum. Genet. 55:175–189.
Jordana, J., Piedrafita, J., Sanchez, A., and Puig, P. (1992). Comparative F statistics analysis of the genetic structure of ten Spanish dog breeds. J. Hered. 83:367–374.
Kandpal, R. P., Kandpal, G., and Weissman, S. M. (1994). Construction of libraries enriched for sequence repeats and jumping clones, and hybridization selection for region-specific markers. Proc. Natl. Acad. Sci. U.S.A. 91:88–92.
Li, S. J., Yang, S. H., Zhao, S. H., and Fan, B. (2004). Genetic diversity analyses of 10 indigenous Chinese pig populations based on 20 microsatellites. J. Anim. Sci. 82:368–374.
MacHugh, D. E., Loftus, R. T., Cunningham, P., and Bradley, D. G. (1998). Genetic structure of seven European cattle breeds assessed using 20 microsatellite markers. Anim. Genet. 29:333–340.
Marshall, T. C., Slate, J., Kruuk, L. E. B., and Pemberton, J. M. (1998). Statistical confidence for likelihood-based paternity inference in natural populations. Mol. Ecol. 9:639–655.
MartoAn-Burriel, I., GarcoAa-Muro, E., and Zaragoza, P. (1999). Genetic diversity analysis of six Spanish native cattle breeds using microsatellites. Anim. Genet. 30:177–182.
Panaud, O., Chen, X., and McCouch, S. R. (1996). Development of microsatellite markers and characterization of simple sequence length polymorphism (SSLP) in rice (Oryza sativa L.). Mol. Genet. 252:597–607.
Raymond, M., and Rousset, F. (1995). Population genetics software for exact tests and ecumenicism GenePop (Version 3.3). J. Hered. 86:248–249.
Schnabel, R. D., Ward, T. J., and Derr, J. N. (2000). Validation of 15 microsatellites for parentage testing in North American bison, Bison bison, and domestic cattle. Anim. Genet. 31:360–366.
Takezaki, P. J., and Nei, M. (1996). Genetic distances and reconstruction of phylogenetic tree from microsatellite DNA. Genetics 144:389–399.
Yu, J. K., Mangor, J., Edwards, E., Thompson, L., and Knapp, S. J. (2000). Simple sequence repeats in sunflower: Length polymorphisms among elite inbred lines for several motifs. In Abstracts of Plant and Animal Genome Conference 8, 9–12 January 2000, San Diego, CA. Available at http://www.intl-pag.org/pag/8/abstracts/pag8735.html.
ACKNOWLEDGMENTS
This study was supported by the National Livestock Research Institute (Project No. NLRI2004-01, “Development of microsatellite loci in animals”). The bovine hybrid mapping panel was provided by Dr. James E. Womack at Texas A&M University.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Chung, H.Y., Kim, T.H., Choi, B.H. et al. Isolation and Characterization of the Bovine Microsatellite Loci. Biochem Genet 44, 518–532 (2006). https://doi.org/10.1007/s10528-006-9055-9
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-006-9055-9