Abstract
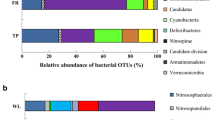
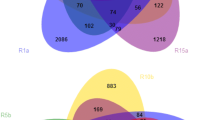
Here we show that both liming the burnt sugarcane and the green harvest practice alter bacterial community structure, diversity and composition in sugarcane fields in northeastern São Paulo state, Brazil. Terminal restriction fragment length polymorphism fingerprinting and 16S rRNA gene cloning and sequencing were used to analyze changes in soil bacterial communities. The field experiment consisted of sugarcane-cultivated soils under different regimes: green sugarcane (GS), burnt sugarcane (BS), BS in soil amended with lime applied to increase soil pH (BSL), and native forest (NF) as control soil. The bacterial community structures revealed disparate patterns in sugarcane-cultivated soils and forest soil (R = 0.786, P = 0.002), and overlapping patterns were shown for the bacterial community structure among the different management regimes applied to sugarcane (R = 0.194, P = 0.002). The numbers of operational taxonomic units (OTUs) found in the libraries were 117, 185, 173 and 166 for NF, BS, BSL and GS, respectively. Sugarcane-cultivated soils revealed higher bacterial diversity than NF soil, with BS soil accounting for a higher richness of unique OTUs (101 unique OTUs) than NF soil (23 unique OTUs). Cluster analysis based on OTUs revealed similar bacterial communities in NF and GS soils, while the bacterial community from BS soil was most distinct from the others. Acidobacteria and Alphaproteobacteria were the most abundant bacterial phyla across the different soils with Acidobacteria Gp1 accounting for a higher abundance in NF and GS soils than burnt sugarcane-cultivated soils (BS and BSL). In turn, Acidobacteria Gp4 abundance was higher in BS soils than in other soils. These differential responses in soil bacterial community structure, diversity and composition can be associated with the agricultural management, mainly liming practices, and harvest methods in the sugarcane-cultivated soils, and they can be detected shortly after harvest.

Source: Google Earth (http://www.google.com.br/intl/pt-BR/earth/)





Similar content being viewed by others
References
Amann RI, Ludwig W, Schleifer KH (1995) Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol Rev 59(1):143–169
Bergmann GT, Bates ST, Eilers KG, Lauber CL, Caporaso JG, Walters WA, Knight R, Fierer N (2011) The under-recognized dominance of Verrucomicrobia in soil bacterial communities. Soil Biol Biochem 43:1450–1455
Brody JR, Kern SE (2004) Sodium boric acid: atriz-less, cooler conductive medium for DNA electrophoresis. Biotechniques 36:214–216
Buckley DH, Schmidt TM (2001) Exploring the biodiversity of soil—a microbial rain forest. In: Staley JT, Reysenbach AL (eds) Biodiversity of microbial life: foundation of Earth’s biosphere. Wiley-Liss Inc., New York, pp 183–208
Buckley DH, Schmidt TM (2003) Diversity and dynamics of microbial communities in soils from agro-ecosystems. Environ Microbiol 5:441–452
Burlage RS, Burlage RS, Atlas R, Stahl D, Geesey G, Sayler G (1998) Techniques in microbial ecology. Oxford University Press, New York, p 468
Catão ECP, Lopes FAC, Araújo JF, Castro AP, Barreto CC, Bustamante MMC et al (2014) Soil acidobacterial 16S rRNA gene sequences reveal subgroup level differences between savanna-like Cerrado and Atlantic forest Brazilian biomes. Int J Microbiol 2014:156341. doi:10.1155/2014/156341
Chao A (1987) Estimating the population size for capture-recapture data with unequal catch ability. Biometrics 43:783–791
Chao A, Lee SM (1992) Estimating the number of classes via sample coverage. J Am Stat Assoc 87:210–217
Clarke KR (1993) Nonparametric multivariate analyses of changes in community structure. Aust J Ecol 18:117–143
Cole JR, Wang Q, Cardenas E, Fish J, Chai B, Farris RJ, Kulam-Syed-Mohideen AS, McgarrelL DM, Marsh T, Garrity GM, Tiedje JM (2009) The Ribossomal Database Project: improved alignments and new tools for rRNA analysis. Nucl Acids Res 37:141–145
Colwell RK, Coddington JA (1994) Estimating terrestrial biodiversity through extrapolation. Philos Trans R Soc B. 345:101–118
CONAB (2013) Acompanhamento de safra brasileira: cana-de-açúcar, segundo levantamento, agosto/2013 Companhia Nacional de Abastecimento. Brasília, p 10
De Figueiredo EB, La Scala N Jr. (2011) Greenhouse gas balance due to the conversion of sugarcane areas from burned to green harvest in Brazil. Agric Ecos Environ 141:77–85
FAO (2016) Crop production. Food and Agriculture Organization of the United Nations. Accessed 06 Jan 2016
Fierer N, Bradford MA, Jackson RB (2007) Toward an ecological classification of soil bacteria. Ecology 88:1354–1364
Garrity GM, Bell JA, Liburun TG (2004) Taxonomic outline of prokaryotes. In: Berey’s manual of systematic bacterriologySpringer, New York
George IF, Liles MR, Hartmann M, Ludwig W, Goodman RM, Agathos SN (2009) Changes in soil Acidobacteria communities after 2,4,6-trinitrotoluene contamination. FEMS Microbiol Lett 296:159–166. doi:10.1111/j.1574-6968.2009.01632.x
Hedlund BP (2011) Phylum XXIII verrucomicrobia phyl. nov. In: Krieg NR, Staley JT, Hedlund BP, Paster BJ, Ward N, Ludwig W, Whitman WB (eds) Bergey’s manual of systematic bacteriology, 2nd edn. Springer, New York, pp 795–799
Janssen PH (2006) Identifying the dominant soil bacterial taxa in libraries of 16S rRNA and 16S rRNA genes. Appl Environ Microbiol 72:1719–1728
Janssen PH, Costa KC, Hedlund BP (2011) Class III. Spartobacteria class. nov. In: Krieg NR, Staley JT, Hedlund BP, Paster BJ, Ward N, Ludwig W, Whitman WB (eds) Bergey’s manual of systematic bacteriology, 2nd edn. Springer, New York, pp 834–836
Jesus EC, Marsh TL, Tiedje JM, Moreira FMS (2009) Changes in land use alter the structure of bacterial communities in Western Amazon soils. ISME J 3:1004–1011
Jones RT, Robeson MS, Lauber CL, Hamady M, Knight R, Fierer N (2009) A comprehensive survey of soil acidobacterial diversity using pyrosequencing and clone library analyses. ISME J 3:442–453
Kuramae E, Gamper H, van Veen J, Kowalchuk G (2011) Soil and plant factors driving the community of soil-borne microorganisms across chronosequences of secondary succession of chalk grasslands with a neutral pH. FEMS Microbiol Ecol 77:285–294
Kuramae EE, Yergeau E, Wong LC, Pijl AS, van Veen JA, Kowalchuk GA (2012) Soil characteristics more strongly influence soil bacterial communities than land-use type. FEMS Microbiol Ecol 79:12–24
Lauber CL, Strickland MS, Bradford MA, Fierer N (2008) The influence of soil properties on the structure of bacterial and fungal communities across land-use types. Soil Biol Biochem 40:2407–2415
Magurran AE (2004) Measuring biological diversity. Blackwell, Oxford
Mendes LW, Taketani RG, Navarrete AA, Tsai SM (2012) Shifts in phylogenetic diversity of archaeal communities in mangrove sediments at different sites and depths in southeastern Brazil. Res Microb 163:366–377
Naether A, Foesel BU, Naegele V, Weinert Wüst PK, Weinert J, Bonkowski M et al (2012) Environmental factors affect acidobacterial communities below the subgroup level in grassland and forest soils. Appl Environ Microbiol 78:7398–7406. doi:10.1128/AEM.01325-12
Navarrete AA, Kuramae EE, de Hollander M, Pijl AS, van Veen JA, Tsai SM (2013) Acidobacterial community responses to agricultural management of soybean in Amazon forest soils. FEMS Microbiol Ecol 83:607–621
Navarrete AA, Diniz TR, Braga LPP, Silva GGZ, Franchini JC, Rossetto R et al (2015a) Multi-analytical approach reveals potential microbial indicators in soil for sugarcane model systems. PLoS One 10(6):e0129765. doi:10.1371/journal.pone.0129765
Navarrete AA, Soares T, Rossetto R, van Veen JA, Tsai SM, Kuramae EE (2015b) Verrucomicrobial community structure and abundance as indicators for changes in chemical factors linked to soil fertility. Antonie Van Leeuwenhoek 108(3):741–752
Navarrete AA, Venturini AM, Meyer KM, Klein AM, Tiedje JM, Bohannan BJ, Nüsslein K, Tsai SM, Rodrigues JL (2015c) Differential response of Acidobacteria subgroups to forest-to-pasture conversion and their biogeographic patterns in the western Brazilian Amazon. Front Microbiol 6:1443. doi:10.3389/fmicb.2015.01443
Panosso AR, Marques JRJ, Milori DMBP, Ferraudo AS, Barbieri DM, Pereira GT, La Scala N (2011) Soil CO2 emission and its relation to soil properties in sugarcane areas under slash-and-burn and Green harvest. Soil Till Res 111:190–196. doi:10.1016/j.still.2010.10.002
Pereira RM, Silveira ÉL, Scaquitto DC, Pedrinho EAN, Val-Moraes SP, Wickert E, Carareto-Alves LM, Lemos EGM (2006) Molecular characterization of bacterial populations of different soils. Braz J Microbiol 37(4):439–447. doi:10.1590/S1517-83822006000400007
Rachid CTCC, Santos AL, Piccolo MC, Balieiro FC, Coutinho HLC et al (2013) Effect of sugarcane burning or green harvest methods on the Brazilian cerrado soil bacterial community structure. PLoS One 8(3):e59342. doi:10.1371/journal.pone.0059342
Raij B, Van Quaggio JA, Cantarella H, Ferreira ME, Lopes AS, Bataglia CO (1987) Análise química do solo para fins de fertilidade. Fundação Cargill Campinas, Campinas
Rasche F, Knapp D, Kaiser C, Koranda M, Kitzler B, Zechmeister-Boltenstern S, Richter A, Sessitsch A (2011) Seasonality and resource availability control bacterial and archaeal communities in soils of a temperate beech forest. ISME J 5:389–402
Rawat SR, Männistö MK, Bromberg Y, Haggblom MM (2012) Comparative genomic and physiological analysis provides insights into the role of Acidobacteria in organic carbon utilization in Arctic Tundra soils. FEMS Microbiol Ecol 82:341–355. doi:10.1111/j.1574-6941.2012.01381.x
Razafimbelo T, Barthes B, Larre-Larrouy MC, De Luca EF, Laurent JY, Cerri CC, Feller C (2006) Effect of sugarcane residue management (mulching versus burning) on organic matter in a clayey oxisol from southern Brazil. Agric Ecos Environ 115(1–4):285–289
Rehm G, Munter R, Rosen C, Schmitt M (1992) Liming materials for Minnesota soils. University of Minnesota Extension Service, St. Paul
Roesch LFW, Passaglia LMP, Bento FM, Triplett EW, Camargo FAO (2007) Diversidade de bactérias diazotróficas endofíticas associadas a plantas de milho. Rev Bras Ciênc Solo 31:1367–1380. doi:10.1590/S0100-06832007000600015
Rousk J, Bååth E, Brookes PC, Lauber CL, Lozupone C, Caporaso JG, Knight R, Fierer N (2010) Soil bacterial and fungal communities across a pH gradient in an Arable soil. ISME J 4(10):1340–1351. doi:10.1038/ismej.2010.58
Sait M, Davis KE, Janssen PH (2006) Effect of pH on isolation and distribution of members of subdivision 1 of the phylum Acidobacteria occurring in soil. Appl Environ Microbiol 72:1852–1857
Sambrook J, Russel DW (2001) Molecular cloning: a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, p 2028
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartman M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Van Horn DJ, Weber CF (2009) Introducing MOTHUR: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541
Sharma SK, Ramesh A, Sharma MP, Joshi OP, Govaerts B, Steenwerth KL, Karlen DL (2011) Microbial community structure and diversity as indicators for evaluating soil quality, 5th edn., Biodiversity, biofuels, agroforestry and conservation agricultureSpringer, New York. doi:10.1007/978-90-481-9513-8
Souza RA, Telles TS, Machado W, Hungria M, Tavares Filho J, Guimarães MF (2012) Effects of sugarcane harvesting with burning on the chemical and microbiological properties of the soil. Agric Ecosyst Environ 155:1–6
Spera ST, Reatto A, Martins ES, Cunha RJF, Correia J (2001) A Desertificação em solos arenosos No Brasil. http://www.cpac.embrapa.br/publicacoes/search_pbl/1?q=solo%20arenoso. Accessed 01 Jul 2015
Tavares Filho J, Barbosa GMC, Ribon AA (2010) Physical properties of dystrophic Red Latosol (oxisol) under different agricultural uses. Rev Bras Ciênc Solo 34:925–933. doi:10.1590/S0100-06832010000300034
ter Braak CJF, Šmilauer P (2002) CANOCO reference manual and CanoDraw for Windows user’s guide: software for canonical community ordination (version 4.5). Microcomputer Power, New York
Togawa RC, Júnior GJ, Quirino BF, Bustamante MM, Williamson L, Handelsman J, Krüger RH (2012) Characterization of soil bacterial assemblies in Brazilian Savanna-like vegetation reveals Acidobacteria dominance. Microb Ecol 64(3):760–770
Torsvik V, Daae FL, Sandaa RA, Ovreas L (1998) Novel techniques for analyzing microbial diversity in natural and perturbed environments. J Biotechnol 64(1):53–62
Uroz S, Buée M, Murat C, Frey-Klett P, Martin F (2010) Pyrosequencing reveals a contrasted bacterial diversity between oak rhizosphere and surrounding soil. Environ Microbiol Rep 2:281–288
Val-Moraes SP, Valarini MJ, Ghini R, Lemos EGM, Carareto-Alves LM (2009) Diversidade de bactérias de solo sob vegetação natural e cultivo de hortaliças. RCA 40(1):7–16
Val-Moraes SP, Pedrinho EAN, Lemos EGM, Carareto-Alves LM (2013) Molecular identification of fungal communities in a soil cultivated with vegetables and soil suppressiveness to Rhizoctonia solani. Appl Environ Soil Sci . doi:10.1155/2013/268768
Venter JC, Remington K, Heidelberg JF et al (2004) Environmental genome shotgun sequencing of the Sargasso Sea. Science. doi:10.1126/science.1093857
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267. doi:10.1128/AEM.00062-07
Ward NL, Challacombe JF, Janssen PH, Henrissat B, Coutinho PM, Wu M et al (2009) Three genomes from the phylum Acidobacteria provide insight into the lifestyles of these microorganisms in soils. Appl Environ Microb 75:2046–2056. doi:10.1128/AEM.02294-08
Woese CR, Stackebrandt E, Macke TJ, Fox GE (1985) A phylogenetic definition of the major eubacterial taxa. Syst Appl Microb 6:143–151
Wright ES, Yilmaz LS, Noguera DR (2012) DECIPHER, a search-based approach to chimera identification for 16S rRNA sequences. Environ Microb Appl 78(3):717–725. doi:10.1128/AEM.06516-11
Zhao N, He N, Wang Q, Zhang X, Wang R, Xu Z, Yu G (2014) The altitudinal patterns of leaf C:N:P stoichiometry are regulated by plant growth form, climate and soil on Changbai Mountain China. PLoS One 9(4):e95196. doi:10.1371/journal.pone.0095196
Acknowledgments
This research was supported by FAPESP and CNPq. AAN was supported by FAPESP grants (FAPESP 12/13321-7).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Silvana Pompeia Val-Moraes and Helena Suleiman de Macedo have contributed equally to this work.
Rights and permissions
About this article
Cite this article
Val-Moraes, S.P., de Macedo, H.S., Kishi, L.T. et al. Liming in the sugarcane burnt system and the green harvest practice affect soil bacterial community in northeastern São Paulo, Brazil. Antonie van Leeuwenhoek 109, 1643–1654 (2016). https://doi.org/10.1007/s10482-016-0764-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-016-0764-8