Abstract
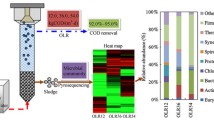
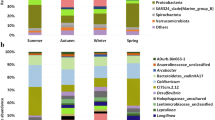
We investigated the microbial community in an up-flow anaerobic sludge blanket (UASB) reactor treating domestic wastewater (DW) during two different periods of organic loading rate (OLR) and food-to-microorganism (F/M) ratio. 16S rDNA clone libraries were generated, and quantitative real-time PCR (qPCR) analyses were performed. Fluctuations in the OLR and F/M ratio affected the abundance and the composition of the UASB prokaryotic community, mainly at the species level, as well as the performance of the UASB reactor. The qPCR analysis suggested that there was a decrease in the bacterial cell number during the rainy season, when the OLR and F/M ratio were lower. However, the bacterial diversity was higher during this time, suggesting that the community degraded more diversified substrates. The diversity and the abundance of the archaeal community were higher when the F/M ratio was lower. Shifts in the methanogenic community composition might have influenced the route of methane production, with methane produced by acetotrophic methanogens (dry season), and by hydrogenotrophic, methylotrophic and acetotrophic methanogens (rainy season). This study revealed higher levels of bacterial diversity, metabolic specialization and chemical oxygen demand removal efficiency of the DW UASB reactor during the rainy season.




Similar content being viewed by others
References
Alves M, Cavaleiro AJ, Ferreira EC, Amaral AL, Mota M, da Motta M, Vivier H, Pons MN (2000) Characterisation by image analysis of anaerobic sludge under shock conditions. Water Sci Technol 41:207–214
APHA, AWWA, WEF (1992) Standard methods for the examination of water and wastewater, 18th edn. American Public Health Association, American Water Works Association and Water Environment Federation, Washington DC
Ariesyady HD, Ito T, Okabe S (2007) Functional bacterial and archaeal community structures of major trophic groups in a full-scale anaerobic sludge digester. Water Res 7:1554–1568
Boone DR, Whitman WB, Rouvière P (1993) Diversity and taxonomy of methanogens. In: Ferry J (ed) Methanogenesis: ecology, physiology, biochemistry & genetics. Champman and Hall, New York, pp 35–80
Cardinali-Rezende J, Debarry RB, Colturato LFDB, Carneiro EV, Chartone-Souza E, Nascimento AMA (2009) Molecular identification and dynamics of microbial communities in digester treating organic household waste. Appl Microbiol Biotechnol 84:777–789
Cardinali-Rezende J, Colturato LFDB, Colturato TDB, Chartone-Souza E, Nascimento AMA, Sanz JL (2012) Prokaryotic diversity and dynamics in a full-scale municipal solid waste anaerobic reactor from start-up to steady-state conditions. Bioresour Technol 119:373–383
Chan OC, Wolf M, Hepperle D, Casper P (2002) Methanogenic archaeal community in the sediment of an artificially partitioned acidic bog lake. FEMS Microbiol Ecol 42:119–129
Delong EF (1992) Archaea in coastal marine environments. Proc Natl Acad Sci USA 9:5685–5689
Díaz EE, Stams AJ, Amils R, Sanz JL (2006) Phenotypic properties and microbial diversity of methanogenic granules from a full scale upflow anaerobic sludge bed reactor treating brewering wastewater. Appl Environ Microbiol 72:4942–4949
Fernandes H, Jungles MK, Hoffmann H, Antonio RV, Costa RHR (2013) Full-scale sequencing batch reactor (SBR) for domestic wastewater: performance and diversity of microbial communities. Bioresour Technol 132:262–268
Good IJ (1953) The population frequencies of species and the estimation of population parameters. Biometrika 40:237–262
Guerrero L, Omil F, Mendez R, Lema JM (1999) Anaerobic hydrolysis and acidogenesis of wastewaters of food industries with high content of organic solids and protein. Water Res 33:3281–3290
Hammes F, Kalogo Y, Verstraete W (2000) Anaerobic digestion technologies for closing the domestic water, carbon and nutrient cycles. Water Sci Technol 41:203–211
Hattori S, Kamagata Y, Hanada S, Shoun H (2000) Thermacetogenium phaeum gen. nov., sp. nov., a strictly anaerobic, thermophilic, syntrophic acetate-oxidizing bacterium. Int J Syst Evol Microbiol 50:1601–1609
Imachi H, Sakai S, Sekiguchi Y, Hanada S, Kamagata Y, Ohashi A, Harada H (2008) Methanolinea tarda gen. nov., sp. nov., a methane producing archaeon isolated from a methanogenic digester sludge. Int J Syst Evol Microbiol 58:294–301
Islam T, Jensen S, Reigstad LJ, Larsen O, Birkeland NK (2008) Methane oxidation at 55°C and pH 2 by a thermoacidophilic bacterium belonging to the Verrucomicrobia phylum. Proc Natl Acad Sci USA 105:300–304
Lane DJ (1991) 16S/23S rDNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, New York, pp 115–148
Levén L, Anders R, Ericksson B, Schurer A (2007) Effect of process temperature on bacterial and archaeal communities in two methanogenic biodigesters treating organic household waste. FEMS Microbiol Ecol 59:683–693
Lobato LCS, Chernicharo CAL, Souza CL (2012) Estimates of methane loss and energy recovery potential in anaerobic reactors treating domestic wastewater. Water Sci Technol 66:2745–2753
Lucena RM, Gavazza S, Florencio L, Kato MT, Morais MAJ (2011) Study of the microbial diversity in a full-scale UASB reactor treating domestic wastewater. J Microbiol Biotechnol 27:2893–2902
Mahmoud N, Zeeman G, Gijzen H, Lettinga G (2003) Solids removal in upflow anaerobic reactors, a review. Bioresour Technol 90:1–9
McMahon KD, Stroot PG, Mackie RI, Raskin L (2001) Anaerobic codigestion of municipal solid waste and biosolids under various mixing conditions-II: microbial population dynamics. Water Res 35:1817–1827
Metcalf and Eddy Inc. (1991) Wastewater engineering: treatment, disposal, reuse. McGraw-Hill Book Company, New York, p 1334
Moyer CL, Dobbs FC, Karl DM (1994) Estimation of diversity and community structure through restriction fragment length polymorphism distribution analysis of bacterial 16S rRNA genes from a microbial mat at an active, hydrothermal vent system, Loihi Seamount, Hawaii. Appl Environ Microbiol 60:871–879
Muyzer G, Waal EC, Uitterlinden AG (1993) Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol 59:695–700
Nelson MC, Morrison M, Yu Z (2011) A meta-analysis of the microbial diversity observed in anaerobic digesters. Bioresour Technol 102:3730–3739
Pauss A, Samson R, Guiot S, Beauchemin C (1990) Continuous measurement of dissolved H2 in an anaerobic reactor using a new hydrogen/air fuel cell detector. Biotechnol Bioeng 35:492–501
Petersen S, Ahring B (1991) Acetate oxidation in a thermophilic anaerobic sewage-sludge digestor: the importance of non-aceticlastic methanogenesis from acetate. FEMS Microbiol Ecol 86:149–158
Petković H, Cullum J, Hranueli D, Hunter IS, Perić-Concha N, Pigac J, Thamchaipenet A, Vujaklija D, Long PF (2006) Genetics of Streptomyces rimosus, the oxytetracycline producer. Microbiol Mol Biol Rev 70:704–728
Reis MP, Barbosa FA, Chartone-Souza E, Nascimento AMA (2013) The prokaryotic community of a historically mining-impacted tropical stream sediment is as diverse as that from a pristine stream sediment. Extremophiles 17:301–309
Rivière D, Desvignes V, Pelletier E, Chaussonnerie S, Guermazi S, Weissenbach J, Li T, Camacho P, Sghir A (2009) Towards the definition of a core of microorganisms involved in anaerobic digestion of sludge. ISME J 3:700–714
Roest K, Altinbas M, Paulo PL, Heilig HGHJ, Akkermans ADL, Smidt H, Vos WM, Stams AM (2005) Enrichment and detection of microorganisms involved in direct and indirect methanogenesis from methanol in an anaerobic thermophilic bioreactor. Microb Ecol 50:440–446
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory, Huntington
Sanz JL, Kochling T (2007) Molecular biology techniques used in wastewater treatment: an overview. Process Biochem 42:119–133
Schloss PD, Handelsman J (2005) Introducing DOTUR, a computer program for defining operational taxonomic units and estimating species richness. Appl Environ Microbiol 71:1501–1506
Schmidt JE, Ahring BK (1997) Treatment of waste water from a multi product food-processing company, in upflow anaerobic sludge blanket (UASB) reactors: the effect of seasonal variation. Pure Appl Chem 69:2447–2452
Sharma S, Radl V, Kloos K, Fuka MM, Engel M, Schauss K, Schloter M (2007) Quantification of functional genes from prokaryotes in soil by PCR. J Microbiol Methods 68:445–452
Shimizu S, Upadhye R, Ishijima Y, Naganuma T (2010) Methanosarcina horonobensis sp. nov., a methanogenic archaeon isolated from a deep subsurface Miocene formation. Int J Syst Evol Microbiol 61:2503–2507
Shin SG, Zhou BW, Lee S, Kim W, Hwang S (2011) Variations in methanogenic population structure under overloading of pre-acidified high-strength organic wastewaters. Process Biochem 46:1035–1038
Urban I, Weichgrebe D, Rosenwinkel KH (2007) Anaerobic treatment of municipal wastewater using the UASB-technology. Water Sci Technol 56:37–44
van Haandel AC, Lettinga G (1994) Anaerobic sewage treatment: a practical guide for regions with a hot climate. Wiley, Chichester, p 226
Wan C-Y, De Wever H, Diels L, Thoeye C, Liang J-B, Huang L-N (2011) Biodiversity and population dynamics of microorganisms in a full-scale membrane bioreactor for municipal wastewater treatment. Water Res 45:1129–1138
Zhou J, Xia B, Treves DS, Wu LY, Marsh TL, O’Neill RV, Palumbo AV, Tiedje JM (2002) Spatial and resource factors influencing high microbial diversity in soil. Appl Environ Microbiol 68:326–334
Acknowledgments
We acknowledge Juliano Leal from Núcleo de Análise de Genoma e Expressão Gênica (NAGE) at the Universidade Federal de Minas Gerais for his technical assistance in the 16S rRNA gene sequencing. We appreciate the financial support provided by the Fundação de Amparo à Pesquisa do Estado de Minas Gerais (FAPEMIG), Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Pró-reitoria de Pesquisa da Universidade Federal de Minas Gerais (PRPq-UFMG) and Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) in the form of a scholarship to Juliana Cardinali Rezende.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
Phylogenetic tree of the bacterial community in the sludge sampled during the dry season (SD), which was constructed using the neighbor-joining method. The numbers at the nodes indicate the percentages of occurrence in 1,000 bootstrapped trees. Methanococcus nannielii (AY196675) was used as the outgroup. Supplementary material 1 (TIFF 72378 kb)
Fig. S2
Phylogenetic tree of the bacterial community in the sludge sampled during the rainy season (SR), which was constructed using the neighbor-joining method. The numbers at the nodes indicate the percentages of occurrence in 1,000 bootstrapped trees. Methanococcus nannielii (AY196675) was used as the outgroup. Supplementary material 2 (TIFF 94325 kb)
Fig. S3
Phylogenetic tree of the archaeal community from sludge samples during the dry (SD) and rainy (SR) seasons, which was constructed using the neighbor-joining method. The numbers at the nodes indicate the percentage of occurrence in 1,000 bootstrapped trees. Methanopyrus kandleri (U57340) was used as the outgroup. Supplementary material 3 (TIFF 53 kb)
Rights and permissions
About this article
Cite this article
Cardinali-Rezende, J., Araújo, J.C., Almeida, P.G.S. et al. Organic loading rate and food-to-microorganism ratio shape prokaryotic diversity in a demo-scale up-flow anaerobic sludge blanket reactor treating domestic wastewater. Antonie van Leeuwenhoek 104, 993–1003 (2013). https://doi.org/10.1007/s10482-013-0018-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-013-0018-y