Abstract
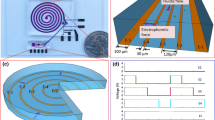
On-chip concentration method for deoxyribonucleic acid (DNA) molecules is preconcentrating DNA molecules before analyses using nanometer-sized structures formed in a microchannel and is effective in improving the sensitivity in DNA analyses using microfluidic devices. Although accurate and predictive theoretical models of concentration profile and concentration factor in on-chip concentration can be used for designing nanostructures and optimizing conditions for concentration, there has been few studies on models. In our previous study, we presented a method of on-chip concentration of DNA molecules using a nanoslit, which is a smaller gap than the diameter of random-coiled DNA molecules. This method is based on the principle of an entropic trap, and we achieved DNA concentration by controlling the applied voltage. In this study, we developed theoretical models of concentration profile and concentration factor in on-chip concentration for DNA molecules using a nanoslit. We conducted concentration experiments of lambda DNA (λ DNA) using our fabricated chip device with a 25-nm nanoslit. The theoretical results of our models were in good agreement with these experimental results. Based on our theoretical models, we determined the optimal applied voltage to be 0.95 V for maximizing the concentration factor in λ DNA concentration by using our chip device.









Similar content being viewed by others
References
Arvanitidou E, Hoagland D (1991) Chain-length dependence of the electrophoretic mobility in random gels. Phys Rev Lett 67:1464–1466. https://doi.org/10.1103/PhysRevLett.67.1464
Azuma N, Itoh S, Fukuzawa K, Zhang H (2016) Separation of large DNA molecules by size exclusion chromatography-based microchip with on-chip concentration structure. Jpn J Appl Phys 55:06GN01
Azuma N, Itoh S, Fukuzawa K, Zhang H (2017) Optimization of applied voltages for on-chip concentration of DNA using nanoslit. Jpn J Appl Phys 56:127001
Azuma N, Itoh S, Fukuzawa K, Zhang H (2018) Separation of large DNA molecules by applying pulsed electric field to size exclusion chromatography-based microchip. Jpn J Appl Phys 57:027002
Broyles BS, Jacobson SC, Ramsey JM (2003) Sample filtration, concentration, and separation integrated on microfluidic devices. Anal Chem 75:2761–2767. https://doi.org/10.1021/ac025503x
Chun H, Chung TD, Ramsey JM (2010) High yield sample preconcentration using a highly ion-conductive charge-selective polymer. Anal Chem 82:6287–6292. https://doi.org/10.1021/ac101297t
Duan C, Majumdar A (2010) Anomalous ion transport in 2-nm hydrophilic nanochannels. Nat Nanotechnol 5:848–852. https://doi.org/10.1038/nnano.2010.233
Fu LM, Hou HH, Chiu PH, Yang RJ (2018) Sample preconcentration from dilute solutions on micro/nanofluidic platforms: a review. Electrophoresis 39:289–310. https://doi.org/10.1002/elps.201700340
Graham SA, Ivanir NM, Paratore F, Scheper T, Bercovici M, Segal E (2017) On chip protein pre-concentration for enhancing the sensitivity of porous silicon biosensors. ACS Sens 2:1767–1773. https://doi.org/10.1021/acssensors.7b00692
Haeberle S, Zengerle R (2007) Microfluidic platforms for lab-on-a-chip applications. Lab Chip 7:1094–1110. https://doi.org/10.1039/B706364B
Han J, Turner SW, Craighead HG (1999) Entropic trapping and escape of long DNA molecules at submicron size constriction. Phys Rev Lett 83:1688–1691. https://doi.org/10.1103/PhysRevLett.83.1688
Hong SA, Kim YJ, Kim SJ, Yang S (2018) Electrochemical detection of methylated DNA on a microfluidic chip with nanoelectrokinetic pre-concentration. Biosens Bioelectron 107:103–110. https://doi.org/10.1016/j.bios.2018.01.067
Jo K, Dhingra DM, Odijk T, Pablo JJ, Graham MD, Runnheim R, Forrest D, Schwartz DC (2007) A single-molecule barcoding system using nanoslits for DNA analysis. Proc Natl Acad Sci USA 104:2673–2678. https://doi.org/10.1073/pnas.0611151104
Kaji N, Tezuka Y, Takamura Y, Ueda M, Nishimoto T, Nakanishi H, Horiike Y, Baba Y (2004) Separation of long DNA molecules by quartz nanopillar chips under a direct current electric field. Anal Chem 76:15–22. https://doi.org/10.1021/ac030303m
Khandurina J, Jacobson SC, Waters LC, Foote RS, Ramsey JM (1999) Microfabricated porous membrane structure for sample concentration and electrophoretic analysis. Anal Chem 71:1815–1819. https://doi.org/10.1021/ac981161c
Kim SM, Burns MA, Hasselbrink EF (2006) Electrokinetic protein preconcentration using a simple glass/poly(dimethylsiloxane) microfluidic chip. Anal Chem 78:4779–4785. https://doi.org/10.1021/ac060031y
Lee SJ, Lee SY (2004) Micro total analysis system (μ-TAS) in biotechnology. Appl Microbiol Biotechnol 64:289–299. https://doi.org/10.1007/s00253-003-1515-0
Lichtenberg J, Verpoorte E, Rooij NF (2001) Sample preconcentration by field amplification stacking for microchip-based capillary electrophoresis. Electrophoresis 22:258–271. https://doi.org/10.1002/1522-2683(200101)22:2<258:AID-ELPS258>3.0.CO;2-4
Lin CC, Hsu JL, Lee GB (2011) Sample preconcentration in microfluidic devices. Microfluid Nanofluid 10:481–511. https://doi.org/10.1007/s10404-010-0661-9
Lion N, Rohner TC, Dayon L, Arnaud IL, Damoc E, Youhnovski N, Wu ZY, Roussel C, Josserand J, Jensen H, Rossier JS, Przybylski M, Girault HH (2003) Microfluidic systems in proteomics. Electrophoresis 24:3533–3562. https://doi.org/10.1002/elps.200305629
Martins D, Levicky R, Song YA (2015) Enhancing the speed of morpholino-DNA biosensor by electrokinetic concentration of DNA in a microfluidic chip. Biosens Bioelectron 72:87–94. https://doi.org/10.1016/j.bios.2015.04.063
Pereira FP, Lavilla I, Bendicho C (2009) Miniaturized preconcentration methods based on liquid–liquid extraction and their application in inorganic ultratrace analysis and speciation: a review. Spectrochim Acta Part B: At Spectrosc 64:1–15. https://doi.org/10.1016/j.sab.2008.10.042
Qi Y, Zeng L, Khorshid A, Hill RJ, Reisner WW (2018) Compression of nanoslit confined polymer solutions. Macromolecules 51:617–625. https://doi.org/10.1021/acs.macromol.7b01894
Scalettar BA, Hearst JE, Klein MP (1989) FRAP and FCS studies of self-diffusion and mutual diffusion in entangled DNA solutions. Macromolecules 22:4550–4559
Shafera KK, Ortiz JH, Potamousis K, Tsvidc G, Place M, Ravindranc P, Jo K, Zhou S, Odijk T, Pablo JJ, Schwartz DC (2017) Electrostatic confinement and manipulation of DNA molecules for genome analysis. Proc Natl Acad Sci USA 114:13400–13405. https://doi.org/10.1073/pnas.1711069114
Song S, Singh AK (2006) On-chip sample preconcentration for integrated microfluidic analysis. Anal Bioanal Chem 384:41–43. https://doi.org/10.1007/s00216-005-0206-3
Sorlie SS, Pécora R (1988) A dynamic light scattering study of a 2311 base pair DNA restriction fragment. Macromolecules 21:1437–1449
Tang Q, Wang X, Hu J (2017) Nano concentration by acoustically generated complex spiral vortex field. Appl Phys Lett 110:104105. https://doi.org/10.1063/1.4978370
Temiz Y, Lovchik RD, Kaigala GV, Delamarche E (2015) Lab-on-a-chip devices: how to close and plug the lab? Microelectron Eng 132:156–175. https://doi.org/10.1016/j.mee.2014.10.013
Vestergaard CL, Mikkelsen MB, Reisner W, Kristensen A, Flyvbjerg H (2016) Transition state theory demonstrated at the micron scale with out-of-equilibrium transport in a confined environment. Nat Commun 7:1–9. https://doi.org/10.1038/ncomms10227
Wu Q, Kaji N, Yasui T, Rahong S, Yanagida T, Kanai M, Nagashima K, Tokeshi M, Kawai T, Baba Y (2017) A millisecond micro-RNA separation technique by a hybrid structure of nanopillars and nanoslits. Sci Rep 7:43877. https://doi.org/10.1038/srep43877
Xu L, Basheer C, Lee HK (2007) Developments in single-drop microextraction. J Chromatogr A 1152:184–192. https://doi.org/10.1016/j.chroma.2006.10.073
Yang RJ, Pu HH, Wang HL (2015) Ion concentration polarization on paper-based microfluidic devices and its application to preconcentrate dilute sample solutions. Biomicrofluidics 9:014122. https://doi.org/10.1063/1.4913366
Acknowledgements
This work was supported in part by JSPS KAKENHI Grant 19K21109 and the Nitto Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
There are no conflicts to declare.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Azuma, N., Itoh, S. & Fukuzawa, K. Optimizing on-chip concentration of DNA molecules against a nanoslit barrier. Microfluid Nanofluid 24, 86 (2020). https://doi.org/10.1007/s10404-020-02392-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10404-020-02392-w