Abstract
The purpose of this paper is to systematically sort out and analyze the cutting-edge research on the eligibility criteria of clinical trials. Eligibility criteria are important prerequisites for the success of clinical trials. It directly affects the final results of the clinical trials. Inappropriate eligibility criteria will lead to insufficient recruitment, which is an important reason for the eventual failure of many clinical trials. We have investigated the research status of eligibility criteria for clinical trials on academic platforms such as arXiv and NIH. We have classified and sorted out all the papers we found, so that readers can understand the frontier research in this field. Eligibility criteria are the most important part of a clinical trial study. The ultimate goal of research in this field is to formulate more scientific and reasonable eligibility criteria and speed up the clinical trial process. The global research on the eligibility criteria of clinical trials is mainly divided into four main aspects: natural language processing, patient pre-screening, standard evaluation, and clinical trial query. Compared with the past, people are now using new technologies to study eligibility criteria from a new perspective (big data). In the research process, complex disease concepts, how to choose a suitable dataset, how to prove the validity and scientific of the research results, are challenges faced by researchers (especially for computer-related researchers). Future research will focus on the selection and improvement of artificial intelligence algorithms related to clinical trials and related practical applications such as databases, knowledge graphs, and dictionaries.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Clinical trials are the most important link in the marketing cycle of new drugs. Eligibility criteria are the most important part of clinical trials. The success of clinical trials depends on the correct eligibility criteria. Clinical trials in the past have often faced multiple problems including under-recruitment, recruiting enough people but failing to demonstrate the efficacy and safety of interventions, and unreasonable experimental designs [1]. These issues are directly or indirectly related to the eligibility criteria. In the second half of the last century, to solve these problems, researchers conducted research through retrospective research, issuing questionnaires [2, 3], etc. Now, with the advent of the era of big data and the dramatic increase in medical-related electronic data such as electronic health records (EHR), researchers are turning to use artificial intelligence to process the increasingly complex data. The urgency of this action is further demonstrated by the emergence of COVID-19. Take vaccine development as an example: In 2020, there are more than 80 potential vaccine candidates being studied [4], but only a few have finally started clinical trials and entered production. The case of Covid-19 demonstrates the need to change the way to run the clinical trials. Compared with the complex, cumbersome and lengthy manual processes, a set of rigorously tested artificial intelligence algorithms has a good chance of replacing most of the manual processes in clinical trials.
Material and methods
Paper selection
The materials of this paper are various published and unpublished papers. The main source of papers is Google Scholar, arXiv, Nature, NIH (National Library of medicine) and other academic related platforms. On these paper platforms, we use keywords such as “Eligibility Criteria” and “Clinical Trial” to query. After reading these papers, we also study their reference papers and incorporate them into our research.
Research methods
We focus on computer and clinical trials related papers and collate the direction, purpose, object, data set, results and shortcomings of these papers. We classify these interdisciplinary papers according to their research directions. We have studied the data sets used in these papers and made a relevant table.
Results
From the beginning to the present, natural language processing (NLP) has been an important research direction. Except the NLP, research on eligibility criteria also includes many research directions such as patient matching, clinical trial evaluation, and clinical trial inquiry. Figure 1 shows the relationship between these research directions.
Research hotspots
Natural language processing (NLP)
Natural language processing (NLP) is one of the important foundations of eligibility criteria research. In NLP, information extraction is the main research direction. The role of information extraction is to convert “human language” into “machine language.” It is to convert the eligibility criteria and patient data in free text form into structured data that can be recognized by computers. NLP is the data source for all computer-related research in this field and an important research basis. Most research focuses on two components of NLP: entity name recognition (NER) and relation extraction (NEL). Because NLP spans many fields, the development and progress of NLP in the field of clinical trials always comes from breakthroughs in other NLP fields.
From more than ten years ago to the present, in the field of clinical medicine, NLP models have undergone a long development process: early semi-automatic methods based on pattern matching and rules: such as the pattern matching and rule-based method proposed by Tu [5] in 2010, object-oriented model such as Elixir proposed by Weng [6] in 2011; various models related to mid-term and machine learning: Criteria2Query model proposed by Yuan [7], which combines machine learning and rule-based methods, which is of great significance for later generations (although this is only a small part of his research); then to the later deep learning period: after the launch of word2vec, the Facebook research team led by Tseo [8] applied it to entity recognition in clinical trials, Pandey [9] proposed a BiLSTM and attention mechanism based on Encoder–Decoder model; and in recent years, after Google proposed two well-known pre-training models for migration learning: Transformer model and Bert model, researchers proposed BioBert [10], BioELECTRA [11], BioALBERT [12], CT-BERT [13], BLURB [14] and other pre-trained models suitable for biomedicine, and incorporate them into the field of clinical trials. The development of NLP in the field of clinical trials has been combined with the state-of-the-art at the time. Regarding clinical trial NLP, there is another organization that must be mentioned: n2c2 (National NLP Clinical Challenges), which publishes a series of clinical trial NLP challenges every year. In addition to having access to large datasets, n2c2 produces a large number of high-quality papers every year [15, 16].
In addition to the various studies mentioned above, there are many systematic summaries of the applications of NLP in medicine and biology. For example: Udo [17] surveyed the fundamental methodological paradigm shift of medical Information Extraction (IE) from standard Machine Learning (ML) to Deep Neural Networks (DNN). Seyedmostafa [18] studied the development and uptake of NLP methods applied to clinical notes related to chronic diseases. Surabhi [19] presented a review of clinical NLP literature for cancer.
Except researching higher-accuracy and more general-purpose NLP models, another important part of NLP research is corpora. Since NLP research requires a large amount of data (this is more important for models based on supervised learning), corpora are needed to provide building applications and training models. And clinical trials are no exception. The data sources of the clinical trial corpus are mainly those texts that are included in the eligibility criteria. But here's the problem: Entity name recognition in NLP requires an already defined entity type. Therefore, when building a corpus, it is necessary to label important features in the data, which is very time consuming. How to mark these contents is a problem that researchers must consider. As the medical profession is involved, the help of medical professionals is indispensable. Although researchers can also refer to authoritative lexicons such as UMLS [20] when annotating texts and use various professional annotation tools (such as BRAT [21]) to speed up annotation, different understandings and different purposes lead to different annotation mode. This eventually led to the birth of various corpora with different uses and different volumes and contents. Researchers usually build corpora according to their needs when conducting NLP research. For example, Mary [22] built a small corpus when they were studying the semantic annotation framework EliXRTIME. Tian [23] also built the corresponding ElilE corpus while developing the inclusion and exclusion criteria for clinical trials (Although its content is limited to Alzheimer's disease, and its dataset is small) and the supporting corpus of BRAT mentioned earlier. There are also researchers who specifically develop corpora that can be used as shared benchmarks by other studies, such as the SUTIME corpus [24] specifically for identifying and normalizing temporal expressions, Chia corpus for machine learning, rule-based or hybrid methods proposed by Fabricio [25], and the LCT corpus [26] recently proposed by Nicholas J.
Patient pre-screening
When a clinical trial faces many participants, manual screening is time-consuming and error prone. Early electronic screening methods were mainly semi-automatic methods, such as developing professional software using the CDSS system of electronic health records, alerting clinicians by e-mail when potential patients are found [27, 28]; using data query to find databases with the assistance of MED software potential patients in and then have the physician manually confirm [29], etc. These methods are still quite time consuming. But with the development of machine learning and the help of artificial intelligence, efficiency will be greatly improved. In Minnesota, for example, in a pilot study at its Mayo Clinic in Rochester, IBM's Watson clinical trial matching system increased the average monthly enrollment in breast cancer trials by 80 percent. How to use computer to quickly and accurately determine patients who meet the inclusion and exclusion criteria is an important direction for the current clinical trial eligibility criteria research.
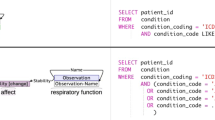
Patient-trial automatic matching is based on a successful application of natural language processing (NLP). The current patient matching research is mainly divided into two categories: project and system. The patient matching project of a clinical trial is divided into two parts in the overall structure: text extraction of patient electronic medical record text clinical trial inclusion and exclusion criteria text; data matching and labeling after extraction is completed. The specific structure is shown in Fig. 2. For current researchers, there are two main technical problems: (1) How to obtain from clinical trial records and patient data. (2) Design a high-precision, high-efficiency data matching model. For the former, clinical trial information extraction has been well developed. Although extracting useful information from patients' electronic medical records remains problematic. Like the eligibility criteria for clinical trials, the content of electronic medical records includes unstructured narrative text and structured coded data [30]. This also means that new annotation standards and new corpora need to be constructed. The latter means that it will be closely linked to various algorithms. For the latter, like NLP, patient matching has gone through a process from machine learning to deep learning to now transfer learning. Kalya [31] proposed a rule-based approach to compare and match structured patient data with a list of eligibility criteria, and Ni [32] used an automated ES algorithm [33] to pre-screen pediatric oncology patients, saving physicians 85–90% workload. Compose and DeepEnroll proposed in recent years are one of the applications of transfer learning. DeepEnroll [34] uses a pre-trained model (BERT) to process test texts, uses a hierarchical embedding model to represent patients’ electronic health records, and the aforementioned Criteria2Query Compared with the accuracy, the accuracy is 7% higher; while Compose [35] uses BERT and CNN for word embedding and semantic acquisition, respectively, and uses multi-granularity medical concept embedding based on learning taxonomy to achieve dynamic patient-trial matching. The final accuracy is as high as 98%. The specific structures of Compose and DeepEnroll are shown in Fig. 3 and Fig. 4. In addition to using EHR data for matching with clinical trials, studies have also been conducted on EMR data. Houssein [36] proposed the medical big data platform EMR2vec, which allows users to match, correlate and query EMR data and clinical trials. Like the NLP in the previous article, there are competitions that release questions related to patient matching every year. For example, the predecessor of n2c2 mentioned above: i2b2 [37], and the TREC text retrieval conference [38].
In addition to patient-matching programs that have been put to the test, websites that automatically match patients to clinical trials are already in use, such as PatientsLikeMe, which requires patients to register and enter relevant information to match clinical trials. In addition, many medical companies have developed their own clinical trial patient matching solutions and put them into the market, such as Circlebase's ACTM solution based on natural language processing and machine learning, CARIS's Caris RIT research system. Unfortunately, it is difficult to know the exact technical details of these solutions because they are commercial products.
Eligibility criteria evaluation
Standard evaluation is to study the influence of the eligibility criteria of clinical trials on the final results, which is consistent with the purpose of earlier research on the content of inclusion and exclusion criteria. The main purpose of the study is to (1) broaden the eligibility criteria, avoid unnecessary exclusions, and benefit more patients (2) develop qualified inclusion and exclusion criteria [39], so that participants in clinical trials can be more in line with the trail, while ensuring safety and efficacy. The impact of eligibility criteria on clinical trials is mainly shown as the number of people recruited to the trial, as well as various parameters used in medicine to evaluate clinical trial results, such as the trial-controlled hazard ratio (HR). In addition to the above two, some people also put forward their own views through their own research, such as the SICE effect proposed by Ma [40], which is suitable for large-scale confirmatory clinical trials, and the GIST2.0 multi-trait metric proposed by Anando [41], which can compute the priori generalizability based on the population representativeness of a clinical study. Early research on eligibility criteria mainly relied on manual comparison and summary. In 2014, when both EHR and NLP in the field of clinical trials became increasingly popular, the research team led by Weng [42] proposed to apply EHR data to the study of clinical trial and eligibility criteria. They used clinical trial data from ClinicalTrials.gov and electronic health records from Columbia University Medical Center of NewYork Presbyterian Hospital to study the differences between the target population of clinical trial and the real-world population. After that, more and more people began to use electronic data resources to analyze clinical trials and eligibility criteria. Nowadays, researchers often need to test the impact of eligibility criteria under different conditions on the results by means of simulation experiments. Since this process needs to be carried out by computer simulations, testing with synthetic data becomes a possible option. But as a research field closely related to evidence-based medicine, real clinical data are very important. At present, researchers in the field of clinical trials mainly use patient data in real-world data (RWD) as research data sources, such as Chen [43] research on Oppenheimer's disease clinical trials, Kim [44] research on COVID-19 clinical trials and Li [45] research on colorectal cancer clinical trials, Li's research also demonstrated the feasibility of using RWD to assess clinical outcomes in patients. In addition to the above studies, others have focused on the optimization of eligibility criteria for clinical trials: Liu [46] from Stanford University developed an open-source artificial intelligence tool called TrialPathfinder that uses EHR data to simulate clinical trials according to different eligibility criteria. Their findings were also striking: Several common inclusion criteria, such as blood pressure, lymphocyte counts, had little effect on trial hazard ratios. However, if it is restricted, it will greatly affect the final number of recruits. Similarly, Liu [47], who proposed the AICO framework, came to similar conclusions when studying several different experimental cases. By widening the thresholds of clinical variables, the coverage of clinical trials can be effectively expanded.
Clinical trial query
The goal of eligibility criteria studies in most clinical trials is to recruit enough patients who meet the trial criteria. To achieve this goal, we need to think from the perspectives of researchers and patients: researchers need to design a reasonable clinical trial plan, and patients need to choose the correct clinical trial. Both steps involve finding a clinical trial that meets the users’ needs from a large amount of past or present clinical trial data.
From the researcher’s point of view, when designing a new clinical trial protocol, the researcher may need to refer to some previous similar cases. To reduce the time and energy spent by researchers in this process, it is obviously a good idea to integrate past clinical trial protocols into a database for systematic review. Such databases would include an NLP model to process the raw textual data, and a user interface (usually a web-based user interface) for the user to use [48]. In addition to data retrieval, knowledge graph is also an important derivative direction. Visualizing the data in the database to the user can speed up the user’s information retrieval, such as the COVID-19 clinical trial knowledge map developed by Jingcheng [49], and the eligibility criteria database with a knowledge map visualization platform developed by Milian [50]. The downside of such databases is that their data accuracy is entirely dependent on the NLP model they use. To ensure a higher level of data accuracy and reusability, extensive testing in collaboration with experienced reviewers and clinicians is required.
From the perspective of patients, it is worth thinking about how to make those patients who are interested in finding a clinical trial that suits their own situation smoothly. With the rapid development of the Internet, today’s patients are more inclined to use the Internet to inquire about clinical trials that may be of interest to them. The conventional query method is to use search engines to search for keywords, and some people have already started research in this area: using FastText, CNN, SVM, KNN and other deep learning networks to test the classification effect of the eligibility criteria [51], identified and classified eligibility criteria in clinical trials by latent Dirichlet allocation (LDA) and logistic regression models in the absence of annotated data [52], or an automated method developed for similar eligibility standard clustering test [53]. Although a search engine that integrates high-precision search methods can make it easier for patients to search, patients may still be faced with finding from hundreds of possible options. This method is time-consuming and labor-intensive, and it is not always possible to find a clinical trial protocol that is suitable for the patient. This situation is very similar to the development of recommendation algorithms and systems. Through unsupervised data mining, Riccardo [54] looks for highly general “labels” from the inclusion and exclusion criteria of clinical trials. On this basis, they proposed the eTACTS system [55]. Targeted at patients and researchers, eTACTS helps users quickly find the clinical trials they need by narrowing down their search by selecting tags from a “tag cloud.” In addition to the static method of tagging, some researchers also consider it from a dynamic point of view such as developing a system that dynamically generates survey questions based on an existing database and then, generates questions based on user responses [56]. These systems greatly save users time, while also making it more likely that clinical trials will recruit eligible patients.
Others
As people increasingly focus on the application of artificial intelligence in clinical trials. There are also many people who are still focusing on some other aspects of research, such as the text specification of eligibility criteria. As early as 2009, when the electronic medical record EHR gradually became popular, Wang [57] studied the standardization of the eligibility criteria. The research lays a solid foundation for future follow-up research. Then, in 2013, the NIH Health Care System Collaborative Laboratories offered their own take on the next generation of clinical trial phenotypic electronic health records [58]: EHR data do not reflect the “real” situation of the patient but reflects the relationship between the patient and the environment, medical care interactions between systems. The same is true for various encoded data in the EHR. Although computers need to convert unstructured sentences into structured sentences, this does not mean that unstructured data are useless. Raghavan [59] studied the importance of unstructured data in clinical trial recruitment. Research on existing disease data shows that structured data alone are not sufficient to address clinical trial recruitment. Structured data need to be further combined with unstructured data to further increase its coverage. Similar to the fact that EHR sometimes fails to reflect the real situation of patients, the eligibility criteria also exist limitations. Averitt [60] studied the eligibility criteria for randomized controlled trials. By comparing the data reported by RCT with the real data of electronic health records of large academic medical centers planned according to RCT qualification criteria, they found that in randomized controlled trials, the qualification criteria may not be sufficient to identify applicable real-world populations.
Dataset
When investigating issues related to clinical trial eligibility criteria, the required datasets fall into two broad categories: clinical trial data and patient data. Researchers obtain eligibility criteria from clinical trial data, and many clinical trials have published their trial content on the Internet. For patient data, due to the extensive development of medical informatization and electronicization, researchers can obtain the required patient data through the patient's electronic health record (EHR).
The sources of clinical trial data used in papers mainly come from the clinicaltrials.gov website, a clinical trial database run by the US National Library of Medicine (NLM) and the US Food and Drug Administration (FDA) and affiliated with the National Institutes of Health (NIH). It is the largest clinical trials registry in the world. The sources of patient EHR data are relatively complex. Except for undisclosed specific sources and data sets provided by conferences or competitions, most of them come from commercial websites or corporate cooperation. For example, Raghavan obtained data from 100 clinical trials, including chronic lymphocytic leukemia and prostate cancer, from the Clinical Trials website. They obtained medical records including 2060 chronic lymphocytic leukemia patients and 1808 prostate cancer patients from The Ohio State University Wexner Medical Center. Table 1 shows the dataset sources for the rest of the reference papers in this paper.
Notably, DeepEnroll researchers have also proposed an automated patient data generator written in Python that can automatically generate patients in batches to meet testing needs. Of course, real patient data are still a must to demonstrate the scientific validity and validity of the findings.
In the early stage of patient data, there was no unified norm and standard, so patient data with complex sources appeared. In recent years, patient data have been gradually unified and standardized. The emergence of large professional databases such as OPTUM and Flatiron has greatly facilitated researchers. At the same time, connecting with these databases has also become one of the ultimate goals of many research projects.
Discussion
Challenges
From the end of the last century to the present, the research on the eligibility criteria of clinical trials has also faced many problems and challenges. During the research process, researchers mainly face three types of problems: interdisciplinary research, research data acquisition and large-scale testing. Since research in this field is closely related to medical clinical trials, understanding, and mastering relevant medical expertise, using scientific and reliable experimental data, and finally conducting a large number of rigorous tests are challenges that must be solved.
Conceptual issues
Before conducting related research work, researchers should understand and be familiar with relevant professional knowledge and concepts of clinical trials, such as interventions, experimental groups and control groups, risk ratios, random blinding, and placebo. The current problem is that many researchers are not fully familiar with the relevant concepts of clinical trials and eligibility criteria when conducting research, which leads to various problems in data processing such as text recognition and text classification, which seriously affects the research.
In addition to the related concepts of clinical trials, the medical concepts themselves are easily confused. Take “osteoporosis” and “osteomalacia” as examples. Osteoporosis is caused by the proportional reduction in bone matrix and bone minerals, and bone resorption is greater than bone formation, which results decrease in bone mass and increase in bone fragility. Osteomalacia refers to no change in the bone matrix due to the decrease in bone mineral mineralization. Many medical terms are not very different in words, but in reality, they are different. Confusion of medical concepts and lack of relevant disease expertise are major problems in the current clinical trials inclusion and exclusion criteria studies.
Dataset problems
The datasets used in the study of eligibility criteria fall into two categories: clinical trial data (ER) and patient electronic medical records (EHR). Patient data are difficult to obtain compared to clinical trial data available from clinicaltrials.gov. Due to personal privacy concerns, it is difficult to obtain free, reliable, open-source patient datasets with a certain sample size on the Internet, except in cooperation with professional institutions. This is the question and challenge that all researchers face.
Another problem with datasets is the uniformity of data specification. Electronic health records may vary from country to country, region to region, or even between departments within the same hospital [61]. Since many papers today involve the study of medical data, how to integrate these data is a problem that must be solved. Although large-scale patient datasets have been widely used and popularized in recent years, and many papers and studies have also begun to actively integrate with these databases. The compatibility of cutting-edge research with different databases has been a new problem, which affects the scalability of research.
Test problems
Whether it is the various clinical trial databases mentioned above or the network questionnaire system. Although they were all tested and released to the public, the main problem they faced was that the number of people who participated in the test was too small, resulting in the findings themselves being very one-sided. While these are helpful for patients who are actively seeking out clinical studies, but not recruiting enough testers remains a serious problem. Only 10 out of 100 trial applicants end up participating in the trial is the normalcy for many clinical trials [62]. From a physician's perspective, the main barriers to clinical trial recruitment are time constraints and the inability of eligible patients to participate in trials for various reasons [3]. It is not only the clinical trial itself that affects the number of recruits, but also economic factors, policy factors, and information asymmetry between the trial and the patients [63]. If the number of participants is not enough, then there will be no practical significance to conduct more studies around the eligibility criteria. The public's attitude toward clinical research is also one of the problems faced by this field. Research in this field, especially those related to clinical trial network platforms and data systems, needs to involve the public in the testing process in addition to real patient data (RWD). In the context of big data, many related studies of artificial intelligence in clinical trials currently lack rigorous large-scale experiments. How to prove that these studies improve clinical trials is one of the major challenges in the field today.
Future outlook
Research in the field of eligibility criteria for clinical trials began as early as the second half of the last century and formed a number of research directions based on natural language recognition.
The current research hotspot in this field—text recognition and information extraction is still an important research direction in the future. As an important foundation for research in the entire field, NLP models with good accuracy are an important prerequisite for most research in this field to be carried out smoothly. In the context of the rapid development of new artificial intelligence technologies such as deep learning and transfer learning, research in the direction of natural language processing (NLP) will continue to be hot. Building NLP suitable for clinical trials based on pre-trained NLP models, building a large corpus sufficient to support other NLP research, and connecting with large clinical databases are all important development directions of NLP in the field of clinical trials in the future. As for patient matching and eligibility criteria evaluation, many projects of the former have now been commercialized and are undergoing trial operation or formal operation in many hospitals and medical institutions and will continue to expand in the future. The latter is not only closely related to the retrospective study of clinical trials, but also closely related to the design of clinical trial qualification standards and personalized medicine. For clinical trial qualification design, in fact, as early as 1999, Daniel [64] have proposed a new tool to write the qualification criteria for clinical trials, but due to the limitations of the times and technology, the research and development in this direction has been in a slower speed. Until recently, the development of various applications and algorithms such as large clinical trial databases, clinical trial data retrieval, provided new possibilities for development in this direction. A large amount of clinical trial data enable people to develop a clinical trial recommendation algorithm in the same way as a commodity recommendation system. If the physician inputs some necessary trial-related data, the system can automatically recommend relevant clinical trials to him as a reference, which helps the physician design eligibility criteria and directly generate relevant templates based on similar cases in the past. For personalized medicine, the development of artificial intelligence in clinical trials has brought new opportunities for it. Data collected in randomized clinical trials can be used to construct individualized treatment rules (ITRs), which can be facilitated by the development of various data management techniques. In the past, building ITRs from large amounts of data was time-consuming and laborious. Today, with the development of artificial intelligence, some researchers are applying active learning to personalized medicine [65, 66]. It reduces the learning cost of ITR and provides more possibilities for the future. In addition to the above two, there will be more large-scale, cross-domain professional clinical trial data platforms in the future. Data visualization, knowledge graphs, and search engines, these calculations closely related to big data will also be closely integrated with clinical trial research and applied.
In addition to the above two categories, there will be more large-scale, cross-disciplinary professional clinical trial data platforms in the future. Building a database for multiple diseases and medical standards is an important research direction in the future. In addition, data visualization, knowledge graphs, search engines, and other technical applications supporting databases are also new research directions.
In short, new research directions will continue to focus on the ultimate goal of “how to help clinical trials.” The “how to help” approach is very diverse and can be any part (or whole) of the clinical trial chain. For researchers, they first need to determine which part of a clinical trial they are studying and how their research will help the clinical trial. The researchers then select the relevant technology and the corresponding data set, and finally conduct the research.
Researchers must collaborate with relevant professional departments in order to obtain valid “real world data.” Although there are many difficulties, the lack of previous research also brings more research directions and research potential.
Conclusion
In summary, this paper systematically analyzes and sorts out the status of studies in clinical trial eligibility criteria. We have selected and studied more than 60 research papers and divided them into four categories: natural language processing, patient matching, evaluation of eligibility criteria, and clinical trial query. The research contents, research methods and research results of these papers are analyzed. The datasets used in each research paper were systematically combed. Models proposed in some computer papers are analyzed. The problems faced in this field are expounded from three perspectives of cross-domain, data acquisition and testing.
The development of this field is closely related to artificial intelligence technology. In the context of the emergence of electronic medical data, machine learning and deep learning allow researchers to process and analyze data more efficiently. In the future, the focus of work will be on practical applications in this field. In addition to the original patient matching and standard evaluation, large-scale applications such as clinical trial personalized medicine, clinical trial database management platform, clinical trial knowledge map, clinical trial retrieval system, and clinical trial recommendation system will be important research objects. We will continue to research how computer technologies, led by artificial intelligence, can impact clinical trials in real life.
References
Marcus W. Trial by artificial intelligence a combination of big data and machine-learning algorithms could help to accelerate clinical testing. Nature. 2019;573
Rahman M, Morita S, Fukui T, Sakamoto J. Physicians reasons for not entering their patients in a randomized controlled trial in Japan. Tohoku J Exp Med. 2004;203(2):105109. https://doi.org/10.1620/tjem.203.105.
Spaar A, Frey M, Turk A, Karrer W, Puhan MA. Recruitment barriers in a randomized controlled trial from the physicians’ perspective–a postal survey. BMC Med Res Methodol. 2009;9(1):1–8.
Acharya S, et al. The COVID-19 pandemic: theories to therapies. Adv Infect Dis. 2020;10(03):16.
Tu SW, Peleg M, Carini S, Bobak M, Ross J, Rubin D, Sim I. A practical method for transforming free-text eligibility criteria intocomputable criteria. J Biomed Inform. 2011;44(2):239–50.
Weng C, Wu X, Luo Z, Boland MR, Theodoratos D, Johnson SB. EliXR: an approach to eligibility criteria extraction and representation. J Am Med Inform Assoc. 2011;8(Supplement_1):i116–24.
Yuan C, Ryan PB, Ta C, Guo Y, Li Z, Hardin J, Makadia R, Jin P, Shang N, Kang T, et al. Criteria2query: a natural language interface to clinical databases for cohort definition. J Am Med Inform Assoc. 2019;26(4):294–305.
Tseo Y, Salkola M, Mohamed A, Kumar A, Abnousi F. Information extraction of clinical trial eligibility criteria. 2020. arXiv preprint http://arxiv.org/2006.07296.
Pandey C, Ibrahim Z, Wu H, Iqbal E, Dobson R. Improving RNN with attention and embedding for adverse drug reactions. In: Proceedings of the 2017 international conference on digital health, 2017. pp. 67–71.
Lee J, Yoon W, Kim S, Kim D, Kim S, So CH, Kang J. BioBERT: a pre-trained biomedical language representation model for biomedical text mining. Bioinformatics. 2020;36(4):1234–40.
Raj Kanakarajan K, Kundumani B, Sankarasubbu M. BioELECTRA: pretrained biomedical text encoder using discriminators: In: Proceedings of the 20th workshop on biomedical language processing, 2021, pp. 143–4.
Naseem U, Dunn AG, Khushi M, Kim J. Benchmarking forbiomedical natural language processing tasks with a domain specific albert. BMC Bioinform. 2022;23(1):1–15.
Liu X, Hersch GL, Khalil I, Devarakonda M. Clinical trial information extraction with Bert. In: 2021 IEEE 9th international conference on healthcare informatics (ICHI). IEEE, 2021, pp. 505–6.
Gu Y, Tinn R, Cheng H, Lucas M, Usuyama N, Liu X, Naumann T, Gao J, Poon H. Domain-specific language model pretraining for biomedical natural language processing. ACM Trans Comput Healthc (HEALTH). 2021;3(1):1–23.
Shi J, Graves K, Hurdle JF. A generic rule-based system for clinical trial patient selection. 2019. arXiv preprint http://arxiv.org/1907.06860
Wang Y, Fu S, Shen F, Henry S, Uzuner O, Liu H, et al. The 2019 n2c2/ohnlp track on clinical semantic textual similarity: overview. JMIR Med Inform. 2020;8(11): e23375.
Hahn U, Oleynik M. Medical information extraction in the age of deep learning. Yearb Med Inform. 2020;29(01):208–20.
Sheikhalishahi S, Miotto R, Dudley JT, Lavelli A, Rinaldi F, Osmani V, et al. Natural language processing of clinical notes on chronic diseases: systematic review. JMIR Med Inform. 2019;7(2): e12239.
Datta S, Bernstam EV, Roberts K. A frame semantic overview of NLP-based information extraction for cancer-related EHR notes. J Biomed Inform. 2019;100: 103301.
Bodenreider O. The unified medical language system (UMLS): integrating biomedical terminology. Nucleic acids Res. 2004;32(suppl_1):D267–70.
Stenetorp P, Pyysalo S, Topić G, Ohta T, Ananiadou S, Tsujii J. BRAT: a web-based tool for NLP-assisted text annotation. In: Proceedings of the demonstrations at the 13th conference of the European chapter of the association for computational linguistics, 2012. pp. 102–7.
Boland MR, Tu SW, Carini S, Sim I, Weng C. Elixr-time: a temporal knowledge representation for clinical research eligibility criteria. AMIA Summits Trans Sci Proc. 2012;2012:71.
Kang T, Zhang S, Tang Y, Hruby GW, Rusanov A, Elhadad N, Weng C. EliiE: an open-source information extraction system for clinical trial eligibility criteria. J Am Med Inform Assoc. 2017;24(6):1062–71.
Chang AX, Manning CD. Sutime: a library for recognizing and normalizing time expressions. In: Lrec, 2012; 3735: 3740.
Kury F, Butler A, Yuan C, Fu L-H, Sun Y, Liu H, Sim I, Carini S, Weng C. Chia, a large annotated corpus of clinical First Author et al.: preprint submitted to Elsevier Page 8 of 10 trial eligibility criteria. Scientific data. 2020;7(1):1–11.
Dobbins NJ, Mullen T, Uzuner Ö, Yetisgen M. The leaf clinical trials corpus: a new resource for query generation from clinical trial eligibility criteria. Sci Data. 2022;9(1):1–15.
Embi PJ, Jain A, Clark J, Bizjack S, Hornung R, Harris CM. Effect of a clinical trial alert system on physician participation in trial recruitment. Arch Intern Med. 2005;165(19):2272–7.
Embi PJ, Jain A, Harris CM. Physician perceptions of an electronic health record-based clinical trial alert system: a survey of study participants. In: AMIA Annual symposium proceedings. American Medical Informatics Association, 2005;2005:949.
Thadani SR, Weng C, Bigger JT, Ennever JF, Wajngurt D. Electronic screening improves efficiency in clinical trial recruitment. J Am Med Inform Assoc. 2009;16(6):869–73.
Van Spall HG, Toren A, Kiss A, Fowler RA. Eligibility criteria of randomized controlled trials published in high-impact general medical journals: a systematic sampling review. JAMA. 2007;297(11):1233–40.
Adupa AK, Garg RP, Corona-Cox J, Shah S, Jonnalagadda SR, et al. An information extraction approach to prescreen heart failure patients for clinical trials. 2016. arXiv preprint http://arxiv.org/1609.01594.
Ni Y, Wright J, Perentesis J, Lingren T, Deleger L, Kaiser M, Kohane I, Solti I. Increasing the efficiency of trial-patient matching: automated clinical trial eligibility pre-screening for pediatric oncology patients. BMC Med Inform Decis Mak. 2015;15(1):1–10.
Ni Y, Kennebeck S, Dexheimer JW, McAneney CM, Tang H, Lingren T, Li Q, Zhai H, Solti I. Automated clinical trial eligibility prescreening: increasing the efficiency of patient identification for clinical trials in the emergency department. J Am Med Inform Assoc. 2015;22(1):166–78.
Zhang X, Xiao C, Glass LM, Sun J. DeepEnroll: patient-trial matching with deep embedding and entailment prediction. In:Proceedings of the web conference 2020, 2020. pp. 1029–37
Gao J, Xiao C, Glass LM, Sun L. Compose: cross-modal pseudo-siamese network for patient trial matching. In: Proceedings of the 26th ACM SIGKDD international conference on knowledge discovery & data mining, 2020. pp. 803–12.
Dhayne H, Kilany R, Haque R, Taher Y. Emr2vec: Bridging the gap between patient data and clinical trial. Comput Ind Eng. 2021;156: 107236.
Murphy SN, Weber G, Mendis M, Gainer V, Chueh HC, Churchill S, Kohane I. Serving the enterprise and beyond with informatics for integrating biology and the bedside (i2b2). J Am Med Inform Assoc. 2010;17(2):124–30.
Truong TH, Otmakhova Y, Mahendra R, Baldwin T, Lau JH, Cohn T, Cavedon L, Spina D, Verspoor K. Ittc@ trec 2021 clinical trials track. 2022. arXiv preprint http://arxiv.org/2202.07858.
U.S. Department of Health and Human Services, Food and Drug Administration, Center for Drug Evaluation and Research (CDER), Center for Biologics Evaluation and Research (CBER). Enhancing the Diversity of Clinical Trial Populations——Eligibility Criteria, Enrollment Practices, and Trial Designs Guidance for Industry, 2020.
Ma J, Holder DJ. Selection induced contrast estimate (SICE) effect: an attempt to quantify the impact of some patient selection criteria in randomized clinical trials. 2020. arXiv preprint http://arxiv.org/2001.02036.
Sen A, Chakrabarti S, Goldstein A, Wang S, Ryan PB, Weng C. Gist 2.0: a scalable multi-trait metric for quantifying population representativeness of individual clinical studies. J Biomed Inform. 2016;63:325–36.
Weng C, Li Y, Ryan P, Zhang Y, Liu F, Gao J, Bigger J, Hripcsak G. A distribution-based method for assessing the differences between clinical trial target populations and patient populations in electronic health records. Appl Clin Inform. 2014;5(02):463–79.
Chen Z, Zhang H, Guo Y, George TJ, Prosperi M, Hogan WR, He Z, Shenkman EA, Wang F, Bian J. Exploring the feasibility of using real-world data from a large clinical data research network to simulate clinical trials of Alzheimer’s disease. NPJ Digit Med. 2021;4(1):1–9.
Kim JH, Ta CN, Liu C, Sung C, Butler AM, Stewart LA, Ena L, Rogers JR, Lee J, Ostropolets A, et al. Towards clinical data-driven eligibility criteria optimization for interventional COVID-19 clinical trials. J Am Med Inform Assoc. 2021;28(1):14–22.
Li Q, He Z, Guo Y, Zhang H, George TJ, Hogan W, Charness N, Bian J. Assessing the validity of a a priori patient-trial generalizability score using real-world data from a large clinical data research network: a colorectal cancer clinical trial case study. In: AMIA annual symposium proceedings. American Medical Informatics Association, 2019, 2019. p. 1101.
Liu R, Rizzo S, Whipple S, Pal N, Pineda AL, Lu M, Arnieri B, Lu Y, Capra W, Copping R, et al. Evaluating eligibility criteria of oncology trials using real-world data and AI. Nature. 2021;592(7855):629–33.
Liu X, Shi C, Deore U, Wang Y, Tran M, Khalil I, Devarakonda M. A scalable AI approach for clinical trial cohort optimization: In: Joint European conference on machine learning and knowledge discovery in databases. Springer, 2021. pp. 479–9.
Liu H, Chi Y, Butler A, Sun Y, Weng C. A knowledge base of clinical trial eligibility criteria. J Biomed Inform. 2021;117: 103771.
Du J, Wang Q, Wang J, Ramesh P, Xiang Y, Jiang X, Tao C. COVID-19 trial graph: a linked graph for COVID-19 clinical trials. J Am Med Inform Assoc. 2021;28(9):1964–9.
Milian K, Hoekstra R, Bucur A, Ten Teije A, van Harmelen F, Paulissen J. Enhancing reuse of structured eligibility criteria and supporting their relaxation. J Biomed Inform. 2015;56:205–19.
Yacoumatos C, Bragaglia S, Kanakia A, Svangård N, Mangion J, Donoghue C, Weatherall J, Khan FM, Shameer K. TrialGraph: Machine intelligence enabled insight from graph modelling of clinical trials. 2021. arXiv preprint http://arxiv.org/2112.08211.
Restificar A, Korkontzelos I, Ananiadou S. A method for discovering and inferring appropriate eligibility criteria in clinical trial protocols without labeled data. In: BMC medical informatics and decision making. BioMed Central, 2013:13 1;1–12.
Hao T, Rusanov A, Boland MR, Weng C. Clustering clinical trials with similar eligibility criteria features. J Biomed Inform. 2014;52:112–20.
Miotto R, Weng C. Unsupervised mining of frequent tags for clinical eligibility text indexing. J Biomed Inform. 2013;46(6):1145–51.
Miotto R, Jiang S, Weng C. eTACTS: a method for dynamically filtering clinical trial search results. J Biomed Inform. 2013;46(6):1060–7.
Liu C, Yuan C, Butler AM, Carvajal RD, Li ZR, Ta CN, Weng C. DQueST: dynamic questionnaire for search of clinical trials. J Am Med Inform Assoc. 2019;26(11):1333–43.
Weng C, Tu SW, Sim I, Richesson R. Formal representation of eligibility criteria: a literature review. J Biomed Inform. 2010;43(3):451–67.
Richesson RL, Hammond WE, Nahm M, Wixted D, Simon GE, Robinson JG, Bauck AE, Cifelli D, Smerek MM, Dickerson J, et al. Electronic health records based phenotyping in next-generation clinical trials: a perspective from the NIH health care systems collaboratory. J Am Med Inform Assoc. 2013;20(e2):e226–31.
Raghavan P, Chen JL, Fosler-Lussier E, Lai AM. How essential are unstructured clinical narratives and information fusion to clinical trial recruitment? AMIA Summits Transl Sci Proc. 2014;2014:218.
Averitt AJ, Weng C, Ryan P, Perotte A. Translating evidence into practice: eligibility criteria fail to eliminate clinically significant differences between real-world and study populations. NPJ Digit Med. 2020;3(1):1–10.
Häyrinen K, Saranto K, Nykänen P. Definition, structure, content, use and impacts of electronic health records: a review of the research literature. Int J Med Inform. 2008;77(5):291–304.
Reynolds T. Clinical trials: can technology solve the problem of low recruitment? BMJ. 2011. https://doi.org/10.1136/bmj.d3662.
Li Y, Slivkins A. Incentivizing participation in clinical trials. 2022. arXiv preprint http://arxiv.org/2202.06191.
Rubin DL, Gennari JH, Srinivas S, Yuen A, Kaizer H, Musen MA, Silva JS. Tool support for authoring eligibility criteria for cancer trials. In: Proceedings of the AMIA Symposium. American Medical Informatics Association, 1999. p. 369.
Minsker S, Zhao Y-Q, Cheng G. Active clinical trials for personalized medicine. J Am Stat Assoc. 2016;111(514):875–87.
Deng K, Pineau J, Murphy S. Active learning for personalizingtreatment. In: 2011 IEEE symposium on adaptive dynamic programming and reinforcement learning (ADPRL). IEEE, 2011. pp. 32–39
Funding
This work was supported by Science and Technology Innovation 2030 – Major Project of "New Generation Artistic Intelligence (2020AAAA0109300)".
Author information
Authors and Affiliations
Contributions
Q.S. conceived the idea. Q.S. and J.H. designed the study. G.C. did the analyses. Q.S. and G.C. wrote the main manuscript text. Q.S. prepared Table 1. G.C. prepared Figs. 1, 2, 3, 4. J.H. provide expertise and guidance in clinical medicine. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
All authors report no conflicts of interest in this work.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Su, Q., Cheng, G. & Huang, J. A review of research on eligibility criteria for clinical trials. Clin Exp Med 23, 1867–1879 (2023). https://doi.org/10.1007/s10238-022-00975-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10238-022-00975-1