Abstract
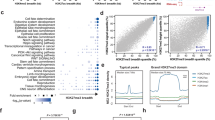
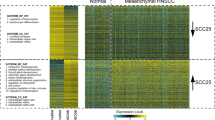
Distinct histone modifications regulate gene expression in certain diseases, but little is known about histone epigenetics in middle ear cholesteatoma. It is known that histone acetylation destabilizes the nucleosome and chromatin structure and induces gene activation. The association of histone acetylation with chronic inflammatory diseases has been indicated in recent studies. In this study, we examined the localization of variously modified histone H3 acetylation at lysine 9, 14, 18, 23, and 27 in paraffin-embedded sections of human middle ear cholesteatoma (cholesteatoma) tissues and the temporal bones of an animal model of cholesteatoma immunohistochemically. As a result, we found that there was a significant increase of the expression levels of H3K27ac both in human cholesteatoma tissues and the animal model. In genetics, super-enhancers are clusters of enhancers that drive the transcription of genes involved in cell identity. Super-enhancers were originally defined using the H3K27ac signal, and then we used H3K27ac chromatin immunoprecipitation followed by sequencing to map the active cis-regulatory landscape in human cholesteatoma. Based on the results, we identified increased H3K27ac signals as super-enhancers of the FOXC2 loci, as well as increased protein of FOXC2 in cholesteatoma. Recent studies have indicated that menin-MLL inhibitor could suppress tumor growth through the control of histone H3 modification. In this study, we demonstrated that the expression of FOXC2 was inhibited by menin-MLL inhibitor in vivo. These findings indicate that FOXC2 expression under histone modifications promoted the pathogenesis of cholesteatoma and suggest that it may be a therapeutic target of cholesteatoma.






Similar content being viewed by others
Availability of Data and Material
The datasets produced and/or analyzed during the current study are available from the corresponding author upon reasonable request.
Code Availability
The study uses data obtained from the UCSC genome browser http://genome.ucsc.edu/index.html.
References
Akiyama N, Yamamoto-Fukuda T, Takahashi H (2014) Influence of continuous negative pressure in the rat middle ear. Laryngoscope 124:2404–2410. https://doi.org/10.1002/lary.24767
Bi Y, Guo S, Xu X, Kong P, Cui H, Yan T, Ma Y, Cheng Y, Chen Y, Liu X, Zhang L, Cheng C, Xu E, Qian Y, Yang J, Song B, Li H, Wang F, Hu X, Liu X, Niu X, Zhai Y, Liu J, Li Y, Cheng X, Cui Y (2020) Decreased ZNF750 promotes angiogenesis in a paracrine manner via activating DANCR/miR-4707-3p/FOXC2 axis in esophageal squamous cell carcinoma. Cell Death Dis 11:296. https://doi.org/10.1038/s41419-020-2492-2
Borkin D, He S, Miao H, Kempinska K, Pollock J, Chase J, Purohit T, Malik B, Zhao T, Wang J, Wen B, Zong H, Jones M, Danet-Desnoyers G, Guzman ML, Talpaz M, Bixby DL, Sun D, Hess JL, Muntean AG, Maillard I, Cierpicki T, Grembecka J (2015) Pharmacologic inhibition of the menin-MLL interaction blocks progression of MLL leukemia in vivo. Cancer Cell 27:589–602. https://doi.org/10.1016/j.ccell.2015.02.016
Bowers EM, Yan G, Mukherjee C, Orry A, Wang L, Holbert MA, Crump NT, Hazzalin CA, Liszczak G, Yuan H, Larocca C, Saldanha SA, Abagyan R, Sun Y, Meyers DJ, Marmorstein R, Mahadevan LC, Alani RM, Cole PA (2010) Virtual ligand screening of the p300/CBP histone acetyl transferase: identification of a selective small molecule inhibitor. Chem Biol 17:471–482. https://doi.org/10.1016/j.chembiol.2010.03.006
Braun S, Krampert M, Bodó E, Kümin A, Born-Berclaz C, Paus R, Werner S (2006) Keratinocyte growth factor protects epidermis and hair follicles from cell death induced by UV irradiation, chemotherapeutic or cytotoxic agents. J Cell Sci 119:4841–4849. https://doi.org/10.1242/jcs.03259
Calo E, Wysocka J (2013) Modification of enhancer chromatin: what, how, and why? Mol Cell 49:825–837. https://doi.org/10.1016/j.molcel.2013.01.038
Cederberg A, Grønning LM, Ahrén B, Taskén K, Carlsson P, Enerbäck S (2001) FOXC2 is a winged helix gene that counteracts obesity, hypertriglyceridemia, andiet-induced insulin resistance. Cell 106:563–573. https://doi.org/10.1016/s0092-8674(01)00474-3
Chapuy B, McKeown MR, Lin CY, Monti S, Roemer MGM, Qi J, Rahl PB, Sun HH, Yeda KT, Doench JG, Reichert E, Kung AL, Rodig SJ, Young RA, Shipp MA, Bradner JE (2013) Discovery and characterization of super-enhancer-associated dependencies in diffuse large b cell lymphoma. Cancer Cell 24(6):777-90. https://doi.org/10.1016/j.ccr.2013.11.003
Chen Y, Deng G, Fu Y, Han Y, Guo C, Yin L, Cai C, Shen H, Wu S, Zeng S (2020) FOXC2 promotes oxaliplatin resistance by inducing epithelial-mesenchymal transition via MAPK/ERK signaling in colorectal cancer. Onco Targets Ther 13:1625–1635. https://doi.org/10.2147/OTT.S241367
Chi Z, Wang Z, Liang Q, Zhu Y, du Q, (2015) Induction of cytokine production in cholesteatoma keratinocytes by extracellular high-mobility group box chromosomal protein 1 combined with DNA released by apoptotic cholesteatoma keratinocytes. Mol Cell Biochem 400:189–200. https://doi.org/10.1007/s11010-014-2275-0
Creyghton MP, Cheng AW, Welstead GG, Kooistra T, Carey BW, Steine EJ, Hanna J, Lodato MA, Frampton GM, Sharp PA, Boyer LA, Young RA, Jaenisch R (2010) Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc Natl Acad Sci U S A 107:21931–21936. https://doi.org/10.1073/pnas.1016071107
Crish JF, Eckert RL (2008) Synergistic activation of human involucrin gene expression by Fra-1 and p300–evidence for the presence of a multiprotein complex. J Invest Dermatol 128:530–541. https://doi.org/10.1038/sj.jid.5701049
Cui YM, Jiao HL, Ye YP, Chen CM, Wang JX, Tang N, Li TT, Lin J, Qi L, Wu P, Wang SY, He MR, Liang L, Bian XW, Liao WT, Ding YQ (2015) FOXC2 promotes colorectal cancer metastasis by directly targeting MET. Oncogene 34:4379–4390. https://doi.org/10.1038/onc.2014.368
Dawson MA, Kouzarides T (2012) Cancer epigenetics: from mechanism to therapy. Cell 150:12–27. https://doi.org/10.1016/j.cell.2012.06.013
Djebali S, Davis CA, Merkel A et al (2012) Landscape of transcription in human cells. Nature 489:101–108. https://doi.org/10.1038/nature11233
Dreijerink KMA, Groner AC, Vos ESM, Font-Tello A, Gu L, Chi D, Reyes J, Cook J, Lim E, Lin CY, de Laat W, Rao PK, Long HW, Brown M (2017) Enhancer-mediated oncogenic function of the menin tumor suppressor in breast cancer. Cell Rep 18:2359–2372. https://doi.org/10.1016/j.celrep.2017.02.025
Ernst J, Kheradpour P, Mikkelsen TS, Shoresh N, Ward LD, Epstein CB, Zhang X, Wang L, Issner R, Coyne M, Ku M, Durham T, Kellis M, Bernstein BE (2011) Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473:43–49. https://doi.org/10.1038/nature09906
Ficz G, Hore TA, Santos F, Lee HJ, Dean W, Arand J, Krueger F, Oxley D, Paul YL, Walter J, Cook SJ, Andrews S, Branco MR, Reik W (2013) FGF signaling inhibition in ESCs drives rapid genome-wide demethylation to the epigenetic ground state of pluripotency. Cell Stem Cell 13:351–359. https://doi.org/10.1016/j.stem.2013.06.004
Finch PW, Cunha GR, Rubin JS, Wong J, Ron D (1995) Pattern of keratinocyte growth factor and keratinocyte growth factor receptor expression during mouse fetal development suggests a role in mediating morphogenetic mesenchymal-epithelial interactions. Dev Dyn 203:223–240. https://doi.org/10.1002/aja.1002030210
Heintzman ND, Stuart RK, Hon G, Fu Y, Ching CW, Hawkins RD, Barrera LO, Van Calcar S, Qu C, Ching KA, Wang W, Weng Z, Green RD, Crawford GE, Ren B (2007) Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat Genet 39:311–318. https://doi.org/10.1038/ng1966
Hnisz D, Abraham BJ, Lee TI, Lau A, Saint-Andre V, Sigova AA, Hoke HA, Young RA (2013) Super-enhancers in the control of cell identity and disease. Cell 155:934–947. https://doi.org/10.1016/j.cell.2013.09.053
Hollier BG, Tinnirello AA, Werden SJ, Evans KW, Taube JH, Sarkar TR, Sphyris N, Shariati M, Kumar SV, Battula VL, Herschkowitz JI, Guerra R, Chang JT, Miura N, Rosen JM, Mani SA (2013) FOXC2 expression links epithelial-mesenchymal transition and stem cell properties in breast cancer. Cancer Res 73:1981–1992. https://doi.org/10.1158/0008-5472.CAN-12-2962
Iida K, Koseki H, Kakinuma H, Kato N, Mizutani-Koseki Y, Ohuchi H, Yoshioka H, Noji S, Kawamura K, Kataoka Y, Ueno F, Taniguchi M, Yoshida N, Sugiyama T, Miura N (1997) Essential roles of the winged helix transcription factor MFH-1 in aortic arch patterning and skeletogenesis. Development 124:4627–4638
Imayama N, Yamada S, Yanamoto S, Naruse T, Matsushita Y, Takahashi H, Seki S, Fujita S, Ikeda T, Umeda M (2015) FOXC2 expression is associated with tumor proliferation and invasion potential in oral tongue squamous cell carcinoma. Pathol Oncol Res 21:783–791. https://doi.org/10.1007/s12253-014-9891-6
Ito K, Ito M, Elliott WM, Cosio B, Caramori G et al (2005) Decreased histone deacetylase activity in chronic obstructive pulmonary disease. N Engl J Med 352:1967–1976. https://doi.org/10.1056/NEJMoa041892
Jiang W, Pang XG, Wang Q, Shen YX, Chen XK, Xi JJ (2012) Prognostic role of Twist, Slug, and Foxc2 expression in stage I non-small-cell lung cancer after curative resection. Clin Lung Cancer 13:280–287. https://doi.org/10.1016/j.cllc.2011.11.005
Kanzaki-Kato N, Tamakoshi T, Fu Y, Chandra A, Itakura T, Uezato T, Tanaka T, Clouthier DE, Sugiyama T, Yanagisawa M, Miura N (2005) Roles of forkhead transcription factor Foxc2 (MFH-1) and endothelin receptor A in cardiovascular morphogenesis. Cardiovasc Res 65:711–718. https://doi.org/10.1016/j.cardiores.2004.10.017
Kempinska K, Malik B, Borkin D, Klossowski S, Shukla S, Miao H, Wang J, Cierpicki T, Grembecka J (2018) Pharmacologic Inhibition of the Menin-MLL interaction leads to transcriptional repression of PEG10 and blocks hepatocellular carcinoma. Mol Cancer Ther 17:26–38. https://doi.org/10.1158/1535-7163.MCT-17-0580
Kouwenhoven EN, Oti M, Niehues H, van Heeringen SJ, Schalkwijk J, Stunnenberg HG, van Bokhoven H, Zhou H (2015) Transcription factor p63 bookmarks and regulates dynamic enhancers during epidermal differentiation. EMBO Rep 16:863–878. https://doi.org/10.15252/embr.201439941
Kume T, Jiang H, Topczewska JM, Hogan BL (2001) Themurine winged helix transcription factors, Foxc1 and Foxc2, are both required for cardiovascular development and somitogenesis. Genes Dev 15:2470–2482. https://doi.org/10.1101/gad.907301
Kuo CL (2015) Etiopathogenesis of acquired cholesteatoma: prominent theories and recent advances in biomolecular research. Laryngoscope 125:234–240. https://doi.org/10.1002/lary.24890
Lee JS, See RH, Deng T, Shi Y (1996) Adenovirus E1A downregulates cJun- and JunB-mediated transcription by targeting their coactivator p300. Mol Cell Biol 16:4312–4326. https://doi.org/10.1128/mcb.16.8.4312
Lee K, Hsiung CC, Huang P, Raj A, Blobel GA (2015) Dynamic enhancer-gene body contacts during transcription elongation. Genes Dev 29:1992–1997. https://doi.org/10.1101/gad.255265.114
Li D, Yan D, Liu W, Li M, Yu J, Li Y, Qu Z, Ruan Q (2011) Foxc2 overexpression enhances benefit of endothelial progenitor cells for inhibiting neointimal formation by promoting CXCR4-dependent homing. J Vasc Surg 53:1668–1678. https://doi.org/10.1016/j.jvs.2011.01.044
Li QL, Wang DY, Ju LG, Yao J, Gao C, Lei PJ, Li LY, Zhao XL, Wu M (2019) The hyper-activation of transcriptional enhancers in breast cancer. Clin Epigenetics 11:48. https://doi.org/10.1186/s13148-019-0645-x
Li W, Fu X, Liu R, Wu C, Bai J, Xu Y, Zhao Y, Xu Y (2013) FOXC2 often overexpressed in glioblastoma enhances proliferation and invasion in glioblastoma cells. Oncol Res 21:111–120. https://doi.org/10.3727/096504013X13814233062171
Lovén J, Hoke HA, Lin CY, Lau A, Orlando DA, Vakoc CR, Bradner JE, Lee TI, Young RA (2013) Selective inhibition of tumor oncogenes by disruption of super-enhancers. Cell 153:320–334. https://doi.org/10.1016/j.cell.2013.03.036
Malik R, Khan AP, Asangani IA, Cieślik M, Prensner JR, Wang X, Iyer MK, Jiang X, Borkin D, Escara-Wilke J, Stender R, Wu YM, Niknafs YS, Jing X, Qiao Y, Palanisamy N, Kunju LP, Krishnamurthy PM, Yocum AK, Mellacheruvu D, Nesvizhskii AI, Cao X, Dhanasekaran SM, Feng FY, Grembecka J, Cierpicki T, Chinnaiyan AM (2015) Targeting the MLL complex in castration-resistant prostate cancer. Nat Med 21:344–352. https://doi.org/10.1038/nm.3830.
Mani SA, Yang J, Brooks M, Schwaninger G, Zhou A, Miura N, Kutok JL, Hartwell K, Richardson AL, Weinberg RA (2007) Mesenchyme Forkhead 1 (FOXC2) plays a key role in metastasis and is associated with aggressive basal-like breast cancers. Proc Natl Acad Sci USA 104:10069–10074. https://doi.org/10.1073/pnas.0703900104
Matsumoto K, Nagayasu T, Hishikawa Y, Tagawa T, Yamayoshi T, Abo T, Tobinaga S, Furukawa K, Koji T (2009) Keratinocyte growth factor accelerates compensatory growth in the remaining lung after trilobectomy in rats. J Thorac Cardiovasc Surg 137:1499–1507. https://doi.org/10.1016/j.jtcvs.2008.11.037
Milne TA, Hughes CM, Lloyd R, Yang Z, Rozenblatt-Rosen O, Dou Y, Schnepp RW, Krankel C, Livolsi VA, Gibbs D, Hua X, Roeder RG, Meyerson M, Hess JL (2005) Menin and MLL cooperatively regulate expression of cyclin-dependent kinase inhibitors. Proc Natl Acad Sci U S A 102:749–754. https://doi.org/10.1073/pnas.0408836102
Nishida N, Mimori K, Yokobori T, Sudo T, Tanaka F, Shibata K, Ishii H, Doki Y, Mori M (2011) FOXC2 is a novel prognostic factor in human esophageal squamous cell carcinoma. Ann Surg Oncol 18:535–542. https://doi.org/10.1245/s10434-010-1274-y
Otsuka-Tanaka Y, Oommen S, Kawasaki M, Kawasaki K, Imam N, Jalani-Ghazani F, Hindges R, Sharpe PT, Ohazama A (2013) Oral lining mucosa development depends on mesenchymal microRNAs. J Dent Res 92:229–234. https://doi.org/10.1177/0022034512470830
Pan K, Xie Y (2020) LncRNA FOXC2-AS1 enhances FOXC2 mRNA stability to promote colorectal cancer progression via activation of Ca (2+)-FAK signal pathway. Cell Death Dis 11:434. https://doi.org/10.1038/s41419-020-2633-7
Peluso S, Douglas A, Hill A, De Angelis C, Moore BL, Grimes G, Petrovich G, Essafi A, Hill RE (2017) Fibroblast growth factors (FGFs) prime the limb specific Shh enhancer for chromatin changes that balance histone acetylation mediated by E26 transformation-specific (ETS) factors. Elife ;6:e28590. https://doi.org/10.7554/eLife.28590
Raisner R, Kharbanda S, Jin L, Jeng E, Chan E, Merchant M, Haverty PM, Bainer R, Cheung T, Arnott D, Flynn EM, Romero FA, Magnuson S, Gascoigne KE (2018) Enhancer activity requires CBP/P300 bromodomain-dependent histone H3K27 acetylation. Cell Rep 24:1722–1729. https://doi.org/10.1016/j.celrep.2018.07.041
Sagiv D, Harari-Steinberg O, Wolf M, Dekel B, Omer D (2019) The feasibility to isolate and expand tympanic membrane squamous epithelium stem cells from scarred perforation margins. Otol Neurotol 40:e1030–e1036. https://doi.org/10.1097/MAO.0000000000002367
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9:671–675. https://doi.org/10.1038/nmeth.2089
Schuettengruber B, Chourrout D, Vervoort M, Leblanc B, Cavalli G (2007) Genome regulation by polycomb and trithorax proteins. Cell 128:735–745. https://doi.org/10.1016/j.cell.2007.02.009
Seo S, Kume T (2006) Forkhead transcription factors, Foxc1 and Foxc2, are required for the morphogenesis of the cardiac outflow tract. Dev Biol 296:421–436. https://doi.org/10.1016/j.ydbio.2006.06.012
Sarkar PL, Lee W, Williams ED, Lubik AA, Stylianou N, Shokoohmand A, Lehman ML, Hollier BG, Gunter JH, Nelson CC (2019) Insulin enhances migration and invasion in prostate cancer cells by up-regulation of FOXC2. Front Endocrinol (Lausanne) 10:481. https://doi.org/10.3389/fendo.2019.00481
Shi A, Murai MJ, He S, Lund G, Hartley T, Purohit T, Reddy G, Chruszcz M, Grembecka J, Cierpicki T (2012) Structural insights into inhibition of the bivalent menin-MLL interaction by small molecules in leukemia. Blood 120:4461–4469. https://doi.org/10.1182/blood-2012-05-429274
Song N, Liu J, An S, Nishino T, Hishikawa Y, Koji T (2011) Immunohistochemical analysis of histone H3 modifications in germ cells during mouse spermatogenesis. Acta Histochem Cytochem 44:183–190. https://doi.org/10.1267/ahc.11027
Sudhoff H, Tos M (2000) Pathogenesis of attic cholesteatoma: clinical and immunohistochemical support for combination of retraction theory and proliferation theory. Am J Otol 21:786–792
Takahashi M, Yamamoto-Fukuda T, Akiyama N, Motegi M, Yamamoto K, Tanaka Y, Yamamoto Y, Kojima H (2019) Partial epithelial-mesenchymal transition was observed under p63 expression in acquired middle ear cholesteatoma and congenital cholesteatoma. Otol Neurotol 40:e803–e811. https://doi.org/10.1097/MAO.0000000000002328
Tambalo M, Anwar M, Ahmed M, Streit A (2020) Enhancer activation by FGF signalling during otic induction. Dev Biol 457:69–82. https://doi.org/10.1016/j.ydbio.2019.09.006
Tanaka Y, Kojima H, Miyazaki H, Koga T, Moriyama H (1999) Roles of cytokines and cell cycle regulating substances in proliferation of cholesteatoma epithelium. Laryngoscope 109:1102–1107. https://doi.org/10.1097/00005537-199907000-00017
Tie F, Banerjee R, Stratton CA, Prasad-Sinha J, Stepanik V, Zlobin A, Diaz MO, Scacheri PC, Harte PJ (2009) CBP-mediated acetylation of histone H3 lysine 27 antagonizes Drosophila Polycomb silencing. Development 136:3131–3141. https://doi.org/10.1242/dev.037127
Vijayakrishnan J, Qian M, Studd JB, Yang W, Kinnersley B, Law PJ, Broderick P, Raetz EA, Allan J, Pui CH, Vora A, Evans WE, Moorman A, Yeoh A, Yang W, Li C, Bartram CR, Mullighan CG, Zimmerman M, Hunger SP, Schrappe M, Relling MV, Stanulla M, Loh ML, Houlston RS, Yang JJ (2019) Identification of four novel associations for B-cell acute lymphoblastic leukaemia risk. Nat Commun 10:5348. https://doi.org/10.1038/s41467-019-13069-6
Wang Z, Zang C, Rosenfeld JA, Schones DE, Barski A, Cuddapah S, Cui K, Roh TY, Peng W, Zhang MQ, Zhao K (2008) Combinatorial patterns of histone acetylations and methylations in the human genome. Nat Genet 40:897–903. https://doi.org/10.1038/ng.154
Whyte WA, Orlando DA, Hnisz D, Abraham BJ, Lin CY, Kagey MH, Rahl PB, Lee TI, Young RA (2013) Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153:307–319. https://doi.org/10.1016/j.cell.2013.03.035
Winnier GE, Kume T, Deng K et al. (1999) Roles for the winged helix transcription factors MF1 and MFH1 in cardiovascular development revealed by nonallelic noncomplementation of null alleles. Dev Biol 213:418–431. https://doi.org/10.1006/dbio.1999.9382
Yamamoto-Fukuda T, Akiyama N, Kojima H (2020) L1CAM-ILK-YAP mechanotransduction drives proliferative activity of epithelial cells in middle ear cholesteatoma. Am J Pathol 190:1667–1679. https://doi.org/10.1016/j.ajpath.2020.04.007
Yamamoto-Fukuda T, Akiyama N, Takahashi M, Kojima H (2018) Keratinocyte growth factor (KGF) modulates epidermal progenitor cell kinetics through activation of p63 in middle ear cholesteatoma. J Assoc Res Otolaryngol 19:223–241. https://doi.org/10.1007/s10162-018-0662-z
Yamamoto-Fukuda T, Akiyama N, Shibata Y, Takahashi H, Ikeda T, Koji T (2015) In vivo over-expression of KGF mimic human middle ear cholesteatoma. Eur Arch Otorhinolaryngol 272:2689–2696. https://doi.org/10.1007/s00405-014-3237-6
Yamamoto-Fukuda T, Akiyama N, Shibata Y, Takahashi H, Ikeda T, Kohno M, Koji T (2014) KGFR as a possible therapeutic target in middle ear cholesteatoma. Acta Otolaryngol 134:1121–1127. https://doi.org/10.3109/00016489.2014.907501
Yamamoto-Fukuda T, Aoki D, Hishikawa Y, Takahashi H, Kobayashi T, Koji T (2003) Possible involvement of keratinocyte growth factor and its receptor in enhanced epithelial-cell proliferation and acquired recurrence of middle-ear cholesteatoma. Lab Invest 83:123–136. https://doi.org/10.1097/01.lab.0000050763.64145.cb
Yamamoto-Fukuda T, Shibata Y, Hishikawa Y, Shin M, Yamaguchi A, Kobayashi T, Koji T (2000) Effects of various decalcification protocols on detection of DNA strand breaks by terminal dUTP nick end labelling. Histochem J 32:697–702. https://doi.org/10.1023/a:1004171517639
Yamamoto-Fukuda T, Akiyama N, Tatsumi N, Okabe M, Kojima H (2021) Menin-MLL inhibitor blocks progression of middle ear cholesteatoma in vivo. Int J Pediatr Otorhinolaryngol 140:110545. https://doi.org/10.1016/j.ijporl.2020.110545
Yang A, Kaghad M, Wang Y, Gillett E, Fleming MD, Dötsch V, Andrews NC, Caput D, McKeon F (1998) p63, a p53 homolog at 3q27-29, encodes multiple products with transactivating, death-inducing, and dominant-negative activities. Mol Cell 2:305–316. https://doi.org/10.1016/s1097-2765(00)80275-0
Zentner GE, Tesar PJ, Scacheri PC (2011) Epigenetic signatures distinguish multiple classes of enhancers with distinct cellular functions. Genome Res 21:1273–1283. https://doi.org/10.1101/gr.122382.111
Zhu JL, Song YX, Wang ZN, Gao P, Wang MX, Dong YL, Xing CZ, Xu HM (2013) The clinical significance of mesenchyme forkhead 1 (FoxC2) in gastric carcinoma. Histopathology 62:1038–1048. https://doi.org/10.1111/his.12132
Acknowledgements
We would like to thank Dr. Jeffrey S. Rubin with the National Cancer Institute/CCR/LCMB for providing the human KGF cDNA construct. We thank Dr. Masahiro Takahashi, Dr. Kazuhisa Yamamoto and Dr. Yutaka Yamamoto (Department of Otorhinolaryngology, Jikei University School of Medicine) for the harvesting of the human tissues. We also thank Miss Shiho Kondo and Mr. Daisuke Endo (Department of Histology and Cell Biology, Nagasaki University Graduate School of Biomedical Sciences) for their excellent technical assistance in this work. We also would like to thank Prof. Takehiko Koji (Department of Histology and Cell Biology, Nagasaki University Graduate School of Biomedical Sciences) for his comments regarding this work.
Funding
This study is supported by JSPS KAKENHI Grant Number JP25462647, JP16K11186, and JP19K09857 to T. Yamamoto-Fukuda, no. JP18K16908 to N. Akiyama).
Author information
Authors and Affiliations
Contributions
Yamamoto-Fukuda T, design of the work; acquisition, analysis, interpretation of data, drafting the work; Akiyama N, analysis and interpretation of data, drafting the work; Kojima H, experimental supports.
Corresponding author
Ethics declarations
Ethics Approval
All the experimental procedures were performed in accordance with the Nagasaki University with the approval of the Institutional Animal Care and Use Committee (Nos. 1209241015-2 and 1404011269), the Guidelines for Animal Experimentation of Jikei University with approval guidelines (No. 2015-139C4), the Human Ethics Review Committee of Jikei University School of Medicine (approval number is 27-344 8229).
Consent for Publication
This study protocol was approved by the Human Ethics Review Committee of Jikei University School of Medicine, and signed informed consent was obtained from all the patients or their guardians for this study (approval number is 27-344 8229).
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Yamamoto-Fukuda, T., Akiyama, N. & Kojima, H. Super-enhancer Acquisition Drives FOXC2 Expression in Middle Ear Cholesteatoma. JARO 22, 405–424 (2021). https://doi.org/10.1007/s10162-021-00801-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10162-021-00801-7