Abstract
Background
As a critical component of exosomes, circular RNAs (circRNAs) have shown great value in cancer diagnosis. This study aimed to identify circRNAs in exosomes for the diagnosis of PTC (papillary thyroid carcinoma).
Methods
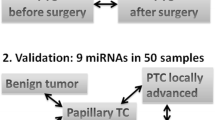
We selected hsa_circ_0082002 and hsa_circ_0003863 based on circRNA microarray. The levels of exosomal hsa_circ_0082002 and hsa_circ_0003863 in the sera of healthy control (n = 68), benign thyroid tumors (n = 60), and PTC without and with Hashimoto’s thyroiditis (n = 164) were quantified by qPCR (quantitative polymerase chain reaction). Receiver operating characteristic analyses were conducted to evaluate the diagnostic sensitivity and specificity. Bioinformatics databases were used to predict the microRNAs and proteins binding with hsa_circ_0082002 and hsa_circ_0003863.
Results
The levels of exosomal hsa_circ_0082002 and hsa_circ_0003863 were positively associated and statistically increased in PTC compared to healthy and benign thyroid tumors. Intriguingly, higher levels of exosomal hsa_circ_0082002 and hsa_circ_0003863 were positively correlated with lymph node metastasis and vascular invasion in PTC. Further stability tests show that exosomal hsa_circ_0082002 and hsa_circ_0003863 could exist stably in sera treated by several freeze–thaw cycles at -20 °C and with a storage time shorter than 24 h at 4 °C. Furthermore, hsa_circ_0082002 and hsa_circ_0003863 were predicted to interact with microRNAs and proteins, suggesting that hsa_circ_0082002 and hsa_circ_0003863 might contribute to the occurrence and progression of PTC through interacting with microRNAs and RNA binding proteins.
Conclusion
Collectively, we identified two PTC-related circRNAs incorporated in exosomes and uncovered their potential as tumor markers to diagnose PTC, in particular, more aggressive PTC.







Similar content being viewed by others
Availability of data and material
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- PTC:
-
Papillary thyroid carcinoma
- circRNA:
-
Circular RNA
- ROC:
-
Receiver operator characteristic
- AUC:
-
Area under the ROC curve
- qPCR:
-
Quantitative polymerase chain reaction
- FNAB:
-
Fine needle aspiration biopsy
- PET:
-
Positron emission tomography
- CT:
-
Computed tomography
- miRNA:
-
MicroRNA
- lncRNA:
-
Long non-coding RNA
References
Ramia de Cap M (2021) Multifocal papillary thyroid carcinoma. Am J Clin Pathol 155(6):913. https://doi.org/10.1093/ajcp/aqab005
Feldkamp J, Führer D, Luster M et al (2016) Fine needle aspiration in the investigation of thyroid nodules. Dtsch Arztebl Int 113(20):353–359. https://doi.org/10.3238/arztebl.2016.0353
Liu T, Tilak M, Awad S et al (2022) A literature review of factors associated with pain from fine needle aspiration biopsy of thyroid nodules. Endocr Pract 28(6):628–636. https://doi.org/10.1016/j.eprac.2022.03.007
Torréns JI, Burch HB (2001) Serum thyroglobulin measurement. Utility in clinical practice. Endocrinol Metab Clin North Am 30 (2):429–467. doi:https://doi.org/10.1016/s0889-8529(05)70194-8
Algeciras-Schimnich A (2018) Thyroglobulin measurement in the management of patients with differentiated thyroid cancer. Crit Rev Clin Lab Sci 55(3):205–218. https://doi.org/10.1080/10408363.2018.1450830
Cho JS, Kim HK (2022) Thyroglobulin Levels as a Predictor of Papillary Cancer Recurrence After Thyroid Lobectomy. Anticancer Res 42(11):5619–5627. https://doi.org/10.21873/anticanres.16070
Kalluri R, LeBleu VS (2020) The biology, function, and biomedical applications of exosomes. Science. https://doi.org/10.1126/science.aau6977
Mashouri L, Yousefi H, Aref AR et al (2019) Exosomes: composition, biogenesis, and mechanisms in cancer metastasis and drug resistance. Mol Cancer 18(1):75. https://doi.org/10.1186/s12943-019-0991-5
Dai J, Su Y, Zhong S et al (2020) Exosomes: key players in cancer and potential therapeutic strategy. Signal Transduct Target Ther 5(1):145. https://doi.org/10.1038/s41392-020-00261-0
Wang Y, Liu J, Ma J et al (2019) Exosomal circRNAs: biogenesis, effect and application in human diseases. Mol Cancer 18(1):116. https://doi.org/10.1186/s12943-019-1041-z
Chen L, Wang C, Sun H et al (2021) The bioinformatics toolbox for circRNA discovery and analysis. Brief Bioinform 22(2):1706–1728. https://doi.org/10.1093/bib/bbaa001
Kristensen LS, Andersen MS, Stagsted LVW et al (2019) The biogenesis, biology and characterization of circular RNAs. Nat Rev Genet 20(11):675–691. https://doi.org/10.1038/s41576-019-0158-7
Chen Y, Wang J, Wang C et al (2022) Deep learning models for disease-associated circRNA prediction: a review. Brief Bioinform. https://doi.org/10.1093/bib/bbac364
Chen L, Shan G (2021) CircRNA in cancer: fundamental mechanism and clinical potential. Cancer Lett 505:49–57. https://doi.org/10.1016/j.canlet.2021.02.004
Panda AC (2018) Circular RNAs Act as miRNA Sponges. Adv Exp Med Biol 1087:67–79. https://doi.org/10.1007/978-981-13-1426-1_6
Huang A, Zheng H, Wu Z et al (2020) Circular RNA-protein interactions: functions, mechanisms, and identification. Theranostics 10(8):3503–3517. https://doi.org/10.7150/thno.42174
Zhu G, Chang X, Kang Y, Zhao X, Tang X, Ma C, Fu S (2021) CircRNA: A novel potential strategy to treat thyroid cancer (Review). Int J Mol Med. https://doi.org/10.3892/ijmm.2021.5034
Chen Y, Ma X, Lou C et al (2022) PLA2G10 incorporated in exosomes could be diagnostic and prognostic biomarker for non-small cell lung cancer. Clin Chim Acta 530:55–65. https://doi.org/10.1016/j.cca.2022.02.016
Chen Y, Lou C, Ma X et al (2022) Serum exosomal hsa_circ_0069313 has a potential to diagnose more aggressive non-small cell lung cancer. Clin Biochem 102:56–64. https://doi.org/10.1016/j.clinbiochem.2022.01.005
Shen X, Yang Y, Chen Y et al (2022) Evaluation of EpCAM-specific exosomal lncRNAs as potential diagnostic biomarkers for lung cancer using droplet digital PCR. J Mol Med (Berl) 100(1):87–100. https://doi.org/10.1007/s00109-021-02145-4
Jiang B, Zhang J, Sun X et al (2022) Circulating exosomal hsa_circRNA_0039480 is highly expressed in gestational diabetes mellitus and may be served as a biomarker for early diagnosis of GDM. J Transl Med 20(1):5. https://doi.org/10.1186/s12967-021-03195-5
Zhi F, Ding Y, Wang R et al (2021) Exosomal hsa_circ_0006859 is a potential biomarker for postmenopausal osteoporosis and enhances adipogenic versus osteogenic differentiation in human bone marrow mesenchymal stem cells by sponging miR-431-5p. Stem Cell Res Ther 12(1):157. https://doi.org/10.1186/s13287-021-02214-y
Yu D, Li Y, Wang M et al (2022) Exosomes as a new frontier of cancer liquid biopsy. Mol Cancer 21(1):56. https://doi.org/10.1186/s12943-022-01509-9
Yang L, Chen Y, Liu N et al (2022) CircMET promotes tumor proliferation by enhancing CDKN2A mRNA decay and upregulating SMAD3. Mol Cancer 21(1):23. https://doi.org/10.1186/s12943-022-01497-w
Huang XY, Zhang PF, Wei CY et al (2020) Circular RNA circMET drives immunosuppression and anti-PD1 therapy resistance in hepatocellular carcinoma via the miR-30-5p/snail/DPP4 axis. Mol Cancer 19(1):92. https://doi.org/10.1186/s12943-020-01213-6
Yao MD, Jiang Q, Ma Y et al (2022) Targeting circular RNA-MET for anti-angiogenesis treatment via inhibiting endothelial tip cell specialization. Mol Ther 30(3):1252–1264. https://doi.org/10.1016/j.ymthe.2022.01.012
Liu J, Dai Z, Li M et al (2022) Circular RNA circMET contributes to tamoxifen resistance of breast cancer cells by targeting miR-204/AHR signaling. Biochem Biophys Res Commun 627:200–206. https://doi.org/10.1016/j.bbrc.2022.07.097
Pei X, Chen SW, Long X et al (2020) circMET promotes NSCLC cell proliferation, metastasis, and immune evasion by regulating the miR-145-5p/CXCL3 axis. Aging (Albany NY) 12(13):13038–13058. https://doi.org/10.18632/aging.103392
Liu Y, Chen L, Liu T et al (2022) Genome-wide circular RNA (circRNA) and mRNA profiling identify a circMET-miR-410-3p regulatory motif for cell growth in colorectal cancer. Genomics 114(1):351–360. https://doi.org/10.1016/j.ygeno.2021.11.038
Acknowledgements
We thank Prof. Junming Guo for providing us with the qPCR instrument.
Funding
This study is supported by the Natural Science Foundation of Ningbo (2023J322, 2023J077, and 2021J313); the Research Foundation of Ningbo No.2 Hospital (2022HMK40); the Young Technical Backbone Talent of Health in Ningbo; the Zhu Xiushan Talent Award Fund of Ningbo NO.2 Hospital (2021HMYQ06); the Project of NINGBO Leading Medical & Health Discipline (2022-F18).
Author information
Authors and Affiliations
Contributions
Study design and supervision: LD, XW, XM. Development of methodology: LD, WH, HJ. Acquisition of data (acquired and managed patients): LD, YW, QL. Analysis and interpretation of data (e.g., statistical analysis, biostatistics): LD. Writing, review, and/or revision of the manuscript: LD, XW, XM.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interest.
Ethics approval and consent to participate
This study was performed following the Declaration of Helsinki. Written informed consent was obtained from all patients, and the study was approved by the ethics committee (the Clinical Research Ethics Committee of Ningbo No.2 Hospital) with the number of. YJ-NBEY-KY-2021-181-01.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
About this article
Cite this article
Dai, L., Hu, W., Jiang, H. et al. The diagnostic potential of two exosome-derived circRNAs for papillary thyroid cancer. Int J Clin Oncol 28, 1461–1474 (2023). https://doi.org/10.1007/s10147-023-02400-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10147-023-02400-3