Abstract
Duck hepatitis A virus 1 (DHAV-1) is one of the main contagious pathogens that causes rapid death of ducklings. To illuminate the potential of DHAV-1-infected underlying mechanisms, we analyzed the mRNA and microRNA (miRNA) expression profiles of duck embryonic hepatocytes (DEHs) in response to DHAV-1. We found 3410 differentially expressed genes (DEGs) and 142 differentially expressed miRNAs (DEMs) at 36 h after DHAV-1 infection. Additionally, DEGs and the target genes of miRNA expression were analyzed and enriched utilizing GO and KEGG, which may be crucial for immune responses, viral resistance, and mitophagy. For instance, the dysregulation of DDX58, DHX58, IRF7, IFIH1, STING1, TRAF3, CALCOCO2, OPTN, PINK1, and MFN2 in DHAV-1-infected DEHs was verified by RT-qPCR. Then, the association analysis of mRNAs and miRNAs was constructed utilizing the protein–protein interaction (PPI) networks, and the expressions of main miRNAs were confirmed, including miR-132c-3p, miR-6542-3p, and novel-mir163. These findings reveal a synthetic characterization of the mRNA and miRNA in DHAV-1-infected DEHs and advance the understanding of molecular mechanism in DHAV-1 infection, which may provide a hint for the interactions of virus and host.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Duck viral hepatitis (DVH) is caused by the duck hepatitis A virus (DHAV), which has fast-spreading and highly infectious disease in ducklings (Sui et al. 2022). There are DHAV-1, DHAV-2, and DHAV-3, three types in DHAV, which is an Avihepatovirus genus (Rohaim et al. 2021). Among three types of DHAV, DHAV-1 is the most widespread and can cause neurological symptoms and acute hepatitis in ducklings (Li et al. 2022a, b, c). The other two types were only found in Asia (Feher et al. 2021). In addition, DHAV-1 infection can also cause egg drop in laying ducks (Lan et al. 2019). In detail, DHAV-1 infection can cause opisthotonos and severe liver damage, leading to rapid death of ducklings and threaten duck production. It is necessary to study the mechanism of DHAV-1-infected host target cells to better control DHAV-1 infection and reduce the economic losses of the duck industry.
RNA-sequencing (RNA-seq) technology is widely used in clinical research, biological research, and related drug development, which can comprehensively analyze the transcription and regulation of genes (Seyama et al. 2022). MicroRNAs (miRNAs) are small-noncoding RNAs involved in gene regulation and cell signaling, which are found in almost all eukaryotes (Qi et al. 2022). It is reported that miRNAs regulate most transcriptional genes by binding to the 3′ untranslated regions (Aass et al. 2022). miRNAs regulate mRNA through a variety of biological pathways such as immunity and metabolism (Movassagh et al. 2022). miRNAs play a role in a variety of biological processes, including viral inhibition of the host immune response, host response to infection, and virus–host interaction (Chatterjee et al. 2022). For instance, miRNA-4776 can regulate the production of influenza A virus in infected cell via regulating NFKBIB expression (Othumpangat et al. 2017). In addition, miRNA-223 regulated the inflammatory response induced by SARS-CoV infection by regulating host mRNAs (Morales et al. 2022). Meanwhile, miR-122 inhibits virus replication by upregulating the expression of type I interferon (Zhang et al. 2022).
Recently, combined mRNA-miRNA analysis has helped advance our understanding of miRNA function and disease mechanisms. For example, after association analysis of mRNA and miRNA in H9N2 avian influenza A virus-infected chicken, miR-126-5p regulates antiviral innate immunity by targeting TRAF3 (Wang et al. 2022). Especially, mRNA and miRNA profiling of DHAV-1-infected primary duck embryo fibroblast cells revealed miR-222a as a key factor in anti-DHAV-1 (Sui et al. 2021). However, the hepatic is the main target organ for DHAV-1-infected ducklings. The mRNA and miRNA expression profiles of DHAV-1 directly infected with DEHs were limited. In this work, we analyzed the expression of mRNA and miRNA in DEHs infected with DHAV-1 by high-throughput sequencing. Additionally, the association analysis of mRNA and miRNA can further understand the potential metabolic and signal pathway in DHAV-1-infected DEHs.
Materials and methods
Cell culture
DEHs were isolated using 14-day-old SPF duck embryos and cultured as previously reported (Qiu et al. 2022). The DEHs were cultured in DMEM (BasaIMedia, China) at 37 °C with 5% CO2, supplemented with 10% fetal calf serum (TIANHANG, China), 100 U/mL penicillin, and 100 μg/mL streptomycin.
TCID50 of DHAV-1
DEHs were cultured in 96-well plates at 37 °C with 5% CO2. DHAV-1 stock solution was diluted and replaced according to tenfold series, and incubated for 12–96 h to observe the existence of cytopathic effect (CPE). Reed Muench method is used to calculate TCID50.
DHAV-1 infection
DEHs were infected using the DHAV-1 LQ2 strain (0.01 multiplicity of infection (MOI) was given by the Shandong Institute of Poultry in China) at 37 °C for 2 h. The virus solutions were discarded, and DEHs were washed by D-Hank’s. Next, the DEHs were cultivated with 1% DMEM at 37 °C with 5% CO2. The DEHs were collected and observed at 12, 24, 36, and 48 h post-infection (hpi) after virus infection.
Sample collection and RNA quality control (QC)
The DEHs were split up into cell control (CC) group and virus control (VC) group, and each group was performed in triplicates (CC1, CC2, and CC3; VC1, VC2, and VC3). The total RNA in all groups was isolated utilizing Trizol (Vazyme, China) at 36 hpi. In addition, RNA QC was evaluated by Standard Sensitivity RNA Analysis Kit and Fragment Analyzer to specifications.
RNA sequencing
The mRNAs were enriched and used as a template to synthesize double-stranded cDNA. Subsequently, the linked products were amplified by PCR. Then, the mRNA libraries were constructed using the quality-qualified RNA. Finally, the mRNA libraries were further sequenced and inspected by DNBSEQ-T7 Genome Analyzer (MGI, China) and Agilent 2100 Bioanalyzer.
MicroRNA sequencing
The miRNA libraries were built and sequenced by DNBSEQ-T7 Genome Analyzer. Finally, the raw data were used to identify and predict miRNAs.
DEG and DEM analysis
We use DESeq2 software for DEG and DEM analysis. A smaller false discovery rate (FDR) value indicates a larger FoldChange, indicating a more significant difference in expression. Analysis of DEGs and DEMs with biological duplication is defined as | FoldChange |≥ 2 and p-adj ≤ 0.05.
Function and pathway analysis
To understand the potential function and pathways of DEGs and DEMs in DEHs, GO and KEGG were analyzed using clusterProfiler software. In addition, the screening criteria for significant results are p-adj < 0.05.
Combined mRNA-miRNA analysis
The mRNA-miRNA networks were created and visualized by Cytoscape 3.9.1, which based on mRNA and miRNA correlation.
RT-qPCR validation
The sequencing results of some screened mRNAs and miRNAs were verified by RT-qPCR. The relative PCR primers were designed based on the NCBI (Table 1). Foremost, total RNAs were extracted with Trizol and quantified with Nanodrop (Thermo, USA). Subsequently, the cDNA was formed using the HiScript® II RT supermix for qPCR Kit (Vazyme, China) and PCR Amplifier. For the miRNA, the cDNA was extracted by miRNA first-strand cDNA synthesis kit. In addition, the mRNA and miRNA analyses were achieved using qPCR mix kit, miRNA qPCR mix kit (Vazyme, China), and Light Cycler 96 instrument (Roche) and quantified using the 2−ΔΔCt method (Xie et al. 2018).
Statistical analysis
All data were expressed as means ± standard deviations (SD), and the statistical tests were performed by IBM SPSS Statistics 26.0 and GraphPad Prism 9.0. The p value < 0.05 was considered statistically significant.
Results
TCID50 of DHAV-1 in DEHs
The TCID50 of DHAV-1 was 10−4.48/0.1 mL (Table 2).
Replication of DHAV-1 in DEHs
CPE and the relative expression of DHAV-1 in DEHs were detected at different time points after infection with DHAV-1. The observation results revealed that CPEs in infected DEHs were visible at 24 hpi and became more pronounced at 36 hpi, and cell fragmentation and detachment were severe at 48 hpi (Fig. 1a). Additionally, the relative expression of DHAV-1 increased rapidly between 24 and 36 hpi (Fig. 1b). However, the infected DEHs were severely shed at 48 hpi. Consequently, in order to ensure definite infection rates and eliminate ultra-CPE, we chose to collect DEHs at 36 hpi for further mRNA and miRNA sequencing.
Characteristics of DEHs following DHAV-1 infection. a Cell morphology of DEHs following DHAV-1 infection at 12, 24, 36, and 48 hpi. b The relative quantification of DHAV-1 on DEHs at 12, 24, 36, and 48 hpi. Values within a column without the same superscripts (a–d) are significant different (p < 0.05)
RNA-seq data analysis
During DHAV-1 infection, the expression of mRNA in DEHs was applied to analyze by the Illumina high-throughput sequencing platform. More than 12.3 billion raw data were obtained of each sample, and clean reads were obtained after filtering and QC. The Q20 and Q30 were > 92.00%, and the GC was > 47.00%, showing the accuracy of the RNA-seq data (Table 3).
miRNA sequencing data analysis
The miRNA expression profile of DEHs infected with DHAV-1 was obtained by the high-throughput sequencing platform. After filtering, clean reads surpassed 10 million and comprised > 93% of the mapped reads in reference genome (Table 4). Most miRNA lengths were 23 nt, which is in line with the typical miRNA length (Fig. 2a) (Jia et al. 2022). Furthermore, the first nucleotide bias of miRNAs indicated that a large proportion of first nucleotide biases were uridine (U), which is consistent with the miRNA typical characteristic (Fig. 2b) (Xie et al. 2015).
DEG and DEM analysis
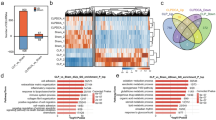
To understand the changes of DEHs infected by DHAV-1, the DEGs and DEMs in two group were compared. There were 2298 upregulated and 1112 downregulated DEGs in CC vs. VC comparison (Fig. 3a). Meanwhile, 97 upregulated and 45 downregulated DEMs were expressed (Fig. 3b). Notably, the hierarchical clustering of DEGs and DEMs revealed a strong correlation between the intra-group samples (Fig. 3c, d).
Functional and pathway enrichment analysis
The functions and pathway of DEGs and DEMs were found using GO and KEGG analyses. In short, DEGs were related primarily to transmembrane transport, ion transport, immune response, and so on (Fig. 4a, b). Simultaneously, the KEGG pathway analysis found that the DEG pathways were mostly in connection with organismal systems, environmental information processing, and metabolism, such as RIG-I like receptor (RLR) pathway, arachidonic acid metabolism, cytokine and cytokine receptor, and so on (Fig. 4c, d).
Moreover, DEM target genes were forecasted to characterize the biological functions of miRNAs. GO analysis revealed that the target genes were mainly related to molecular functions such as oxidoreductase activity, ATPase-coupled intramembrane, and voltage-gated calcium channel activity (Fig. 5a, b). Meanwhile, cGMP-PKG, cAMP, and calcium signaling pathway were the top KEGG pathways related to the target genes, indicating that the energy metabolism was significantly changes (Fig. 5c, d).
mRNA–miRNA correlation network construction
To clarify the co-expression of mRNA and miRNA, the PPI network was constructed (Fig. 6a). In addition, a miRNA can concurrently target multiple genes, and the expression of miRNAs and mRNAs was multi-directional. Later, the differential expression of mRNA-miRNA pairs in the main pathways was visual (Fig. 6b, c). For instance, the downregulated novel-mir163 regulates TRAF3, and the upregulated gga-miR-132c-3p regulates MFN2.
mRNA–miRNA correlation networks in DHAV-1-infected DEHs. a All differential expression of mRNA-miRNA pairs. b Differential miRNA-mRNA interaction networks in RIG-I-like receptor signaling pathway. c Differential miRNA-mRNA interaction networks in mitophagy. Red: upregulated miRNA. Yellow: downregulated miRNA. Blue: upregulated mRNA. Green: downregulated mRNA.
DEG and DEM validation
To test the accuracy of the RNA-seq and miRNA-seq results, the pivotal antiviral-related and mitophagy-related genes and miRNAs were examined. Briefly, the level of selected mRNAs was consistent with the RNA-seq results (Fig. 7a). Simultaneously, the tendency of selected miRNAs was in conformity with the miRNA-seq results (Fig. 7b). Moreover, the validation results of miRNA target genes were consistent with RNA-seq results (Fig. 7c).
Discussion
As a viral pathogen, DHAV-1 has emerged as the leading cause of ducklings’ death (Xie et al. 2019). DHAV-1 is the most ordinary serotype and gives rise to serious acute hepatitis in ducklings (Liu et al. 2021). The typical symptom is punctate or ecchymotic bleeding on liver surface (Zhang et al. 2020). The current research revealed that m6A modification motifs and unique motifs in attenuated DHAV-1 infected duckling livers by m6A-Seq (Wu et al. 2022). Nevertheless, the DHAV-1-infected target cell molecular mechanism is unclarified. Some researchers found that miRNA plays a necessary role in the mutual effect between virus and host (Chirayil et al. 2018). Herein, we analyzed the mRNA and miRNA in DHAV-1-infected DEHs to illuminate duckling-DHAV-1 interaction mechanisms.
DHAV-1 infection causes the dysregulation of mRNAs expressions on DEHs. A total of 3410 DEGs were found in DHAV-1-infected DEHs. Subsequently, DEGs were forecasted for further functional enrichment analysis. GO analysis specified that the DEGs were centralized in multiple biotic processes, including transmembrane transport, ion transport, and immune response. Moreover, KEGG analysis revealed that DEGs involved in PI3K-Akt pathway, Ras pathway, RLR pathway, and mitophagy. The RLRs are necessary for discovering viral RNA and starting the innate immune response (Kell and Gale 2015). Briefly, in RNA viruses’ infections, the viral RNA binds to RLRs after entering the host cells and stimulates the activation of RIG-I or MDA5 (Zhang et al. 2021). The sensitized RIG-I or MDA5 recruits the MAVS at the mitochondrial membrane to promote the constitution of MAVS filaments (Li et al. 2021). Whereafter, MAVS filaments recruit TBK1, leading to the assembly of a complex crucial to the excitation of IRF3 and IRF7, thereby activating type I interferon responses and forming an antiviral state (Wang et al. 2021a, b, c). A previous study implicates RLR pathway as a main antiviral pathway against Kaposi’s sarcoma-associated herpesvirus infection (Zhao et al. 2018). In our study, several genes of RLR pathway, including DDX58 (RIG-I), DHX58 (LGP2), IFIH1 (MDA5), STING1, TRAF3, and IRF7, were significantly increased whether RNA-seq or RT-qPCR. The recent study demonstrated that long noncoding RNAs play essential roles in the RLR pathway by regulating related genes, defending DHAV-1 infection (Sui et al. 2022). For instance, DDX58 is a crucial factor in the cytoplasmic pattern recognition receptor family that initiates innate immune responses by detecting viral RNA (Frietze et al. 2022). Generally, IFIH1 (MDA5) can detect RNA from particular viral species, such as Picornaviridae (Stok et al. 2022). Hence, we speculate that RLR pathway is the key pathway for DEHs against DHAV-1 infection.
DHAV-1 infected motivates the dysregulation of miRNAs. Specifically, in DHAV-1-infected DEHs, a majority of the miRNA lengths ranged from 21 to 24 nt and most first nucleotide bases were U, indicating that the DEH miRNA characteristics were consistent with previous research (McGeary et al. 2022). In CC vs. VC comparison, we found a total of 142 DEMs. Then, the DEM target genes were predicted and performed functional annotation. In a word, DEM target genes have enriched energy metabolism. Moreover, mitochondria are the main organelles responsible for energy metabolism and ATP production in cells (Yan et al. 2021). Notably, not only DEGs but also DEM target genes has enriched mitophagy. In this study, the upregulated CALCOCO2 (NDP52) and OPTN and the downregulated PINK1 and MFN2 in mitophagy were found from RNA/miRNA-seq and PT-qPCR. Previous studies have reported that mitophagy is induced during virus infection, thereby destroying the function of mitochondria in managing inflammation and immune response (Li et al. 2022a, b, c; Wang et al. 2021a, b, c). For example, some investigators elaborated that syndrome coronavirus 2 ORF10 inhibits antiviral innate immune response by reducing MAVS through mitophagy (Li et al. 2022a, b, c). Additionally, another study indicated that influenza A virus induced mitophagy via interactions with TUFM and LC3B, thereby suppressing the antiviral innate immune response (Wang et al. 2021a, b, c). Therefore, the DHAV-1 infection may affect the antiviral pathway (RLR pathway) through mitochondria and mitophagy.
Next, the relevance between mRNA and miRNA was used for constituting the networks and implementing a comprehensively analysis. The correlation analysis identified 115 mRNA-miRNA pairs in CC vs. VC comparisons. Universally, majority of miRNAs could target more than one gene. Furthermore, the antiviral-related and mitophagy-related mRNA-miRNAs were established. The gga-miR-132c-3p (targeted by MFN2), gga-miR-6542-3p (targeted by PINK1), and novel-mir163 (targeted by TRAF3) were selected to verify, and the trend was consistent with miRNA-seq. However, these miRNAs have not been reported in antiviral studies. During RNA virus infection, MFN2 was responsible for IL-1β generation and interacted with MAVS to inhibit antiviral immunity (Silwal et al. 2021). It is reported that classical swine fever virus replication is accomplished by regulating PINK1 to induce mitophagy (Chengcheng et al. 2022). In addition, several studies elucidated that antiviral immunity is activated by catalyzing TRAF3 (Gao et al. 2021; Sun et al. 2020). Thus, these miRNAs may be prospective candidates for antiviral immunity.
In summary, this work is the first to identify mRNA and miRNA expression patterns in DHAV-1-infected DEHs for a further exploration of DHAV-1-target cell interactions. Moreover, we screen several dysregulated genes and miRNAs during DHAV-1 infection, involved in antiviral immunity and mitophagy. These findings provide serviceable mRNA and miRNA sequencing to better understand the changes of target cells after DHAV-1 infection. Further research will focus on the mechanisms of antiviral immunity and mitophagy during DHAV-1 infection, and find the potential therapeutic targets against DHAV-1.
Abbreviations
- RNA-seq:
-
RNA-sequencing
- miRNA-seq:
-
MicroRNA-sequencing
- miRNAs:
-
MicroRNAs
- DHAV-1:
-
Duck hepatitis A virus 1
- DEHs:
-
Duck embryonic hepatocytes
- DEGs:
-
Differentially expressed genes
- DEMs:
-
Differentially expressed miRNAs
- GO:
-
Gene Ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- PPI:
-
Protein-protein interaction
- DVH:
-
Duck viral hepatitis
- DMEM:
-
Dulbecco’s modified Eagle’s medium
- TCID50 :
-
50% Tissue culture infective doses
- RT-qPCR:
-
Real-time qualitative PCR
- RIG-I:
-
Retinoic acid-inducible gene I
- LGP2:
-
Laboratory of Genetics and Physiology 2
- MDA5:
-
Melanoma differentiation-associated gene 5
- TUFM:
-
Tu translation elongation factor, mitochondrial
- LC3B:
-
Microtubule-associated protein 1 light chain 3 beta
- DDX58:
-
DExD/H-box helicase 58
- DHX58:
-
DExH-box helicase 58
- IRF7:
-
Interferon regulatory factor 7
- IFIH1:
-
Interferon induced with helicase C domain 1
- STING1:
-
Stimulator of interferon response cGAMP interactor 1
- TRAF3:
-
TNF receptor associated factor 3
- CALCOCO2:
-
Calcium binding and coiled-coil domain 2
- OPTN:
-
Optineurin
- PINK1:
-
PTEN-induced kinase 1
- MFN2:
-
Mitofusin 2
References
Aass KR, Nedal TMV, Tryggestad SS, Haukas E, Slordahl TS, Waage A, Standal T, Mjelle R (2022) Paired miRNA- and messenger RNA-sequencing identifies novel miRNA-mRNA interactions in multiple myeloma. Sci Rep 12:12147. https://doi.org/10.1038/s41598-022-16448-0
Chatterjee S, Mukherjee I, Bhattacharjee S, Bose M, Chakrabarti S, Bhattacharyya SN (2022) Target-dependent coordinated biogenesis of secondary microRNAs by miR-146a balances macrophage activation processes. Mol Cell Biol 42:e0045221. https://doi.org/10.1128/mcb.00452-21
Chengcheng Z, Xiuling W, Jiahao S, Mengjiao G, Xiaorong Z, Yantao W (2022) Mitophagy induced by classical swine fever virus nonstructural protein 5A promotes viral replication. Virus Res 320:198886. https://doi.org/10.1016/j.virusres.2022.198886
Chirayil R, Kincaid RP, Dahlke C, Kuny CV, Dalken N, Spohn M, Lawson B, Grundhoff A, Sullivan CS (2018) Identification of virus encoded microRNAs in divergent Papillomaviruses. PLoS Pathog 14:e1007156. https://doi.org/10.1371/journal.ppat.1007156
Feher E, Jakab S, Bali K, Kaszab E, Nagy B, Ihasz K, Balint A, Palya V, Banyai K (2021) Genomic epidemiology and evolution of duck hepatitis A virus. Viruses 13. https://doi.org/10.3390/v13081592
Frietze KK, Brown AM, Das D, Franks RG, Cunningham JL, Hayward M, Nickels JT Jr (2022) Lipotoxicity reduces DDX58/Rig-1 expression and activity leading to impaired autophagy and cell death. Autophagy 18:142–160. https://doi.org/10.1080/15548627.2021.1920818
Gao P, Ma X, Yuan M, Yi Y, Liu G, Wen M, Jiang W, Ji R, Zhu L, Tang Z, Yu Q, Xu J, Yang R, Xia S, Yang M, Pan J, Yuan H, An H (2021) E3 ligase Nedd4l promotes antiviral innate immunity by catalyzing K29-linked cysteine ubiquitination of TRAF3. Nat Commun 12:1194. https://doi.org/10.1038/s41467-021-21456-1
Jia B, Wang X, Ma F, Li X, Han X, Zhang L, Li J, Diao N, Shi K, Ge C, Yang F, Du R (2022) The combination of SMRT sequencing and Illumina sequencing highlights organ-specific and age-specific expression patterns of miRNAs in Sika Deer. Front Vet Sci 9:1042445. https://doi.org/10.3389/fvets.2022.1042445
Kell AM, Gale M Jr (2015) RIG-I in RNA virus recognition. Virology 479–480:110–121. https://doi.org/10.1016/j.virol.2015.02.017
Lan J, Zhang R, Yu H, Wang J, Xue W, Chen J, Lin S, Wang Y, Xie Z, Jiang S (2019) Quantitative proteomic analysis uncovers the mediation of endoplasmic reticulum stress-induced autophagy in DHAV-1-infected DEF cells. Int J Mol Sci 20. https://doi.org/10.3390/ijms20246160
Li J, Wang M, Zhou S, Cheng A, Ou X, Sun D, Wu Y, Yang Q, Gao Q, Huang J, Tian B, Mao S, Zhang S, Zhao X, Jia R, Liu M, Zhu D, Chen S, Liu Y, Yu Y, Zhang L, Pan L (2022a) The DHAV-1 protein VP1 interacts with PI3KC3 to induce autophagy through the PI3KC3 complex. Vet Res 53:64. https://doi.org/10.1186/s13567-022-01081-6
Li X, Hou P, Ma W, Wang X, Wang H, Yu Z, Chang H, Wang T, Jin S, Wang X, Wang W, Zhao Y, Zhao Y, Xu C, Ma X, Gao Y, He H (2022b) SARS-CoV-2 ORF10 suppresses the antiviral innate immune response by degrading MAVS through mitophagy. Cell Mol Immunol 19:67–78. https://doi.org/10.1038/s41423-021-00807-4
Li S, Kuang M, Chen L, Li Y, Liu S, Du H, Cao L, You F (2021) The mitochondrial protein ERAL1 suppresses RNA virus infection by facilitating RIG-I-like receptor signaling. Cell Rep 34:108631. https://doi.org/10.1016/j.celrep.2020.108631
Li Y, Wu K, Zeng S, Zou L, Li X, Xu C, Li B, Liu X, Li Z, Zhu W, Fan S, Chen J (2022c) The role of mitophagy in viral infection. Cells 11. https://doi.org/10.3390/cells11040711
Liu Y, Cheng A, Wang M, Mao S, Ou X, Yang Q, Wu Y, Gao Q, Liu M, Zhang S, Huang J, Jia R, Zhu D, Chen S, Zhao X, Yu Y, Liu Y, Zhang L, Tian B, Pan L (2021) Duck hepatitis A virus type 1 induces eIF2alpha phosphorylation-dependent cellular translation shutoff via PERK/GCN2. Front Microbiol 12:624540. https://doi.org/10.3389/fmicb.2021.624540
McGeary SE, Bisaria N, Pham TM, Wang PY, Bartel DP (2022) MicroRNA 3′-compensatory pairing occurs through two binding modes, with affinity shaped by nucleotide identity and position. Elife 11. https://doi.org/10.7554/eLife.69803
Morales L, Oliveros JC, Enjuanes L, Sola I 2022 Contribution of host miRNA-223-3p to SARS-CoV-induced lung inflammatory pathology. mBio 13:e0313521. https://doi.org/10.1128/mbio.03135-21
Movassagh M, Morton SU, Hehnly C, Smith J, Doan TT, Irizarry R, Broach JR, Schiff SJ, Bailey JA, Paulson JN (2022) mirTarRnaSeq: an R/bioconductor statistical package for miRNA-mRNA target identification and interaction analysis. BMC Genomics 23:439. https://doi.org/10.1186/s12864-022-08558-w
Othumpangat S, Bryan NB, Beezhold DH, Noti JD (2017) Upregulation of miRNA-4776 in influenza virus infected bronchial epithelial cells is associated with downregulation of NFKBIB and increased viral survival. Viruses 9. https://doi.org/10.3390/v9050094
Qi J, Han W, Zhong N, Gou Q, Sun C (2022) Integrated analysis of miRNA-mRNA regulatory network and functional verification of miR-338-3p in coronary heart disease. Funct Integr Genomics 23:16. https://doi.org/10.1007/s10142-022-00941-w
Qiu T, Shi Y, Wang R, Wang J, Wang W, Zhu J, Wang W, Wu Y, Li K, Liu J (2022) Treatment effects of phosphorylated Chrysanthemum indicum polysaccharides on duck viral hepatitis by protecting mitochondrial function from oxidative damage. Vet Microbiol 275:109600. https://doi.org/10.1016/j.vetmic.2022.109600
Rohaim MA, Naggar RFE, AbdelSabour MA, Ahmed BA, Hamoud MM, Ahmed KA, Zahran OK, Munir M (2021) Insights into the genetic evolution of duck hepatitis A virus in Egypt. Animals (Basel) 11. https://doi.org/10.3390/ani11092741
Seyama R, Uchiyama Y, Ceroni JRM, Kim VEH, Furquim I, Honjo RS, Castro MAA, Pires LVL, Aoi H, Iwama K, Hamanaka K, Fujita A, Tsuchida N, Koshimizu E, Misawa K, Miyatake S, Mizuguchi T, Makino S, Itakura A, Bertola DR, Kim CA, Matsumoto N (2022) Pathogenic variants detected by RNA sequencing in Cornelia de Lange syndrome. Genomics 114:110468. https://doi.org/10.1016/j.ygeno.2022.110468
Silwal P, Kim JK, Jeon SM, Lee JY, Kim YJ, Kim YS, Seo Y, Kim J, Kim SY, Lee MJ, Heo JY, Jung MJ, Kim HS, Hyun DW, Han JE, Whang J, Huh YH, Lee SH, Heo WD, Kim JM, Bae JW, Jo EK (2021) Mitofusin-2 boosts innate immunity through the maintenance of aerobic glycolysis and activation of xenophagy in mice. Commun Biol 4:548. https://doi.org/10.1038/s42003-021-02073-6
Stok JE, Oosenbrug T, Ter Haar LR, Gravekamp D, Bromley CP, Zelenay S, Reis ESC, van der Veen AG (2022) RNA sensing via the RIG-I-like receptor LGP2 is essential for the induction of a type I IFN response in ADAR1 deficiency. EMBO J 41:e109760. https://doi.org/10.15252/embj.2021109760
Sui N, Zhang R, Jiang Y, Yu H, Xu G, Wang J, Zhu Y, Xie Z, Hu J, Jiang S (2021) Integrated miRNA and mRNA expression profiles reveal differentially expressed miR-222a as an antiviral factor against duck hepatitis A virus type 1 infection. Front Cell Infect Microbiol 11:811556. https://doi.org/10.3389/fcimb.2021.811556
Sui N, Zhang R, Jiang Y, Yu H, Xu G, Wang J, Zhu Y, Xie Z, Hu J, Jiang S (2022) Long noncoding RNA expression rofiles elucidate the potential roles of lncRNA- XR_003496198 in duck hepatitis A virus type 1 infection. Front Cell Infect Microbiol 12:858537. https://doi.org/10.3389/fcimb.2022.858537
Sun N, Jiang L, Ye M, Wang Y, Wang G, Wan X, Zhao Y, Wen X, Liang L, Ma S, Liu L, Bu Z, Chen H, Li C (2020) TRIM35 mediates protection against influenza infection by activating TRAF3 and degrading viral PB2. Protein Cell 11:894–914. https://doi.org/10.1007/s13238-020-00734-6
Wang R, Zhu Y, Ren C, Yang S, Tian S, Chen H, Jin M, Zhou H (2021c) Influenza A virus protein PB1-F2 impairs innate immunity by inducing mitophagy. Autophagy 17:496–511. https://doi.org/10.1080/15548627.2020.1725375
Wang J, Cheng Y, Wang L, Sun A, Lin Z, Zhu W, Wang Z, Ma J, Wang H, Yan Y, Sun J (2022) Chicken miR-126-5p negatively regulates antiviral innate immunity by targeting TRAF3. Vet Res 53:82. https://doi.org/10.1186/s13567-022-01098-x
Wang C, Ling T, Zhong N, Xu LG (2021a) N4BP3 regulates RIG-I-like receptor antiviral signaling positively by targeting mitochondrial antiviral signaling protein. Front Microbiol 12:770600. https://doi.org/10.3389/fmicb.2021.770600
Wang H, Zheng Y, Huang J, Li J (2021) Mitophagy in antiviral immunity Front Cell. Dev Biol 9:723108. https://doi.org/10.3389/fcell.2021.723108
Wu L, Quan W, Zhang Y, Wang M, Ou X, Mao S, Sun D, Yang Q, Wu Y, Wei Y, Jia R, Chen S, Zhu D, Liu M, Zhao X, Zhang S, Huang J, Gao Q, Tian B, Cheng A (2022) Attenuated duck hepatitis A virus infection is associated with high mRNA maintenance in duckling liver via m6A modification. Front Immunol 13:839677. https://doi.org/10.3389/fimmu.2022.839677
Xie F, Wang Q, Zhang B (2015) Global microRNA modification in cotton (Gossypium hirsutum L.). Plant Biotechnol J 13:492–500. https://doi.org/10.1111/pbi.12271
Xie J, Zeng Q, Wang M, Ou X, Ma Y, Cheng A, Zhao XX, Liu M, Zhu D, Chen S, Jia R, Yang Q, Wu Y, Zhang S, Liu Y, Yu Y, Zhang L, Chen X (2018) Transcriptomic characterization of a chicken embryo model infected with duck hepatitis A virus type 1. Front Immunol 9:1845. https://doi.org/10.3389/fimmu.2018.01845
Xie J, Wang M, Cheng A, Zhao XX, Liu M, Zhu D, Chen S, Jia R, Yang Q, Wu Y, Zhang S, Liu Y, Yu Y, Zhang L, Chen X (2019) DHAV-1 inhibits type I interferon signaling to assist viral adaption by increasing the expression of SOCS3. Front Immunol 10:731. https://doi.org/10.3389/fimmu.2019.00731
Yan W, Diao S, Fan Z (2021) The role and mechanism of mitochondrial functions and energy metabolism in the function regulation of the mesenchymal stem cells. Stem Cell Res Ther 12:140. https://doi.org/10.1186/s13287-021-02194-z
Zhang R, Yang Y, Lan J, Lin S, Xie Z, Zhang X, Jiang S (2020) A novel peptide isolated from a phage display peptide library modeling antigenic epitope of DHAV-1 and DHAV-3. Vaccines (Basel) 8. https://doi.org/10.3390/vaccines8010121
Zhang X, Yang F, Li K, Cao W, Ru Y, Chen S, Li S, Liu X, Zhu Z, Zheng H (2021) The insufficient activation of RIG-I-like signaling pathway contributes to highly efficient replication of porcine picornaviruses in IBRS-2 cells. Mol Cell Proteomics 20:100147. https://doi.org/10.1016/j.mcpro.2021.100147
Zhang J, Li F, Sun P, Wang J, Li K, Zhao Z, Bai X, Cao Y, Bao H, Li D, Zhang J, Liu Z, Lu Z (2022) Downregulation of miR-122 by porcine reproductive and respiratory syndrome virus promotes viral replication by targeting SOCS3. Vet Microbiol 275:109595. https://doi.org/10.1016/j.vetmic.2022.109595
Zhao Y, Ye X, Dunker W, Song Y, Karijolich J (2018) RIG-I like receptor sensing of host RNAs facilitates the cell-intrinsic immune response to KSHV infection. Nat Commun 9:4841. https://doi.org/10.1038/s41467-018-07314-7
Funding
The project was supported by the Open Project Program of Beijing Key Laboratory of Traditional Chinese Veterinary Medicine at Beijing University of Agriculture (No. BUAPSP202204) and National Natural Science Foundation of China (Grant No. 31772784), the project funded by the Priority Academic Program Development (PAPD) of Jiangsu Higher Education Institutions.
Author information
Authors and Affiliations
Contributions
Weiran Wang: conceptualization, methodology, software, and writing—original draft. Kun Li: data curation and methodology. Tao Zhang: data curation. Hong Dong: conceptualization and resources. Jiaguo Liu: conceptualization, funding acquisition, resources, and supervision.
Corresponding authors
Ethics declarations
Ethical approval
All animal experiments were approved by the Animal Care and Use Committee of Nanjing Agricultural University.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, W., Li, K., Zhang, T. et al. RNA-seq and microRNA association analysis to explore the pathogenic mechanism of DHAV-1 infection with DEHs. Funct Integr Genomics 23, 99 (2023). https://doi.org/10.1007/s10142-023-01022-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10142-023-01022-2