Abstract
Objectives
To investigate the clinical characteristics and molecular epidemiology of CRKP infection in neonatal patients in a children’s hospital in China from 2017 to 2021.
Methods
Species identification and antibiotic susceptibilities were tested with matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) and VITEK 2 systems. The clinical data were collected from medical records. Carbapenem-resistant Klebsiella pneumoniae (CRKP) isolates were investigated by antimicrobial susceptibility testing, carbapenemase genes and multilocus sequence typing.
Results
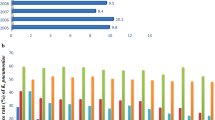
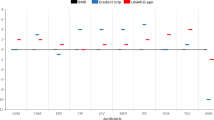
Six kinds of resistant genes and 23 STs were detected. BlaNDM-1 (n=83, 55.3%) was the predominant carbapenemase gene, followed by blaKPC-2 (n=45, 30.0%), blaNDM-5 (n=7, 4.7%), blaIMP-38 (n=6, 4.0%). BlaNDM-1 was predominant in 2017 and 2018, whereas blaKPC-2 increased in 2019 and became the predominant gene from 2020 to 2021. ST11 accounted for most infections (n=35, 23.3%), followed by ST278 (n=23, 15.3%), ST17 (n=17, 11. 3%) and ST2735 (n=16, 10.7%). ST278 and ST17 were predominant in 2017 and 2018, whereas ST11 increased in 2019 and became the predominant sequence type from 2020 to 2021. Compared with blaNDM-1, the CRKP strains producing blaKPC-2 were characterized by high resistance to gentamicin, amikacin and levofloxacin and the change trend of drug resistance rate before and after COVID-19 was consistent with that of blaNDM-1 and blaKPC-2.
Conclusions
The main sequence type of CRKP infection changed dynamically from ST278-NDM-1 to ST11-KPC-2 during the years 2017–2021 in the newborns. Antibiotic exposure and the prevalence of COVID-19 since 2020 may have led to changes in hospital population and lead to the changes.




Similar content being viewed by others
Data availability
The data presented in this study are available on reasonable request from the corresponding author.
References
Ahmad N et al (2018) Occurrence of blaNDM variants among Enterobacteriaceae from a neonatal intensive care unit in a Northern India Hospital. Front microbiol 9:407. https://doi.org/10.3389/fmicb.2018.00407
Aguilera-Alonso D et al (2020) Carbapenem-resistant gram-negative bacterial infections in children. Antimicrob agent chemot 64:e02183–e02119. https://doi.org/10.1128/AAC.02183-19
Boni S et al (2022) Association between consumption of fluoroquinolones and carbapenems and their resistance rates in Pseudomonas aeruginosa in Argentina. Interdiscip Perspect Infect Dis 2(2022):3924212. https://doi.org/10.1155/2022/3924212
Castagnola E et al (2019) Epidemiology of carbapenemase-producing Enterobacteriaceae in a pediatric hospital in a country with high endemicity. J Infect Public Health 12:270–274. https://doi.org/10.1016/j.jiph.2018.11.003
Clinical and Laboratory Standards Institute (2020) Performance standards for antimicrobial susceptibility testing. M100-S30
Deshpande LM et al (2006) Emergence of serine carbapenemases (KPC and SME) among clinical strains of Enterobacteriaceae isolated in the United States Medical Centers: report from the MYSTIC Program (1999-2005). Diagnostic microbiol infect disease 56:367–372. https://doi.org/10.1016/j.diagmicrobio.2006.07.004
Diestra K, Miró E, Martí C, Navarro D, Cuquet J, Coll P, Navarro F (2011) Multiclonal epidemic of Klebsiella pneumoniae isolates producing DHA-1 in a Spanish hospital. Clin microb infect: the official pub Europ Soc Clin Microbiol Infect Diseas 17(7):1032–1036. https://doi.org/10.1111/j.1469-0691.2010.03319.x
Doi Y (2019) (2019) Treatment options for carbapenem-resistant gram-negative bacterial infections. Clin Infect Dis 69(Suppl 7):S565–S575. https://doi.org/10.1093/cid/ciz830
Ernst CM et al (2020) Adaptive evolution of virulence and persistence in carbapenem-resistant Klebsiella pneumoniae. Nat med 26:705–711. https://doi.org/10.1038/s41591-020-0825-4
Fu P et al (2021) Bacterial epidemiology and antimicrobial resistance profiles in children reported by the ISPED program in China, 2016 to 2020. Microbiol Spectr 9:e0028321. https://doi.org/10.1128/Spectrum.00283-21
Gaillot O, Clément C, Simonet M, Philippon A (1997) Novel transferable beta-lactam resistance with cephalosporinase characteristics in Salmonella enteritidis. J antimicrob chemoth 39(1):85–87. https://doi.org/10.1093/jac/39.1.85
Gu D et al (2018) A fatal outbreak of ST11 carbapenem-resistant hypervirulent Klebsiella pneumoniae in a Chinese hospital: a molecular epidemiological study. Lancet Infect diseas 18:37–46
Horan TC, Andrus M, Dudeck MA (2008) CDC/NHSN surveillance definition of health care-associated infection and criteria for specific types of infections in the acute care setting. Am j infect control 36(5):309–332. https://doi.org/10.1016/j.ajic.2008.03.002
Hu F et al (2018) Current status and trends of antibacterial resistance in China. Clin Infect Dis 67(Suppl 2):S128–S134. https://doi.org/10.1093/cid/ciy657
Hu F et al (2022) (2021) CHINET surveillance of bacterial resistance in China: 21 report. Chin J Infect Chemother 22:521–530
Kis Z et al (2016) Countrywide dissemination of a DHA-1-type plasmid-mediated AmpC β-lactamase-producing Klebsiella pneumoniae ST11 international high-risk clone in Hungary, 2009-2013. J med microbiol 65:1020–1027. https://doi.org/10.1099/jmm.0.000302
Logan LK, Weinstein RA (2017) The epidemiology of carbapenem-resistant Enterobacteriaceae: the impact and evolution of a global menace. J infect diseas 215(suppl_1):S28–S36. https://doi.org/10.1093/infdis/jiw282
Lutgring JD (2019) Carbapenem-resistant Enterobacteriaceae: an emerging bacterial threat. Semina diagnos pathol 36(3):182–186. https://doi.org/10.1053/j.semdp.2019.04.011
Liao W et al (2020) (2020) Virulence evolution, molecular mechanisms of resistance and prevalence of ST11 carbapenem-resistant Klebsiella pneumoniae in China: a review over the last 10 years. J Glob Antimicrob Resist 23:174–180. https://doi.org/10.1016/j.jgar.2020.09.004
Mei YF et al (2017) Virulence and genomic feature of a virulent Klebsiella pneumoniae sequence type 14 strain of serotype K2 harboring blaNDM-5 in China. Front microbiol 8:335. https://doi.org/10.3389/fmicb.2017.00335
Mukherjee S et al (2021) Neonatal sepsis: the impact of carbapenem-resistant and hypervirulent Klebsiella pneumoniae. Front med 8:634349. https://doi.org/10.3389/fmed.2021.634349
Nordmann P et al (2009) The real threat of Klebsiella pneumoniae carbapenemase-producing bacteria. Lancet Infect diseases 9:228–236
Qin S et al (2014) High incidence and endemic spread of NDM-1-positive Enterobacteriaceae in Henan Province, China. Antimicrob agents chemoth 58:4275–4282. https://doi.org/10.1128/AAC.02813-13
Rodríguez-Baño J et al (2018) Treatment of infections caused by extended-spectrum-betalactamase-, AmpC-, and carbapenemase-producing Enterobacteriaceae. Clin Microbiol Rev 31:e00079–e00017. https://doi.org/10.1128/CMR.00079-17
Tang Y et al (2020) Absence of the type I-E CRISPR-Cas system in Klebsiella pneumoniae clonal complex 258 is associated with dissemination of IncF epidemic resistance plasmids in this clonal complex. J antimicrob chemoth 75:890–895. https://doi.org/10.1093/jac/dkz53827
Vanwynsberghe T et al (2007) Outbreak of Klebsiella pneumoniae strain harbouring an AmpC (DHA-1) and a blaSHV-11 in a Belgian hospital, August-December 2006. Euro surveillance: bulletin Europeen sur les maladies transmissibles =. Europ commun disease bullet 12:E070201.3. https://doi.org/10.2807/esw.12.05.03130-en
Veeraraghavan B et al (2019) Antimicrobial susceptibility profile & resistance mechanisms of Global Antimicrobial Resistance Surveillance System (GLASS) priority pathogens from India. Indian J Med Res 149:87–96
Wyres K et al (2021, 2022) Regional differences in carbapenem-resistant Klebsiella pneumoniae. Lancet Infect Dis:309–310
Yin D et al (2018) Clinical and molecular epidemiologic characteristics of carbapenem-resistant Klebsiella pneumoniae infection/colonization among neonates in China. J hospital infect 100:21–28. https://doi.org/10.1016/j.jhin.2018.05.005
Yin L et al (2020) Actively surveillance and appropriate patients placements' contact isolation dramatically decreased Carbapenem-Resistant Enterobacteriaceae infection and colonization in pediatric patients in China. J hospital infect S0195-6701:30130–30134. Advance online publication. https://doi.org/10.1016/j.jhin.2020.03.031
Yin L et al (2021) Carbapenem-resistant Enterobacterales colonization and subsequent infection in a neonatal intensive care unit in Shanghai China. Infect prevent pract 3:100147. https://doi.org/10.1016/j.infpip.2021.100147
Zhang D et al (2019) Antibiotic consumption versus the prevalence of carbapenem-resistant Gram-negative bacteria at a tertiary hospital in China from 2011 to 2017. J infect public health 12:195–199. https://doi.org/10.1016/j.jiph.2018.10.003
Zhou F et al (2020 Mar 28) (2020) Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: a retrospective cohort study. Lancet 395:1054–1062. https://doi.org/10.1016/S0140-6736(20)30566-3
Zhang WM et al (2022) Correlation between the consumption of carbapenems and the resistance of gram-negative bacteria [J]. World Clini Drug 43:1405–1411. https://doi.org/10.13683/j.wph.2022.11.004
Acknowledgements
We thank the staff of the Institute Pasteur MLST and whole genome MLST databases for curating the data and making them publicly available at http://bigsdb.pasteur.fr.
Funding
This work was supported by the National Key Research and Development Program of China (grant numbers 2021YFC2701800,2021YFC2701805)
Author information
Authors and Affiliations
Contributions
Lijun Yin performed data analysis and wrote the paper. Leiyan He carried out the bacteria identification. Lijun Yin and Lu lu prepared the Tables and figures. Laishuan Wang, Guoping Lu, Yun Cao, Xiaowen Zhai and Chuanqing Wang contributed to experiment conception and design.All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Ethics approval
The patient informed consent was waived because only bacterial isolates recovered from routine diagnostic laboratory tests were assessed. The study was approved by the Ethics Committee of the Children's Hospital of Fudan University, Shanghai, China (approval number (2021)372).
Consent for publication
All authors have read and agreed to the published version of the manuscript.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yin, L., Lu, L., He, L. et al. Shift in the dominant sequence type of carbapenem-resistant Klebsiella pneumonia infection from ST278-NDM-1 to ST11-KPC-2 in neonatal patients in a children’s hospital in Shanghai, China, 2017–2021. Int Microbiol (2023). https://doi.org/10.1007/s10123-023-00436-z
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10123-023-00436-z