Abstract
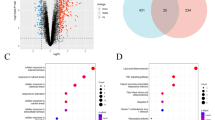
The majority of the biomarkers were associated with the diagnosis of epilepsy and few of them can be applied to predict the response to antiseizure medications (ASMs). In this study, we identified 26 significantly up-regulated genes and 32 down-regulated genes by comparing the gene expression profiles of patients with epilepsy that responded to valproate with those without applying any ASM. The results of gene set enrichment analysis indicated that the ferroptosis pathway was significantly impacted (p = 0.0087) in patients who responded to valproate. Interestingly, the gene NCOA4 in this pathway exhibited significantly different expression levels between the two groups, indicating that NCOA4 could serve as a potential biomarker to better understand the mechanism of valproate resistance. In addition, six up-regulated genes SF3A2, HMGN2, PABPN1, SSBP3, EFTUD2, and CREB3L2 as well as six down-regulated genes ZFP36L1, ACRC, SUB1, CALM2, TLK1, and STX2 also showed significantly different expression patterns between the two groups. Moreover, based on the gene expression profiles of the patients with the treatment of valproate, carbamazepine, and phenytoin, we proposed a strategy for predicting the response to the ASMs by using the Connectivity Map scoring method. Our findings could be helpful for better understanding the mechanisms of drug resistance of ASMs and improving the clinical treatment of epilepsy.





Similar content being viewed by others
Data availability
This datasets analyzed during the current study are available in the Gene Expression Omnibus, accession number: GSE143272 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE143272).
Abbreviations
- ASMs:
-
Antiseizure medications
- VPA:
-
Valproate
- CBZ:
-
Carbamazepine
- PHT:
-
Phenytoin
- CMap:
-
Connectivity Map
- DEGs:
-
Differentially expressed genes
- GEO:
-
Gene Expression Omnibus
- DAVID:
-
Visualization and Integrated Discovery
- SNM:
-
Supervised Normalization of Microarrays
- WTCS:
-
Weighted connectivity score
- AUC:
-
Area under the receiver operating characteristic curve
- FTH1:
-
Ferritin heavy chain 1
- NCOA4:
-
Nuclear receptor coactivator 4
- PCBP1:
-
Poly r(C) binding protein 1
- OPMD:
-
Oculopharyngeal muscular dystrophy
- IQR:
-
Interquartile range
References
Fisher RS et al (2014) ILAE official report: a practical clinical definition of epilepsy. Epilepsia 55(4):475–482
Rizvi S et al (2017) Epidemiology of early stages of epilepsy: risk of seizure recurrence after a first seizure. Seizure 49:46–53
Chen Z et al (2018) Treatment outcomes in patients with newly diagnosed epilepsy treated with established and new antiepileptic drugs: a 30-year longitudinal cohort study. JAMA Neurol 75(3):279–286
Kalilani L et al (2018) The epidemiology of drug-resistant epilepsy: a systematic review and meta-analysis. Epilepsia 59(12):2179–2193
Fisher RS et al (2017) Instruction manual for the ILAE 2017 operational classification of seizure types. Epilepsia 58(4):531–542
Scheffer IE et al (2017) ILAE classification of the epilepsies: position paper of the ILAE Commission for Classification and Terminology. Epilepsia 58(4):512–521
Specchio LM, Beghi E (2004) Should antiepileptic drugs be withdrawn in seizure-free patients? CNS Drugs 18(4):201–212
Schmidt D (2011) AED discontinuation may be dangerous for seizure-free patients. J Neural Transm (Vienna) 118(2):183–186
Beghi E et al (2013) Withdrawal of antiepileptic drugs: guidelines of the Italian League Against Epilepsy. Epilepsia 54(Suppl 7):2–12
Wang J et al (2015) Genome-wide circulating microRNA expression profiling indicates biomarkers for epilepsy. Sci Rep 5:9522
An N et al (2016) Elevated serum miR-106b and miR-146a in patients with focal and generalized epilepsy. Epilepsy Res 127:311–316
Wang D et al (2016) GC-MS-Based metabolomics discovers a shared serum metabolic characteristic among three types of epileptic seizures. Epilepsy Res 126:83–89
Yan S et al (2017) Altered microRNA profiles in plasma exosomes from mesial temporal lobe epilepsy with hippocampal sclerosis. Oncotarget 8(3):4136–4146
Li R, Hu J, Cao S (2020) The clinical significance of miR-135b-5p and its role in the proliferation and apoptosis of hippocampus neurons in children with temporal lobe epilepsy. Dev Neurosci 42(5–6):187–194
Martins-Ferreira R et al (2020) Circulating microRNAs as potential biomarkers for genetic generalized epilepsies: a three microRNA panel. Eur J Neurol 27(4):660–666
Raoof R et al (2018) Dual-center, dual-platform microRNA profiling identifies potential plasma biomarkers of adult temporal lobe epilepsy. EBioMedicine 38:127–141
Conte G et al (2021) Circulating P2X7 receptor signaling components as diagnostic biomarkers for temporal lobe epilepsy. Cells 10(9):2444
Mo L et al (2019) Association of cerebrospinal fluid zinc-alpha2-glycoprotein and tau protein with temporal lobe epilepsy and related white matter impairment. NeuroReport 30(8):586–591
Maiti R et al (2018) Effect of anti-seizure drugs on serum S100B in patients with focal seizure: a randomized controlled trial. J Neurol 265(11):2594–2601
Walker LE et al (2022) High-mobility group box 1 as a predictive biomarker for drug-resistant epilepsy: a proof-of-concept study. Epilepsia 63(1):e1–e6
Toscano ECB et al (2019) Circulating levels of adipokines are altered in patients with temporal lobe epilepsy. Epilepsy Behav 90:137–141
Wang J et al (2015) Circulating microRNAs are promising novel biomarkers for drug-resistant epilepsy. Sci Rep 5:10201
Niu X et al (2021) MiR-194-5p serves as a potential biomarker and regulates the proliferation and apoptosis of hippocampus neuron in children with temporal lobe epilepsy. J Chin Med Assoc 84(5):510–516
Sun Y et al (2016) Expression of microRNA-129-2-3p and microRNA-935 in plasma and brain tissue of human refractory epilepsy. Epilepsy Res 127:276–283
Wang X et al (2016) Serum microRNA-4521 is a potential biomarker for focal cortical dysplasia with refractory epilepsy. Neurochem Res 41(4):905–912
Avansini SH et al (2017) MicroRNA hsa-miR-134 is a circulating biomarker for mesial temporal lobe epilepsy. PLoS ONE 12(4):e0173060
De Benedittis S et al (2021) Circulating microRNA: the potential novel diagnostic biomarkers to predict drug resistance in temporal lobe epilepsy, a pilot study. Int J Mol Sci 22(2):702
Shen CH et al (2019) Expression of plasma microRNA-145-5p and its correlation with clinical features in patients with refractory epilepsy. Epilepsy Res 154:21–25
Raoof R et al (2017) Cerebrospinal fluid microRNAs are potential biomarkers of temporal lobe epilepsy and status epilepticus. Sci Rep 7(1):3328
Hogg MC et al (2019) Elevation in plasma tRNA fragments precede seizures in human epilepsy. J Clin Invest 129(7):2946–2951
Mirzajani S et al (2020) Expression analysis of lncRNAs in refractory and non-refractory epileptic patients. J Mol Neurosci 70(5):689–698
Wang R et al (2016) Evaluation of serum matrix metalloproteinase-3 as a biomarker for diagnosis of epilepsy. J Neurol Sci 367:291–297
Wang R et al (2016) Serum matrix metalloproteinase-2: a potential biomarker for diagnosis of epilepsy. Epilepsy Res 122:114–119
Zhang H et al (2019) Chitinase-3-like protein 1 may be a potential biomarker in patients with drug-resistant epilepsy. Neurochem Int 124:62–67
Asadollahi M, Simani L (2019) The diagnostic value of serum UCHL-1 and S100-B levels in differentiate epileptic seizures from psychogenic attacks. Brain Res 1704:11–15
Asia P et al (2021) The study of ischemia modified albumin as an early biomarker of epilepsy in adolescent population: a cross-sectional study. Horm Mol Biol Clin Investig 42(2):183–187
Lai Q et al (2022) GluR3B antibody was a biomarker for drug-resistant epilepsy in patients with focal to bilateral tonic-clonic seizures. Front Immunol 13:838389
Stocklin B et al (2015) Copeptin as a serum biomarker of febrile seizures. PLoS ONE 10(4):e0124663
Wang N et al (2021) Serum neuropeptide Y level is associated with post-ischemic stroke epilepsy. J Stroke Cerebrovasc Dis 30(2):105475
Kopczynska M et al (2018) Complement system biomarkers in epilepsy. Seizure 60:1–7
Hong Z et al (2014) Serum brain-derived neurotrophic factor levels in epilepsy. Eur J Neurol 21(1):57–64
Rawat C et al (2020) Downregulation of peripheral PTGS2/COX-2 in response to valproate treatment in patients with epilepsy. Sci Rep 10(1):2546
Mootha VK et al (2003) PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet 34(3):267–273
Subramanian A et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102(43):15545–15550
Subramanian A et al (2017) A next generation connectivity map: L1000 platform and the first 1,000,000 profiles. Cell 171(6):1437-1452 e17
Dang X et al (2022) Correlation of ferroptosis and other types of cell death in neurodegenerative diseases. Neurosci Bull 38(8):938–952
Curinha A et al (2014) Implications of polyadenylation in health and disease. Nucleus 5(6):508–519
Weis S et al (2017) Metabolic adaptation establishes disease tolerance to sepsis. Cell 169(7):1263-1275 e14
Muhoberac BB, Vidal R (2019) iron, ferritin, hereditary ferritinopathy, and neurodegeneration. Front Neurosci 13:1195
Akyuz E et al (2021) Myocardial iron overload in an experimental model of sudden unexpected death in epilepsy. Front Neurol 12:609236
Philpott CC, Jadhav S (2019) The ins and outs of iron: escorting iron through the mammalian cytosol. Free Radic Biol Med 133:112–117
Philpott CC (2018) The flux of iron through ferritin in erythrocyte development. Curr Opin Hematol 25(3):183–188
Santana-Codina N, Gikandi A, Mancias JD (2021) The role of NCOA4-mediated ferritinophagy in ferroptosis. Adv Exp Med Biol 1301:41–57
Cai Y, Yang Z (2021) Ferroptosis and its role in epilepsy. Front Cell Neurosci 15:696889
Sticht C et al (2018) miRWalk: an online resource for prediction of microRNA binding sites. PLoS One 13(10):e0206239
Brodie MJ et al (2012) Patterns of treatment response in newly diagnosed epilepsy. Neurology 78(20):1548–1554
Gesche J et al (2017) Resistance to valproic acid as predictor of treatment resistance in genetic generalized epilepsies. Epilepsia 58(4):e64–e69
Sills GJ, Rogawski MA (2020) Mechanisms of action of currently used antiseizure drugs. Neuropharmacology 168:107966
Funding
This work was supported by grants from the National Natural Science Foundation of China (No. 81871018), the National Natural Science Foundation of China (No. 21575094), and Med-X Center for Informatics funding project (No. YGJC001).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection, and analysis were performed by YD, LK, and YH. The first draft of the manuscript was written by YD, LK, and YH. All authors contributed to discussions regarding the results and the manuscript. LC, ZW, ML, and TL revised the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethical approval
Not available.
Informed consent
No informed consent statement was not required for this study.
Conflict of interest
None.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Duan, Y., Kang, L., He, Y. et al. A pilot study on identifying gene signatures as markers for predicting patient response to antiseizure medications. Neurol Sci 44, 2137–2148 (2023). https://doi.org/10.1007/s10072-023-06605-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10072-023-06605-2