Abstract
Objectives
This healthy volunteer control-based study was conducted to explore alterations of compositions and function of gut microbiota in Chinese pSS patients.
Method
The high-throughput Illumina Miseq sequencing method, targeting the V3-V4 region of the 16S ribosomal RNA (rRNA) gene, was used to compare the microbiota communities between 30 pSS patients and 30 age-matched healthy volunteers. The intestinal dysbiosis of pSS patients was evaluated and its correlation with some disease phenotypes was analyzed. Furthermore, we performed the amino acid sequence alignment analysis to illustrate the molecular mimicry patterns of new microbial peptides.
Results
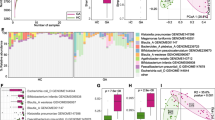
Compared with that in healthy controls, the composition and function of the gut microbiota significantly differed in pSS patients. Certain genera and species, including genera: Escherichia-Shigella, Sardovia, Veillonella, Insteinimonas, and Lactobacillales; species: Escherichia coli, Lactobacillus phage Sal3, Lactobacillus reuteri, Lactobacillus gasseri, Streptococcus lutetiensis, Streptococcus mutans, Scardovia wiggsiae, and Fusobacterrium ulcerans were found to be enriched in the feces of pSS patients, while butyrate-producing bacteria were less abundant in pSS patients. Certain genera (including Lactobacillales) and species (including Lactobacillus gasseri) were associated with disease severity and therapy resistance parameters. Autoantigen epitopes of “WPSALPT, NPARSFG, MNPARSFG, and AFGLAIGT” from aquaporin-5 were aligned perfectly with one enriched microbiota of patients with pSS, namely Escherichia coli.
Conclusions
The composition and function of the gut microbiota significantly differed in pSS patients compared with that in healthy controls. Our study would facilitate the possible research on the role of gut microbiota in the pathogenesis of pSS.
Key Points • The composition and function of the gut microbiota significantly differed in pSS patients compared with that in healthy controls. • Certain genera (including Lactobacillales) and species (including Lactobacillus gasseri) were associated with disease severity and therapy resistance parameters. • The amino acid sequence of Escherichia coli, an enriched species in pSS patients, is similar to those of the pSS-relevant auto-epitopes, such as WPSALPT, NPARSFG, MNPARSFG, and AFGLAIGT, which provides us new clues to explore the exact mechanisms by which Escherichia coli contributes to pSS pathogenesis. |




Similar content being viewed by others
Abbreviations
- pSS:
-
Primary Sjögren’s syndrome
- Treg:
-
FoxP3 + regulatory T cells
- SPF:
-
Specific pathogens
- GF:
-
Germ-free
- SCFA:
-
Short-chain fatty acids
- ACR-EULAR:
-
American Congress of Rheumatology–European League Against Rheumatism
- ESSDAI:
-
EULAR Sjögren’s syndrome disease activity index
- IgG:
-
Immunoglobulin G
- IgA:
-
Immunoglobulin A
- IgM:
-
Immunoglobulin M
- Ro/SSA:
-
Ro/Sjögren’s syndrome autoantibody A
- La/SSB:
-
La/Sjögren’s syndrome autoantibody B
- ANA:
-
Antinuclear antibody
- pSS-FC:
-
PSS with a favourable course
- pSS-STR:
-
Most severe and therapy-resistant pSS
- pSS-R:
-
PSS with therapy-resistant
- pSS-NR:
-
Non-therapy resistant pSS
- AGE:
-
Gel electrophoresis
- OTU:
-
Operational taxonomic units
- RDP:
-
Ribosomal Database Project
- PCoA:
-
Principal coordinate analysis
- PICRUSt:
-
Phylogenetic Investigation of Communities by Reconstruction of Unobserved States
- KEGG:
-
Kyoto Encyclopedia of Genes and Genome
- KO:
-
Orthologics
- IEDB:
-
Immune epitope database
- pDCs:
-
Plasmacytoid dendritic cells
- IFN:
-
Interferon
- SLE:
-
Systemic lupus erythematosus
- OmpA:
-
Outer membrane protein
References
Saraux A, Pers JO, Devauchelle-Pensec V (2016) Treatment of primary Sjogren syndrome. Nat Rev Rheumatol 12(8):456–471
Miossec P, Korn T, Kuchroo VK (2009) Interleukin-17 and type 17 helper T cells. N Engl J Med 361(9):888–898
Nguyen CQ, Hu MH, Li Y, Stewart C, Peck AB (2008) Salivary gland tissue expression of interleukin-23 and interleukin-17 in Sjogren’s syndrome: findings in humans and mice. Arthritis Rheum 58(3):734–743. https://doi.org/10.1002/art.23214
Manfrè V, Cafaro G, Riccucci I, Zabotti A, Perricone C, Bootsma H, De Vita S, Bartoloni E (2020) One year in review 2020: comorbidities diagnosis and treatment of primarySjogren’s syndrome.Clin Exp Rheumatol 38 Suppl 126(4):10–22
Björk A, Mofors J, Wahren-Herlenius M (2020) Environmental factors in the pathogenesis of primary Sjogren’s syndrome. J Intern Med 287(5):475–492
Atarashi K, Tanoue T, Shima T et al (2011) Induction of colonic regulatory T cells by indigenousClostridium species [J]. Science 331:337–341
Cheng H, Guan X, Chen D, Ma W (2019) The Th17/Treg Cell balance: a gut microbiota-modulated story. Microorganisms 7(12):583
Flannigan KL, Denning TL (2018) Segmented filamentous bacteria-induced immune responses: a balancing act between host protection and autoimmunity. Immunology 154(4):537–546
Xu M, Pokrovskii M, Ding Y, Yi R, Au C, Harrison OJ, Galan C, Belkaid Y, BonneauR Littman DR (2018) c-MAF-dependent regulatory T cells mediate immunological tolerance to a gut pathobiont. Nature 554(7692):373–377
Gao Y, Chen Y, Zhang Z, Yu X, Zheng J (2020) Recent advances in mouse models of Sjögren’s syndrome. Front Immunol 11:1158
Vitali C, Bombardieri S, Jonsson R et al (2002) Classification criteria for Sjogren s syndrome: a revised version of the European criteria proposed by theAmerican-European Consensus Group. Ann Rheum Dis 61:554–558
Seror R, Ravaud P, Bowman SJ et al (2010) EULAR Sjogrens syndrome disease activity index: development of a consensus systemic disease activity index for primary Sjogren’s syndrome. Ann Rheum Dis 69:1103–1109
Qian Y, Yang X, Shaoqing Xu, Chunyan Wu, Song Y, Qin N, Chen S-D, Xiao Q (2018) “Alteration of the fecal microbiota in Chinese patients with Parkinson’s disease”, brain, behavior, and immunity. Brain Behav Immun 70:194–202
Edga R (2013) UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10(10):996–8
Yilmaz P, Parfrey LW, Yarza P, Gerken J, Pruesse E, Quast C, Schweer T, Peplies J, Ludwig W, Glöckner FO (2014) The SILVA and “All-species Living Tree Project (LTP)” taxonomic frameworks. Nucleic Acids Res 42(Database issue):D643-8
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215(3):403–410
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7(5):335–6
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215(3):403–410
de Paiva CS, Jones DB, Stern ME, Bian F, Moore QL, Corbiere S, Streckfus CF, Hutchinson DS, Ajami NJ, Petrosino JF, Pflugfelder SC (2016) Altered mucosal microbiome diversity and disease severity in Sjögren syndrome. Sci Rep 6:23561
Cano-Ortiz A, Laborda-Illanes A, Plaza-Andrades I, Membrillo Del Pozo A, Villarrubia Cuadrado A, de Rodríguez Calvo Mora M, Leiva-Gea I, Sanchez-Alcoholado L, Queipo-Ortuño MI (2020) Connection between the gut microbiome, systemic inflammation, gut permeability and FOXP3 expression in patients with primary Sjögren’s syndrome. Int J Mol Sci 21(22):8733
Zegarra-Ruiz DF, El Beidaq A, Iñiguez AJ, Lubrano Di Ricco M, Manfredo Vieira S, Ruff WE, Mubiru D, Fine RL, Sterpka J, Greiling TM, Dehner C, Kriegel MA (2019) A diet-sensitive commensal lactobacillus strain mediates TLR7-dependent systemic autoimmunity. Cell Host Microbe 25(1):113–127
Xu X, Hicks C, Li Y, Su J, Shiloach J, Kaufman JB, Fitz Y, Eichacker PQ, Cui X (2014) Purified cell wall from the probiotic bacterium Lactobacillus gasseri activates systemic inflammation and at higher doses, produces lethality in a rat model. Crit Care 18(4):R140
Tochio T, Kadota Y, Tanaka T, Koga Y (2018) 1-Kestose, the smallest fructooligosaccharide component, which efficiently stimulates Faecalibacterium prausnitzii as well as bifidobacteria in humans. Foods 7(9):140
Zhao H, Xu H, Chen S, He J, Zhou Y, Nie Y (2021) Systematic review and meta-analysis of the role of Faecalibacterium prausnitzii alteration in inflammatory bowel disease. J Gastroenterol Hepatol 6(2):320–328
Hiippala K, Barreto G, Burrello C, Diaz-Basabe A, Suutarinen M, Kainulainen V, Bowers JR, Lemmer D, Engelthaler DM, Eklund KK, Facciotti F, Satokari R (2020) Novel Odoribacter splanchnicus Strain and its outer membrane vesicles exert immunoregulatory effects in vitro. Front Microbiol 11:575455
Nylund L, Nermes M, Isolauri E, Salminen S, de Vos WM, Satokari R (2015Feb) Severity of atopic disease inversely correlates with intestinal microbiota diversity and butyrate-producing bacteria. Allergy 70(2):241–244
Yanagisawa N, Ueshiba H, Abe Y, Kato H, Higuchi T, Yagi J (2018) Outer membrane protein of gut commensal microorganism induces autoantibody production and extra-intestinal gland inflammation in mice. Int J Mol Sci 19(10):3241
Winter SE, Winter MG, Xavier MN, Thiennimitr P, Poon V, Keestra AM, Laughlin RC, Gomez G, Wu J, Lawhon SD, Popova IE, Parikh SJ, Adams LG, Tsolis RM, Stewart VJ, Bäumler AJ (2013) Host-derived nitrate boosts growth of E coli in the inflamed gut. Science 339(6120):708–11
Yanagisawa N, Ueshiba H, Abe Y, Kato H, Higuchi T, Yagi J (2018) Outer membrane protein of gut commensal microorganism induces autoantibody production and extra-intestinal gland inflammation in mice. Int J Mol Sci 19(10):3241
Chen BD, Jia XM, Xu JY, Zhao LD, Ji JY, Wu BX, Ma Y, Li H, Zuo XX, Pan WY, Wang XH, Ye S, Tsokos GC, Wang J, Zhang X (2021) An autoimmunogenic and proinflammatory profile defined by the gut microbiota of patients with untreated systemic Lupus Erythematosus. Arthritis Rheumatol 73(2):232–243
Acknowledgements
We thank the patients and healthy volunteers who made this study possible and provided their fecal samples. We thank Professor Yudong Liu from Beijing Hospital for the manuscript editing.
Funding
This work was supported by grants from the Natural Science Foundation of China (31140008), CAMS Innovation Fund for Medical Science (2021-12 M-1–050), and Beijing Hospital fund (BJ2014033).
Author information
Authors and Affiliations
Contributions
Clinical analyses and manuscript writing: Yongjing Cheng, Zhe Chen.
Sample and data collection: Fang Wang.
Statistical analyses: Yongjing Cheng, Yunzhi Zhufeng, Jun Xu.
Study design:Fang Wang, Yongjing Cheng.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, F., Zhufeng, Y., Chen, Z. et al. The composition and function profile of the gut microbiota of patients with primary Sjögren’s syndrome. Clin Rheumatol 42, 1315–1326 (2023). https://doi.org/10.1007/s10067-022-06451-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10067-022-06451-1