Abstract
The tumor suppressor p53 plays a crucial role in the cellular response to DNA damage by transcriptional activation of numerous downstream genes. Although a considerable number of p53 target genes have been reported, the precise mechanism of p53-regulated tumor suppression still remains to be elucidated. Here, we report a novel role of the DFNA5 gene in p53-mediated etoposide-induced cell death. The DFNA5 gene has been previously reported to be responsible for autosomal-dominant, nonsyndromic hearing impairment. The expression of the DFNA5 gene was strongly induced by exogenous and endogenous p53. The chromatin immunoprecipitation assay indicated that a potential p53-binding sequence is located in intron 1 of the DFNA5 gene. Furthermore, the reporter gene assay revealed that the sequence displays p53-dependent transcriptional activity. The ectopic expression of DFNA5 enhanced etoposide-induced cell death in the presence of p53; however, it was inhibited in the absence of p53. Finally, the expression of DFNA5 mRNA was remarkably induced by gamma-ray irradiation in the colon of p53(+/+) mice but not in that of p53(−/−) mice. These results suggest that DFNA5 plays a role in the p53-regulated cellular response to genotoxic stress probably by cooperating with p53.
Similar content being viewed by others
Introduction
Mutations in the p53 gene are the most common genetic alterations found in human cancers. A normal cell expresses a low level of p53 protein because of its short half-life. However, the expression level and functional activity of p53 increases in response to various cellular stresses. The activated p53 then functions as a transcription factor that activates the transcription of many target genes; this leads to the expression of multiple physiological functions of p53 (Vogelstein et al. 2000; Vousden 2002; Nakamura 2004; Arakawa 2005). Thus far, a number of p53 transcriptional targets have been identified. These include cell-cycle-regulating proteins (p21/WAF1, El-Deiry et al.1993; and hCDC4b, Kimura et al. 2003a), apoptosis-regulating proteins (BAX, Miyashita and Reed 1995; p53AIP1, Oda et al. 2000; Matsuda et al. 2002; Yoshida et al. 2004; and STAG1 Anazawa et al. 2004), DNA damage-repair proteins (p53R2, Tanaka et al. 2000; Yamaguchi et al. 2001; Kimura et al. 2003; and GADD45, Kastan et al. 1992), negative regulators of p53 (MDM2, Barak et al. 1993; and Pirh2, Leng et al. 2003), positive regulators of p53 (p53DINP1, Okamura et al. 2001), and inhibitors of angiogenesis (TSP-1, Dameron et al. 1994; and BAI1, Nishimori et al. 1997). On the other hand, the analysis of a microarray including 6,000 human genes indicated that 107 genes were up-regulated and 54 were down-regulated in cells infected with the adenovirus vector designed to express p53 (Zhao et al. 2000). These observations clearly suggested that a considerable number of p53 target genes have yet to be isolated. Moreover, the identification of these additional p53 target genes is likely to be indispensable for understanding the mechanism of the p53-regulated cellular response in order to prevent tumor formation and progression.
The DFNA5 gene was initially identified as one of the genes whose expression is inversely correlated with that of the estrogen receptor in breast carcinomas, and therefore, it was initially designated as ICERE-1 (inversely correlated with estrogen receptor expression) (Thompson and Weigel 1998). However, later on, a mutation in the DFNA5 gene was reported to be associated with autosomal-dominant, nonsyndromic hearing impairment in an extended Dutch family (Van Laer et al. 1998). An insertion/deletion mutation in intron 7 of this gene caused the skipping of exon 8, which resulted in the premature termination of the open reading frame (Van Laer et al. 1998). Thus far, two additional mutations in DFNA5 have been reported in a Chinese family and another Dutch family. Interestingly, these mutations also cause skipping of exon 8, thereby resulting in the generation of the same aberrant transcript as in the case of the first Dutch family identified with the mutations (Yu et al. 2003; Bischoff et al. 2004). Therefore, it was speculated that hearing impairment may not be due to haploinsufficiency but rather due to a gain-of-function mutation. Furthermore, mutant DFNA5 may have a new deleterious function. Despite these observations, the physiological function of the DFNA5 protein still remains to be elucidated.
Here, we report an unexpected role of the DFNA5 gene in the p53-mediated cellular response to DNA damage; this implies the involvement of DFNA5 in tumorigenesis. These findings will shed light on the mechanism of the p53-dependent pathway for suppressing cancer.
Materials and methods
Cell lines and transfections
Human cancer cell lines HepG2 (hepatoblastoma), H1299 (lung cancer), and COS7 (monkey kidney fibroblast) were purchased from American Type Culture Collection. Human cancer cell line T98G (glioblastoma) was purchased from Human Science Research Resource Bank (HSRRB, Japan). All cell lines were cultured under conditions recommended by their respective depositors.
For transfection, cells were seeded at 2×105 cells per well of six-well plates. After 24 h, cells were transfected with 1 μg of plasmid mixtures pre-incubated for 15 min with 6 μl of Fugene6 transfection reagent (Roche).
DNA-damaging treatments
Cells were seeded 12 h before treatment and were 60–70% confluent at the time of treatment. To examine the expression of DFNA5 in response to genotoxic stresses, HepG2 cells were continuously treated with 1.0 μg/ml adriamycin for 2 h, UV-irradiated at 30 J/m2 using a UV cross-linker (Stratagene), or γ-irradiated at 50 Gy using a 60Co source.
DNA microarray
Total cellular RNA was extracted at the indicated time from HepG2 cells infected with adenovirus vector at 30 moi, which was designed to express wild-type p53 (Ad-p53-WT), a mutant p53 (Ad-p53-46F) or EGFP (Ad-EGFP) (Nakamura et al. 2006). For gene-expression profiling, GeneChip Human Genome U133A and U133B microarrays (Affymetrix, Santa Clara, CA, USA) were used that contain 22,215 and 22,577 probe sets, respectively, to examine a total of about 39,000 transcripts. Target cRNA for microarray hybridization was prepared from 5 μg of total RNA according to the manufacturer’s instructions using a BioArray RNA transcript labeling kit (Enzo Diagnostics, Farmingdale, NY, USA). Hybridization to the microarrays, washing and staining with the antibody amplification procedure, and scanning were also carried out according to the manufacturer’s instructions. The expression value of each gene was calculated and normalized using Affymetrix Microarray Suite software version 5.0.
RNA interference
We established the p53 knock-down (HepG2-p53-KD) and the control (HepG2-p53-Cont) cell lines as described previously (Hara et al. 2004), which were derived from a hepatoblastoma cell line (HepG2 that contains wild-type p53). In brief, HepG2 cells were infected with SI-MSCV-puro-H1R-p53Ri retrovirus for down-regulation of p53 expression and with SI-MSCV-puro-H1R retrovirus for negative control. Then the infected cells were selected with 1 μg/ml puromycin for 2 weeks, and the single clones were isolated.
Northern blotting
Using TRIZOL reagent (GIBCO BRL), total RNAs were extracted at various time points from HepG2 infected with Ad-p53-WT, Ad-p53-46F or Ad-EGFP, and also form HepG2-p53-KD and HepG2-p53-Cont cells subjected to DNA damaging treatments. Total RNAs were further purified to poly(A)+ RNA using mRNA purification kits (TAKARA) according to the manufacturer’s instructions. A 3 μg aliquot of each poly(A)+ RNA was separated on 1% agarose gel containing 1× 4-morpholinepropanesulfonic acid buffer (MOPS) and 2% formaldehyde, and transferred onto a nylon membrane. The blots were hybridized with a random-primed 32P-labeled DNA fragment carrying the coding-sequence of DFNA5, p21/WAF1, or β-actin.
ChIP assay
ChIP assay was performed using the chip assay kit (Upstate Biotechnology) as recommended by the manufacture. HepG2 cells (3×105 cells) were seeded on a 10-cm dish, and infected with Ad-p53-WT, Ad-p53-46F and Ad-EGFP at 30 moi. After 24 h, genomic DNA and protein were cross-linked by adding formaldehyde (1% final concentration) directly into the culture medium and incubated for 15 min at 37°C. Cells were lysed in 200 μl SDS lysis buffer with a protease inhibitor cocktail and sonicated to generate DNA fragments 200–1,000 bp long. After centrifugation, the cleared supernatant was diluted 10-fold with ChIP assay buffer and incubated at 4°C overnight with the specific antibody at 4°C for 16 h with rotation. Immune complexes were precipitated, washed, and eluted as recommended. DNA–protein cross-links were reversed by heating to 65°C for 5 h. DNA was phenol-extracted, ethanol-precipitated and resuspended in 50 μl of Tris–EDTA. We used 2 μl of each sample as a template for PCR amplification. PCR amplifications of DFNA5 intron 1, containing a predictive p53 binding-sites, were performed on immunoprecipitated chromatin using specific primers: 5′-TCTGTGATTGAGTATCCTCGT-3′ (p53BS1, forward) and 5′-CCTGGTGTTGCTGACCATG-3′ (p53BS1, reverse), or 5′-GATGCCCCTCTGTTCTAATT-3′ (p53BS2, forward) and 5′-GCCTCATATGACTCATTCTGT-3′ (p53BS2, reverse), or 5′-GCCAGGCTGGTTTCAAACTC-3′ (p53BS3, forward) and 5′-ATGAAGGATCTAGAACTCAAGT-3′ (p53BS3, reverse), or 5′-ATTCTGATCACAGATCCTCCA-3′ (p53BS4, forward) and 5′-GATGCTGTAGCTGAGGAGC-3′ (p53BS4, reverse), or 5′-ACCTTTCACCATTCCCCTAC-3′ (p21/WAF1, forward) and 5′-GCCCAAGGACAAAATAGCCA-3′ (p21/WAF1, reverse). To ensure that PCR was performed in linear range, template DNA was amplified for a maximum of 30 cycles.
Gene reporter assay
A DNA fragment, including potential p53-binding sites of DFNA5, was amplified by PCR, and cloned into the pGL3-promoter vector (Promega, Madison, WI, USA). The same primers were used for PCR as for the ChIP assay. H1299 cells were plated in six-well culture plates (1.5×105 cells per well) 24 h before cotransfection of 1 μg of reporter plasmid and either 1 μg of wild-type (p53 WT) or mutant p53 (p53 MT) expression vector in combination with 50 ng of pRL-CMV vector (Promega). At 48 h after transfection, cells were rinsed with PBS and lysed in 500 μl of a passive lysis buffer (Promega). Cell lysates were used directly in the Dual Luciferase assay system (Promega), which depends on sequential measurements of firefly and Renilla luciferase activities in specific substrates (beetle luciferin and coelenterazine, respectively). Quantification of luciferase activities and calculation of relative ratios were carried out manually with a luminometer.
Plasmids
The entire coding sequence of DFNA5 cDNA was amplified by PCR using KOD-Plus DNA polymerase (Toyobo), and initially cloned into pCR-Blunt II-TOPO (Invitrogen). The construct plasmid was confirmed by sequencing. Then the EcoRI-digested DNA fragment from the pCR-Blunt II-TOPO plasmid, which includes the entire coding region of DFNA5, was inserted into the EcoRI site of pcDNA3.1(+) (Invitrogen) to prepare the plasmids of pcDNA3.1(+)-sense-DFNA5 and pcDNA3.1(+)-antisense-DFNA5 for colony formation assay, or into the EcoRI site of pCMV-tag2A (Stratagene) to prepare the plasmid of pCMVtag2A/DFNA5 for examination of subcellular localization using anti-FLAG antibody, or into the EcoRI site of pIRES2-EGFP (BD Bioscience) to prepare the plasmid of pIRES2-DFNA5 for establishing the stable transformant expressing the EGFP signal that represents the expression level of DFNA5 mRNA. Primer sequences are forward, 5′-AATGTTTGCCAAAGCAACCAG-3′; reverse, 5′-CATCATGAATGTTCTCTGCCT-3′.
Western blotting
For preparation of whole-cell lysates, adherent and detached cells were collected and lysed in chilled RIPA buffer [1% NP40, 50 mM Tris–HCl at pH8.0, 150 mM NaCl, 0.5% DOC, 0.1% SDS, 1 mM phenyl methylsulphonyl fluoride (PMSF)] for 30 min on ice. Homogenates were centrifuged for 15 min in a microcentrifuge at 4°C and the supernatants were collected and boiled in 5× SDS sample buffer (125 mM Tris–HCl at pH 6.8, 4% SDS, 20% glycerol, 10% β-mercaptoethanol and 0.4 mg/ml Bromophenol Blue). Each sample (10 μg) was loaded onto a 12% SDS–polyacrylamide gel electrophoresis (PAGE) gel and blotted onto a nitrocellulose membrane (Amersham). Protein bands on Western blots were visualized by chemiluminescent detection (ECL plus; Amersham).
Immunocytochemistry
Adherent cells were fixed with 4% paraformaldehyde in PBS and permeabilized with 0.2% Triton X-100 in PBS for 5 min at room temperature. The cells were then covered with blocking solution (3% BSA/PBS containing 0.2% Triton X-100) for 30 min at room temperature and incubated with mouse anti-FLAG antibody (diluted 1:250) in blocking solution for 60 min at room temperature. Primary antibody was stained with rat anti-mouse secondary antibody conjugated to fluorescein isothiocyanate (FITC; diluted 1:1,000) for 1 h at room temperature, stained with PI and visualized with a Radiance 2000 (Bio-Rad).
Semi-quantitative RT-PCR analysis
The RT-PCR exponential phase was determined on 15–30 cycles to allow semi-quantitative comparisons among cDNAs developed from identical reactions. Each PCR regime involved a 5-min initial denaturing step at 94°C, followed by 23 cycles (for DFNA5) or 17 cycles (for β2MG) at 94°C for 30 s, 55°C for 30 s and 72°C for 30 s on a Gene Amp PCR system 9700 (Applied Biosystems, Foster City, CA, USA). Primer sequences were, for DFNA5: forward, 5′-ACCTTGTGGCCTTCCTGCA-3′ and reverse, 5′-GACTAATGTCAGCTGAGGCA-3′; and for β2MG: forward, 5′-TAGCTGTGCTCGCGCTACT-3′ and reverse, 5′-GCGCTACTCTCTCTTCTG-3′.
Colony-formation assay
H1299 and T98G cells were plated at 1×105 cells per dish in 10-cm culture dishes 24 h before transfection. For each expression vector [pcDNA3.1(+)-sense-DFNA5, pcDNA3.1(+)-antisense-DFNA5, or pcDNA3.1(+)], 6 μg was transfected using 18 μl of FuGENE6 reagent (Roche). Forty-eight hours later, the harvested cells were diluted and replated in 10-cm culture dishes. Transfected cells were allowed to grow in the presence of 0.8 mg/ml G418 (GIBCO) for 2 weeks. The colonies formed from each cell were fixed with 10% formalin, stained with crystal violet, and counted. To confirm the expression of DFNA5 in each dish, RT-PCR was constantly performed.
Stable transformants
H1299 cells (3×105cells) were plated in a 6-cm dish 12 h before transfection of 2 μg of each expression vector (pIRES2-DFNA5 or pIRES2-EGFP) using 6 μl of FuGENE6 reagent (Roche). Forty-eight hours later, the harvested cells were diluted and replated in 10-cm culture dishes. Stable transformants were selected in the presence of 0.8 mg/ml G418 (GIBCO). Resistant clones were expanded, selected for EGFP and tested for DFNA5 by RT-PCR. At least 20 colonies were tested in each case.
Apoptosis assay
Cells were trypsinized, washed, collected and fixed with 70% ethanol at the indicated times. Fixed samples were centrifuged, treated with RNase (1 mg/ml), and resuspended in propidium iodide (50 μg/ml). The stained cells were analyzed on a Becton Dickinson FACScan flow cytometer. The sub-G1 fraction of the cells was counted as the apoptotic cells, and the proportion of the apoptotic cells to the total cells was indicated as percentage.
p53 knock-out mice
p53-deficient mice were a gift from Dr. Aizawa, Center for Developmental Biology, RIKEN (Tsukada et al. 1993). p53 (+/+) and p53 (−/−) mice were irradiated by 10 Gy gamma-rays, and the indicated organs were isolated 24 h after irradiation. Total RNAs were purified from the tissues, from which cDNAs were synthesized, and were subjected to RT-PCR analysis. All mouse procedures were carried out according to the recommendations of the Institutional Animal Care and Use Committee of the National Cancer Center at Tsukiji, Japan.
Results
In order to identify the additional p53-regulated genes, we applied microarray analysis for screening the p53-inducible transcripts in a hepatocellular carcinoma cell line (HepG2) infected with adenovirus p53 (Ad-p53), adenovirus p53-46F (Ad-p53-46F), or adenovirus EGFP (Ad-EGFP). p53-46F was previously demonstrated to be an activated form of p53 with an enhanced ability to induce p53-dependent apoptosis (Nakamura et al. 2006). The RNAs were isolated from the infected cells at 0, 6, 12, 24, and 48 h post-infection and were subjected to microarray analysis. As shown in Fig. 1a, the expression of the p53 mRNA increased in a time-dependent manner in the cells that were infected with Ad-p53 or Ad-p53-46F (Fig. 1a). The p21/WAF1 mRNA was also inducible in concert with the expression of p53 in these cells (Fig. 1a). In this system, although nearly 100 genes were likely to be up-regulated by p53, we focused on the expression of the DFNA5 gene, because the expression of DFNA5 showed a clear increase in a p53- or p53-46F-dependent manner (Fig. 1a). The gene was previously reported to be responsible for autosomal-dominant, nonsyndromic hearing impairment.
Identification of DFNA5 as a p53-inducible gene. a Identification of DFNA5 as a p53-inducible gene by microarray analysis. The expression of DFNA5, p21/WAF1, and p53 mRNA in HepG2 cells at the indicated times after infection with Ad-p53-WT, Ad-p53-46F, and Ad-EGFP are indicated by the green and red intensities of the microarray analysis. The red and green intensities indicate up- and down-regulated expression of the genes, respectively. b, c Northern blot analysis of DFNA5 mRNA. The expression of DFNA5 mRNA in HepG2 cells infected with Ad-p53-WT, Ad-p53-46F, and Ad-EGFP at the indicated times after infection (b), and in HepG2-p53-Cont and HepG2-p53-KD cells at the indicated times after treatment with 1 μg/ml adriamycin (c). p21/WAF1 and β-actin were used as the positive and loading controls, respectively
To confirm the results of the microarray analysis, we performed a Northern blot analysis. As shown in Fig. 1b, the expression of DFNA5 was specifically induced by exogenous p53 protein derived from either Ad-p53 or Ad-p53-46F; these observations are consistent with those of the microarray analysis.
To examine whether endogenous p53 activates the transcription of DFNA5 in response to genotoxic stresses, we established a p53 knock-down (HepG2-p53-KD) cell line derived from a hepatoblastoma cell line (HepG2 that contains wild-type p53) in which the expression level of p53 was inhibited by siRNA. We also established a control cell line that was infected with a retrovirus and was devoid of any knock-down sequences (HepG2-p53-Cont). The expression level of p53 protein was clearly elevated in the HepG2-p53-Cont cells in response to DNA damage whereas it was not at all in the HepG2-p53-KD cells (data not shown). In concert with the expression of p53, the expression of DFNA5 showed an increase in the HepG2-p53-Cont cells but not in the HepG2-p53-KD cells, although both the cell types were treated with 1 μg/ml adriamycin (Fig. 1c). This implies that endogenous activated p53 transactivated the DFNA5 gene.
Further, we examined whether the expression of DFNA5 is inducible in response to various cellular stresses in a p53-dependent manner. Both HepG2-p53-Cont and HepG2-p53-KD cells were treated with H2O2, UV (30 J/m2) or gamma rays (50 Gy), and the expression level of DFNA5 was examined by RT-PCR. As shown in Fig. 2, the expression of DFNA5 as well as p21/WAF1 mRNA was elevated in response to all three types of cellular stresses in the HepG2-p53-Cont cells but not in the HepG2-p53-KD cells. These results suggest that the transcription of DFNA5 is activated by endogenous p53 in response to various cellular stresses.
Endogenous p53-dependent induction of DFNA5 transcription in response to various cellular stresses. The expression of the DFNA5 mRNA is indicated by semi-quantitative RT-PCR in HepG2-p53-Cont and HepG2-p53-KD cells at the indicated times after treatment with H2O2 (100 μM), UV (30 J/m2), or γ-rays (50 Gy). p21/WAF1 and β-MG were used as the positive and loading control, respectively. The PCR cycles for DFNA5, p21/WAF1, and β-MG were 24, 19, and 18, respectively
To determine whether the transcription of DFNA5 is directly regulated by p53, we searched for candidate p53-binding sequence(s) in the genomic region of the DFNA5 gene. Eventually, we found four possible sequences in intron 1 that showed an approximately 75% or greater match to the consensus p53-binding sequence, i.e., two copies of the 10-bp motif, 5′-PuPuPuC(A/T)(A/T)GpyPyPy-3′, separated by any 0- to 13-bp sequence (Fig. 3a). To evaluate the in vivo binding of p53 with these sequences, we performed a chromatin immunoprecipitation (ChIP) assay for these sequences. A protein complex including the p53 protein and its associated genomic DNAs was immunoprecipitated with an anti-p53 antibody, and the genomic DNAs were subjected to PCR analysis for the four candidate sequences. As indicated in Fig. 3b, the fragments containing the BS1, BS2, and BS3 sequences were amplified with the precipitated genomic DNAs, whereas that containing the BS4 sequence was not; this suggests that BS1, BS2, and BS3 probably interact with the p53 protein in vivo.
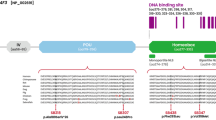
DFNA5 is a direct target gene for p53. a Identification of potential p53-binding site (p53BS) in the genomic DNA of DFNA5. Four candidate p53BS sequences (p53BS1 to p53BS4) were found in intron 1 of DFNA5. The p53BS1, p53BS2, p53BS3, and p53BS4 sequences are shown here. b p53 interacts with p53BS1, p53BS2, and p53BS3 of DFNA5. The ChIP assay was performed on genomic fragments containing p53BS1, p53BS2, p53BS3, and p53BS4. The cell lysates were isolated from Ad-p53-infected (lanes 1–4), Ad-EGFP-infected (lanes 5–8), or noninfected (lanes 9–12) HepG2 cells. A protein complex including p53 and the genomic DNA was precipitated with the anti-p53 antibody (lanes 2, 6, 10) before PCR amplification. The input chromatin represents a portion of the sonicated chromatin before immunoprecipitation (lanes 1, 5, 9). The immunoprecipitates with the anti-FLAG antibody (lanes 3, 7, 11) or in the absence of an antibody (lanes 4, 8, 12) were used as negative controls. p21/WAF1 was used as the positive control. c The p53-responsive sequence of DFNA5. The heterologous luciferase reporter plasmid containing p53BS1, p53BS2, or p53BS3 was cotransfected with the plasmid designed to express wild-type p53 (p53WT), mutant p53 (p53MT), or no p53 (mock) in the H1299 cells. The luciferase activity 24 h after transfection is shown in relation to the activity of the pGL3-promoter vectors without p53BSs
In order to determine whether BS1, BS2, or BS3 is responsible for p53-dependent inducibility of DFNA5, each 300- to 400-bp DNA fragment containing either BS1, BS2, or BS3 was cloned upstream to the SV40-promoter in the pGL3-promoter vector (Promega, Madison, WI, USA); the new vectors were designated as pGL-pro-BS1, pGL-pro-BS2 and pGL-pro-BS3, respectively. Either pGL-pro-BS1, pGL-pro-BS2, or pGL-pro-BS3 was cotransfected with the wild-type p53 expression plasmid (WT-p53) into H1299 (p53-null) cells. As shown in Fig. 3c, the transcriptional activity of luciferase was strongly enhanced only when pGL-pro-BS1 was cotransfected with WT-p53. In contrast, the cotransfection of pGL-pro-BS2 or BS3 with WT-p53 did not reveal any increase in the luciferase activity (Fig. 3c). Moreover, mutant p53 did not activate the transcription of pGL-pro-BS1 at all. Taken together, these results indicate that BS1 is clearly a p53-responsive sequence of the DFNA5 gene and that DFNA5 is a bona fide target of the tumor suppressor p53.
To investigate the role of DFNA5 in p53-mediated functions, we initially examined the subcellular localization of the DFNA5 protein. For this purpose, we designed a plasmid to express DFNA5 with a FLAG-tag at the N-terminus (N-FLAG-DFNA5). The N-FLAG-DFNA5 plasmid was transfected into HepG2 cells, and the protein was immunostained with an anti-FLAG antibody. As indicated by Western blotting in Fig. 4a, the DFNA5 protein was highly expressed in HepG2 cells transfected with N-FLAG-DFNA5. Furthermore, in the same cells, an enormous proportion showed strong staining of the protein in the cytoplasm, and in a subset of the cells, the signal was detected in the nucleus as well (Fig. 4b). Since the previous study reported that GFP-DFNA5 was localized in cytoplasm (Van Laer et al.2004), and the antibody against endogenous DFNA5 is not still available, we suggest that DFNA5 may be a cytoplasmic protein; however, the observation should be confirmed by the antibody that reacts with endogenous DFNA5.
Subcellular localization of the DFNA5 protein. a Western blot analysis of the DFNA5 protein. N-FLAG-DFNA5 and N-FLAG-empty plasmids were transfected into H1299 cells and the cell lysates were subjected to Western blot analysis. The expression of the DFNA5 protein was detected with the anti-FLAG antibody in the cells transfected with N-FLAG-DFNA5 but not in the cells with N-FLAG-empty plasmids. b Immunocytochemical analysis of the DFNA5 protein. In the same cells as in a, the DFNA5 protein was immunostained with the anti-FLAG antibody and detected by the FITC signal (green). The nuclear DNA was stained with PI (red). The nuclear staining of DFNA5 is indicated by arrows
Next, we examined if overexpression of DFNA5 affected the growth of cancer cells. We prepared two plasmids that were designed to express sense and antisense DFNA5, i.e., pcDNA3.1(+)-sense-DFNA5 and pcDNA3.1(+)-antisense-DFNA5, respectively. These plasmids and the empty vector were transfected into a lung-cancer cell line (H1299) and a glioblastoma cell line (T98G). As shown in Fig. 5a, although we were not able to examine the protein level of DFNA5, we confirmed that the mRNAs of the sense and antisense DFNA5 sequences were expressed in the H1299 cells. The cells were then cultured in the presence of G418 and the colonies were counted 2 weeks later. We observed the growth suppression effect of DFNA5 in the H1299 cells to some extent (Fig. 5b). In the T98G cells, there was no significant difference in the colony number between the sense and antisense DFNA5 plasmids (Fig. 5b).
The effect of ectopically expressed DFNA5 on cell growth of two cancer cell lines, H1299 and T98G. a The expression of DFNA5 mRNA in the transfected cell lines. RT-PCR results indicated the expression levels of DFNA5 mRNA in H1299 cell lines 24 h after transfection of the pcDNA3.1(+)-sense-DFNA5, pcDNA3.1(+)-antisense-DFNA5, or pcDNA3.1(+). b Colony formation assay. G418-resistant colonies grown from H1299 or T98G cells transfected with either pcDNA3.1(+), pcDNA3.1(+)-sense-DFNA5, or pcDNA3.1(+)-antisense-DFNA5. The experiments were performed three times using triplicate samples, and the average scores are indicated with error bars on the histogram below
The down-regulation of DFNA5 expression was previously reported to contribute to acquired etoposide resistance in melanoma cells, implying that DFNA5 may play a positive role in etoposide-induced apoptosis (Lage et al. 2001). On the other hand, in the present study, we have clearly demonstrated that the DFNA5 gene is a direct target gene of the tumor suppressor p53. Therefore, we postulated that DFNA5 might be involved in the p53-regulated cellular response to DNA damage. To validate this hypothesis, we have established a stable transformant expressing DFNA5 using the H1299 cells (p53-null). The DFNA5 sense sequence was cloned into the pIRES2-EGFP vector that has the internal ribosomal site of the encephalomyocarditis virus between the multiple cloning site (MCS) and the EGFP gene to be translated from a single bicistronic mRNA. Therefore, it is possible to monitor the expression of DFNA5 by checking the EGFP signal. Eventually, we selected two independent clones: pIRES-DFNA5-18 and pIRES-DFNA5-21. As indicated in Fig. 6a and b, the former expresses DFNA5 mRNA at very low levels, whereas the latter expresses it at higher levels. Using these cell lines, we examined the apoptotic cells after treatment with 1 μg/ml etoposide in the presence or absence of wild-type p53. Unexpectedly, the expression of DFNA5 inhibited etoposide-induced apoptosis in the absence of p53 (Fig. 6c). However, in the presence of p53, high levels of DFNA5 expression enhanced etoposide-induced apoptosis in the late phase (Fig. 6d). These results suggest that DFNA5 may play dual roles in etoposide-induced apoptosis. That is, in the absence of p53, DFNA5 contributes to the resistance of etoposide-induced apoptosis, whereas in the presence of p53, DFNA5 may be positively involved in p53-mediated apoptosis in response to etoposide-induced DNA damage.
Positive and negative roles of DFNA5 in etoposide-induced cell death and its p53-dependency. a The expression levels of EGFP in the transformed cell lines. The expression levels of EGFP in two independent clones, pIRES2-DFNA5-18 and pIRES-DFNA5-21, were found to monitor the expression levels of DFNA5. b The expression levels of DFNA5 mRNA in the transformed cell lines. RT-PCR results indicated the expression levels of DFNA5 mRNA in pIRES2-DFNA5-18 and pIRES2-DFNA5-21. c Antiapoptotic role of DFNA5 in etoposide-induced cell death in the absence of p53. Each cell line was treated with 1 μg/ml etoposide. Apoptotic cells were evaluated by FACS scan analysis at the indicated times after the treatment. d Proapoptotic role of DFNA5 in etoposide-induced cell death in the presence of p53. Each cell line was infected with Ad-p53 (10 moi), and 12 h later, the cells were treated with 1 μg/ml etoposide. Apoptotic cells were evaluated by FACS scan analysis at the indicated times after the treatment. All experiments were repeated independently three times in order to confirm the reproducibility of the results (c, d). The averages of apoptotic cells are shown with error bars. The representative result is shown
The most important question is how and where does DFNA5 function as a mediator of p53? To address this issue, we examined the p53-dependent induction of DFNA5 in various tissues of mice. We irradiated p53(+/+) and p53(−/−) mice with gamma rays, and the RNAs were isolated from the thymus, colon, brain, spleen, and small intestine 24 h after radiation. Interestingly, p53-dependent induction of DFNA5 in response to DNA damage was observed in the colon (Fig. 7). Moreover, p53-dependent expression of DFNA5 was observed in the brain and colon in the absence of DNA damage (Fig. 7). These data strongly suggest that DFNA5 may play an important role in the p53-mediated cellular response to DNA damage in various tissues other than the cochlea.
Potential in vivo role of DFNA5 in p53-regulated response to DNA damage. RT-PCR results indicated expression levels of m-DFNA5 and m-p21/WAF1 mRNA in various tissues, including thymus, colon, lung, brain, spleen, and small intestine tissues. Each tissue was isolated from p53+/+ or p53−/− mice 24 h after irradiation with γ-ray (10 Gy). The number of cycles for each RT-PCR is indicated in parentheses
Discussion
It has been previously reported that the down-regulation of DFNA5 contributes to acquired etoposide-resistance in melanoma cells (Lage et al. 2001). Etoposide is a DNA topoisomerase II inhibitor that induces double-strand DNA breaks, resulting in apoptosis in cancer cells. Therefore, etoposide is one of the commonly used anticancer drugs in cancer therapy. In one study, DFNA5 mRNA was identified by utilizing a differential display method, and its expression in the etoposide-resistant human melanoma cell line MeWo-ETO-1 was found to be remarkably decreased as compared to that in the parental nonresistant human melanoma cell line MeWo (Lage et al. 2001). Further, it was demonstrated that the enforced expression of DFNA5 in MeWo-ETO-1 cells showed approximately 30–35% increased sensitivity to etoposide as compared to that in the parental MeWo-ETO-1 cells; this increased sensitivity is responsible for the increase in the number of apoptotic cells and the caspase-3 activity (Lage et al. 2001). Therefore, it was suggested that DFNA5 is positively involved in DNA damage-induced apoptosis. In contrast, in the present study, the ectopic expression of DFNA5 alone in H1299 cells reduced the number of etoposide-induced apoptotic cells (Fig. 6c, left), thereby suggesting that DFNA5 may have some role in cell survival following the genotoxic stress caused by etoposide in H1299 cells. Conversely, the coexpression of DFNA5 with wild-type p53 in the H1299 cells enhanced cell death 72 h after DNA damage (Fig. 6c, right), as seen in the case of MeWo-ETO-1 cells.
How do we explain such contrasting dual roles of DFNA5 in the cellular response to DNA damage? Our data in the case of the H1299 cells clearly indicated that DFNA5 might contribute to cell survival in the absence of p53, whereas in the presence of p53, DFNA5 may exert a positive role in DNA damage-induced cell death. The role of DFNA5 in genotoxin-induced cell death is probably dependent on the presence or absence of p53.
Although p53 mutations are very rare in most melanomas, the melanoma cell line MeWo was reported to contain a p53 nonsense mutation at codon 341 (Albino et al. 1994). According to our hypothesis, theoretically, if the function of p53 is completely impaired in MeWo cells, DFNA5 may not contribute to DNA damage-induced apoptosis in these cells. In other words, if MeWo cells do not contain mutant p53, the expression of DFNA5 in the etoposide-resistant MeWo-ETO-1 cells might increase the number of apoptotic cells in response to etoposide-induced DNA damage. On the other hand, it may be possible that the MeWo mutant might not cause a complete loss of the p53 functions. Thus, a small increase (30–35%) in sensitivity was obtained by the expression of DFNA5 despite the presence of mutant p53. Consistent with this notion, a very recent study reported that a small pharmacological compound activated the transcription of p21/WAF1, Bax, and PUMA in MeWo cells, resulting in the activation of the p53-dependent apoptotic pathway (Ho and Li 2005). This observation is in agreement with the idea that the function of p53 in MeWo cells may not be completely impaired.
Thus far, the physiological function of DFNA5 remains to be elucidated. The sequence of the DFNA5 protein revealed no significant homology with any other protein. However, a study has revealed that DFNA5 shares sequence similarity with the Mcm10 family of DNA replication proteins in a small region (~50 a.a.) of a conserved zinc finger-like motif referred to as the CCCH domain, which is essential for the Mcm10 function (Gregan et al. 2003). Previously, it was known that a mutation in DFNA5 is associated with nonsyndromic hearing impairment and that it generates a new protein that is derived from the DFNA5 transcript lacking exon 8. In the above-mentioned study, the ectopic expression of mutant DFNA5 led to cell-cycle arrest at the G1 or early S phase in yeast cells, whereas overexpression of wild-type DFNA5 did not reveal any change in the phenotype, implying that DFNA5 mutation results in a gain of function and that mutant DFNA5 may have a new deleterious function (Gregan et al. 2003). Further, another study has recently demonstrated that overexpression of disease-associated mutant DFNA5 in mammalian cells induces cell death; however, wild-type DFNA5 does not (Van Laer et al. 2004). Therefore, mutations of the DFNA5 gene that occur in patients with hearing impairment are unlikely to result in loss of function of wild-type DFNA5.
Next, in order to address the question regarding the physiological function of the DFNA5 protein, DFNA5 knock-out (KO) mice were generated and analyzed (Van Laer et al. 2005). Although specific deletion of exon 8 alone was attempted in order to mimic the human disease in these mice, the DFNA5 protein that is derived from the resulting transcript lacking exon 8 was not detected; this implies that the DFNA5-KO-mice lacked the DFNA5 protein completely (Van Laer et al. 2005). Surprisingly, although there were some significant differences in the number of fourth row outer hair cells between DFNA5-WT and DFNA5-KO mice, neither hearing impairment nor other remarkable abnormalities were detected in the DFNA5-KO mice, as observed in the case of p53-KO-mice (Van Laer et al. 2005). These observations supported the notion that the mutation of the DFNA5 gene that occurs in hearing impairment does not cause loss of function of wild-type DFNA5 but rather acquires a new toxic function. The data also suggest the possibility that the normal function of wild-type DFNA5 is not related to hearing and that wild-type DFNA5 may play a small role in the cochlea.
Furthermore, does DFNA5 mediate the p53-regulated cellular response outside the cochlea? To address this question, we examined the p53-regulated induction of DFNA5 mRNA in response to gamma-ray irradiation in various tissues, including the thymus, colon, brain, spleen, lung, and small intestine. Interestingly, the expression level of DFNA5 mRNA was remarkably increased 24 h after gamma-ray irradiation in the colon of p53(+/+) mice but not in that of p53(−/−) mice, implying that DFNA5 may play a role in p53-mediated cellular response to DNA damage in the colon. Intestinal tissues, including the colon and small intestine, are well-known target organs for p53-induced apoptosis following gamma-ray irradiation (Merritt et al. 1994; Fei et al. 2002). Therefore, along with our observations on the involvement of DFNA5 in etoposide-induced cell death and p53-regulated expression, we speculate that DFNA5 might be one of the mediators of p53 that is involved in p53-induced apoptosis in response to DNA damage.
In the present study, we reported an unexpected role of DFNA5 as a p53 target gene in the p53-regulated cellular response to DNA damage. The mechanism for this phenomenon still remains unclear. However, a clarification on the role of DFNA5 in the cellular response to DNA damage will certainly shed light on the mechanism of the p53-regulated apoptotic pathway. Further, careful investigation is necessary to unravel the role of this interesting molecule in tumorigenesis.
References
Albino AP, Vidal MJ, McNutt NS et al (1994) Mutation and expression of the p53 gene in human malignant melanoma. Melanoma Res 4:35–45
Anazawa Y, Arakawa H, Nakagawa H, Nakamura Y (2004) Identification of STAG1 as a key mediator of a p53-dependent apoptotic pathway. Oncogene 23:7621–7627
Arakawa H (2005) p53, apoptosis and axon-guidance molecules. Cell Death Differ 12:1057–1065
Barak Y, Juven T, Haffner R, Oren M (1993) Mdm2 expression is induced by wild type p53 activity. EMBO J 12:461–468
Bischoff AM, Luijendijk MW, Huygen PL et al (2004) A novel mutation identified in the DFNA5 gene in a Dutch family: a clinical and genetic evaluation. Audiol Neurootol 9:34–46
Dameron KM, Volpert OV, Tainsky MA, Bouck N (1994) Control of angiogenesis in fibroblasts by p53 regulation of thrombospondin-1. Science 265:1582–1584
El-Deiry W, Tokino T, Velculescu VE et al (1993) WAF1, a potential mediator of p53 tumor suppression. Cell 75:817–825
Fei P, Bernhard EJ, El-Deiry WS (2002) Tissue-specific induction of p53 targets in vivo. Cancer Res 62:7316–7327
Gregan J, Van Laer L, Lieto LD, Van Camp G, Kearsey SE (2003) A yeast model for the study of human DFNA5, a gene mutated in nonsyndromic hearing impairment. Biochim Biophys Acta 1638:179–186
Hara S, Nakashima S, Kiyono T et al (2004) p53-independent ceramide formation in human glioma cells during gamma-irradiation-induced apoptosis. Cell Death Differ 11:853–861
Ho CK, Li G (2005) Mutant p53 melanoma cell lines respond differentially to CP-31398-induced apoptosis. Br J Dermatol 153:900–910
Kastan MB, Zhan Q, El-Deiry WS et al (1992) A mammalian cell cycle checkpoint pathway utilizing p53 and GADD45 is defective in ataxia-telangiectasia. Cell 71:587–597
Kimura T, Gotoh M, Nakamura Y, Arakawa H (2003a) hCDC4b, a regulator of cyclin E, as a direct transcriptional target of p53. Cancer Sci 94:431–436
Kimura T, Takeda S, Sagiya Y, Gotoh M, Nakamura Y, Arakawa H (2003b) Impaired function of p53R2 in Rrm2b-null mice causes severe renal failure through attenuation of dNTP pools. Nat Genet 34:440–445
Lage H, Helmbach H, Grottke C, Dietel M, Schadendorf D (2001) DFNA5 (ICERE-1) contributes to acquired etoposide resistance in melanoma cells. FEBS Lett 494:54–59
Leng RP, Lin Y, Ma W et al (2003) Pirh2, a p53-inducible ubiquitin-protein ligase, promotes p53 degradation. Cell 112:779–791
Matsuda K, Yoshida K, Nakamura K, Nakamura Y, Arakawa H (2002) p53AIP1 regulates the mitochondrial apoptotic pathway. Cancer Res 62:2883–2889
Merritt AJ, Potten CS, Kemp CJ et al (1994) The role of p53 in spontaneous and radiation-induced apoptosis in the gastrointestinal tract of normal and p53-deficient mice. Cancer Res 54:614–617
Miyashita T, Reed JC (1995) Tumor suppressor p53 is a direct transcriptional activator of the human BAX gene. Cell 80:293–299
Nakamura Y (2004) Isolation of p53-target genes and their functional analysis. Cancer Sci 95:7–11
Nakamura Y, Futamura M, Kamino H, Yoshida K, Nakamura Y, Arakawa H (2006) Identification of p53–46F as a super p53 with an enhanced ability to induce p53-dependent apotosis. Cancer Sci 97:633–641
Nishimori H, Shiratsuchi T, Urano T et al (1997) A novel brain-specific p53-target gene, BAI1, containing thrombospondin type 1 repeats inhibits experimental angiogenesis. Oncogene 15:2145–2150
Oda K, Arakawa H, Tanaka T et al (2000) p53AIP1, a potential mediator of p53-dependent apoptosis, and its regulation by Ser-46-phophorylated p53. Cell 102:849–862
Okamura S, Arakawa H, Tanaka T et al (2001) p53DINP1, a p53-inducible gene, regulates p53-dependent apoptosis. Mol Cell 8:85–94
Tanaka H, Arakawa H, Yamaguchi T et al (2000) A ribonucleotide reductase gene involved in a p53-dependent cell-cycle checkpoint for DNA damage. Nature 404:42–49
Thompson DA, Weigel RJ (1998) Characterization of a gene that is inversely correlated with estrogen receptor expression (ICERE-1) in breast carcinomas. Eur J Biochem 252:169–177
Tsukada T, Tomooka Y, Takai S et al (1993) Enhanced proliferative potential in culture of cells from p53-deficient mice. Oncogene 8:3313–3322
Van Laer L, Huizing EH, Verstreken M et al (1998) Nonsyndromic hearing impairment is associated with a mutation in DFNA5. Nat Genet 20:194–197
Van Laer L, Vrijiens K, Thys S et al (2004) DFNA5: hearing impairment exon instead of hearing impairment gene? J Med Genet 41:401–406
Van Laer L, Pfister M, Thys S et al (2005) Mice lacking Dfna5 show a diverging number of cochlear fourth row outer hair cells. Neurobiol Dis 19:386–399
Vogelstein B, Lane D, Levine AJ (2000) Surfing the p53 network. Nature 408:307–310
Vousden KH (2002) Live or let die: the cell’s response to p53. Nat Rev Cancer 2:594–604
Yamaguchi T, Matsuda K, Sagiya Y et al (2001) p53R2-dependent pathway for DNA synthesis in a p53-regulated cell cycle checkpoint. Cancer Res 61:8256–8262
Yoshida K, Monden M, Nakamura Y, Arakawa H (2004) Adenovirus-mediated p53AIP1 gene transfer as a new strategy for treatment of p53-resistant tumors. Cancer Sci 95:91–97
Yu C, Meng X, Zhang S, Zhao G, Hu L, Kong X (2003) A 3-nucleotide deletion in the polypyrimidine tract of intron 7 of the DFNA5 gene causes nonsyndromic hearing impairment in a Chinese family. Genomics 82:575–579
Zhao R, Gish K, Murphy M et al (2000) Analysis of p53-regulated gene expression patterns using oligonucleotide arrays. Genes Dev 14:981–993
Acknowledgments
We thank S. Usuda, A. Ishizaka and I. Hyo for their excellent technical assistance. We also thank S. Aizawa for the p53-deficient mice. This work was supported in part by grants from the Ministry of Health, Labor and Welfare, Japan, and the Ministry of Education, Culture, Sports, Science and Technology, Japan, and supported in part by the program for promotion of Fundamental Studies in Health Sciences of the Pharmaceuticals and Medical Devices Agency (PMDA).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Masuda, Y., Futamura, M., Kamino, H. et al. The potential role of DFNA5, a hearing impairment gene, in p53-mediated cellular response to DNA damage. J Hum Genet 51, 652–664 (2006). https://doi.org/10.1007/s10038-006-0004-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-006-0004-6
Keywords
This article is cited by
-
Pyroptosis: a double-edged sword in lung cancer and other respiratory diseases
Cell Communication and Signaling (2024)
-
The multifaceted roles of GSDME-mediated pyroptosis in cancer: therapeutic strategies and persisting obstacles
Cell Death & Disease (2023)
-
GSDME-mediated pyroptosis promotes the progression and associated inflammation of atherosclerosis
Nature Communications (2023)
-
The emerging role of pyroptosis in pediatric cancers: from mechanism to therapy
Journal of Hematology & Oncology (2022)
-
Inverse regulation of GSDMD and GSDME gene expression during LPS-induced pyroptosis in RAW264.7 macrophage cells
Apoptosis (2022)