Abstract
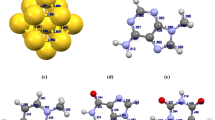
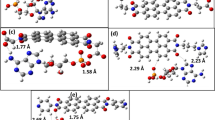
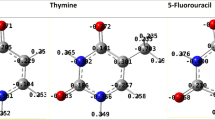
In the present work, CHARMM force field parameters are generated for a cationic oligomer of N, N, N-trimethyl-3-(4-methylthiophen-3-yl) oxy) propan-1-aminium) which has the potential for sensing biological molecules such as nucleic acids, nucleobases. We have used ffTK (force field tool kit) to obtain potential parameters. MD simulations are performed for 20-mer and its complexes with AMP and ATP. The simulation results are analyzed to see the number of phosphates in adenosine nucleotides effects on the structure of the backbone of oligomer. The UV-VIS calculations for the conformers which possess the most probable radius of gyration are carried out and compared to the experimental ones to validate the generated force field.

Recent studies have shown that, biologically important anions (ATP, AMP, vb.) change the spectroscopic properties of cationic polythiophenes (CPT) in the solutions. This work aims to generate CHARMM compatible force field parameters for a CPT to explain experimental studies. The type of interactions will be investigated deeply to lead new biosensor studies by examining the formation and the structure of complexes that consist of a oligothiophene and biological molecules, ATP, AMP by molecular dynamic simulations.









Similar content being viewed by others
References
Brooks BR, Bruccoleri RE, Olafson BD, States DJ, Swaminathan S, Karplus M (1983) Charmm: a program for macromolecular energy, minimization, and dynamics calculations. J Comput Chem 4 (2):187–217. https://doi.org/10.1002/jcc.540040211. https://onlinelibrary.wiley.com/doi/abs/10.1002/jcc.540040211
Carreon AC, Santos WL, Matson JB, So RC (2014) Cationic polythiophenes as responsive dna-binding polymers. Polym. Chem. 5:314–317. https://doi.org/10.1039/C3PY01069D
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Scalmani G, Barone V, Petersson GA, Nakatsuji H, Li X, Caricato M, Marenich AV, Bloino J, Janesko BG, Gomperts R, Mennucci B, Hratchian HP, Ortiz JV, Izmaylov AF, Sonnenberg JL, Williams-Young D, Ding F, Lipparini F, Egidi F, Goings J, Peng B, Petrone A, Henderson T, Ranasinghe D, Zakrzewski VG, Gao J, Rega N, Zheng G, Liang W, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Vreven T, Throssell K, Montgomery JA Jr, Peralta JE, Ogliaro F, Bearpark MJ, Heyd JJ, Brothers EN, Kudin KN, Staroverov VN, Keith TA, Kobayashi R, Normand J, Raghavachari K, Rendell AP, Burant JC, Iyengar SS, Tomasi J, Cossi M, Millam JM, Klene M, Adamo C, Cammi R, Ochterski JW, Martin RL, Morokuma K, Farkas O, Foresman JB, Fox DJ (2016) Gaussian0~9 Revision A.02. Gaussian Inc. Wallingford CT
Galindo-Murillo R, Robertson JC, Zgarbová M, Šponer J, Otyepka M, Jurecka P, Cheatham TE (2016) Assessing the current state of amber force field modifications for dna. J Chem Theory Comput 12(8):4114–4127. https://doi.org/10.1021/acs.jctc.6b00186. PMID: 27300587
Ho HA, Béra-Abérem M, Leclerc M (2005) Optical sensors based on hybrid dna/conjugated polymer complexes. Chemistry 11(6):1718–1724. https://doi.org/10.1002/chem.200400537. https://onlinelibrary.wiley.com/doi/abs/10.1002/chem.200400537
Li C, Numata M, Takeuchi M, Shinkai S (2006) Unexpected chiroptical inversion observed for supramolecular complexes formed between an achiral polythiophene and atp. Chemistry 1(1-2):95–101. https://doi.org/10.1002/asia.200600039. https://onlinelibrary.wiley.com/doi/abs/10.1002/asia.200600039
MacKerell AD, Bashford D, Bellott M, Dunbrack RL, Evanseck JD, Field MJ, Fischer S, Gao J, Guo H, Ha S, Joseph-McCarthy D, Kuchnir L, Kuczera K, Lau F TK, Mattos C, Michnick S, Ngo T, Nguyen DT, Prodhom B, Reiher WE, Roux B, Schlenkrich M, Smith JC, Stote R, Straub J, Watanabe M, Wiórkiewicz-Kuczera J, Yin D, Karplus M (1998) All-atom empirical potential for molecular modeling and dynamics studies of proteins. J Phys Chem B 102(18):3586–3616. https://doi.org/10.1021/jp973084f. PMID: 24889800
Mackerell AD Jr, Feig M, BrooksIII CL (2004) Extending the treatment of backbone energetics in protein force fields: limitations of gas-phase quantum mechanics in reproducing protein conformational distributions in molecular dynamics simulations. J Comput Chem 25(11):1400–1415. https://doi.org/10.1002/jcc.20065. https://onlinelibrary.wiley.com/doi/abs/10.1002/jcc.20065
Mayne CG, Saam J, Schulten K, Tajkhorshid E, Gumbart JC (2013) Rapid parameterization of small molecules using the force field toolkit. J Comput Chem 34(32):2757–2770. https://doi.org/10.1002/jcc.23422. https://onlinelibrary.wiley.com/doi/abs/10.1002/jcc.23422
Miranda OR, You C-C, Phillips R, Kim I-B, Ghosh PS, Bunz U HF, Rotello VM (2007) Array-based sensing of proteins using conjugated polymers. J Am Chem Soc 129(32):9856–9857. https://doi.org/10.1021/ja0737927. PMID: 17658813
Oostenbrink C, Villa A, Mark AE, VanGunsteren WF (2004) A biomolecular force field based on the free enthalpy of hydration and solvation: The gromos force-field parameter sets 53a5 and 53a6. J Comput Chem 25(13):1656–1676. https://doi.org/10.1002/jcc.20090. https://onlinelibrary.wiley.com/doi/abs/10.1002/jcc.20090
Pavlova A, Gumbart JC (2015) Parametrization of macrolide antibiotics using the force field toolkit. J Comput Chem 36(27):2052–2063. https://doi.org/10.1002/jcc.24043. https://onlinelibrary.wiley.com/doi/abs/10.1002/jcc.24043
Pavlova A, Parks JM, Gumbart JC (2018) Development of charmm-compatible force-field parameters for cobalamin and related cofactors from quantum mechanical calculations. J Chem Theory Comput 14(2):784–798. https://doi.org/10.1021/acs.jctc.7b01236. PMID: 29334459
Rajwar D, Ammanath G, Cheema JA, Palaniappan A, Yildiz UH, Liedberg B (2016) Tailoring conformation-induced chromism of polythiophene copolymers for nucleic acid assay at resource limited settings. ACS Appl Mater Interfaces 8(13):8349–8357. https://doi.org/10.1021/acsami.5b12171. PMID: 26956217
Rubio-Magnieto J, Thomas A, Richeter S, Mehdi A, Dubois P, Lazzaroni R, Clément S, Surin M (2013) Chirality in dna–π-conjugated polymer supramolecular structures: insights into the self-assembly. Chem Commun 49:5483–5485. https://doi.org/10.1039/C3CC42108B
Singh UC, Kollman PA (1984) An approach to computing electrostatic charges for molecules. J Comput Chem 5(2):129–145. https://doi.org/10.1002/jcc.540050204. https://onlinelibrary.wiley.com/doi/abs/10.1002/jcc.540050204
Thomas SW, Joly GD, Swager TM (2007) Chemical sensors based on amplifying fluorescent conjugated polymers. Chem Rev 107(4):1339–1386. https://doi.org/10.1021/cr0501339. PMID: 17385926
Tocci G, Joly L, Michaelides A (2014) Friction of water on graphene and hexagonal boron nitride from ab initio methods: Very different slippage despite very similar interface structures. Nano Lett 14 (12):6872–6877. https://doi.org/10.1021/nl502837d. PMID: 25394228
Vanommeslaeghe K, Hatcher E, Acharya C, Kundu S, Zhong S, Shim J, Darian E, Guvench O, Lopes P, Vorobyov I, Mackerell AD Jr (2010) Charmm general force field: a force field for drug-like molecules compatible with the charmm all-atom additive biological force fields. J Comput Chem 31(4): 671–690. https://doi.org/10.1002/jcc.21367. https://onlinelibrary.wiley.com/doi/abs/10.1002/jcc.21367
Wang X, Feng Q, Wang L, Pei M, Zhao J, Zhang G (2014) A novel polythiophene derivative as a sensitive colorimetric and fluorescent sensor for the detection of atp. Des Monomers and Polym 17(1):26–32. https://doi.org/10.1080/15685551.2013.771315
Yao Z, Li C, Shi G (2008) Optically active supramolecular complexes of water-soluble achiral polythiophenes and folic acid: spectroscopic studies and sensing applications. Langmuir 24(22):12829–12835. https://doi.org/10.1021/la802086d
Acknowledgments
The numerical calculations reported in this paper were fully performed at TUBITAK ULAKBIM, High Performance and Grid Computing Center (TRUBA resources). We would like to give thanks to Ümit Hakan Yıldız for inspirations to this study.
Funding
This work was supported by the “Scientific and Technological Research Council of Turkey,” TUBITAK, under the project numbers 119Z100.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kıbrıs, E., Barbak, N.N. & Irmak, N.E. CHARMM force field generation for a cationic thiophene oligomer with ffTK. J Mol Model 27, 34 (2021). https://doi.org/10.1007/s00894-020-04610-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-020-04610-2