Abstract
Wnt and Frizzled (Fzd) family members play crucial roles in the self-renewal of tumor-initiating cells. Until now, only a few studies have addressed the distinct mechanism of Wnt–Fzd interactions. In this study, we suggest a possible interaction mode of Wnt2 with the Fzd7 cysteine-rich domain (CRD)—both of which are up-regulated in some types of cancer. A combination of homology modeling, molecular docking and molecular dynamics (MD) simulations was carried out to study this ligand–receptor complex in great detail. The results demonstrated the unique dynamic behavior of Wnt2 upon binding to Fzd7. Interestingly, the β-strand content of the C-terminal binding site of Wnt2 was obviously reduced when bound to Fzd7 CRD. Moreover, the N-terminal and C-terminal binding sites of Wnt2 appeared to interact with the C-terminal and N-terminal binding sites of Fzd7, respectively. Calculation of the binding energies uncovered the pivotal role of electrostatic and hydrophobic interactions in the binding of Wnt2 to Fzd7 CRD. In conclusion, this study provides valuable insights into the mechanism of the Wnt2-Fzd7 CRD interaction for application in colorectal cancer prevention programs.

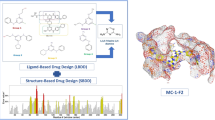
Flowchart representation of different steps used in this study.










Similar content being viewed by others
Abbreviations
- 3D:
-
Three dimensional
- CN:
-
Cluster number
- CRC:
-
Colorectal cancer
- CRD:
-
Cysteine-rich domain
- CTD:
-
C-terminal domain
- DKK:
-
Dikpoff
- DOPE:
-
Discrete optimized potential energy
- FZD:
-
Frizzled
- HM:
-
Homology modeling
- LRP:
-
Lipoprotein receptor related protein
- LE:
-
Lowest energy
- mFZD:
-
mouse frizzled
- MD:
-
Molecular dynamics
- NTD:
-
N-terminal domain
- PDB:
-
Protein Data Bank
- Pdf:
-
Probability density function
- Rg:
-
Radius of gyration
- RMSD:
-
Root mean square deviation
- RMSF:
-
Root mean square fluctuations
- SASA:
-
Solvent accessible surface area
- SPC:
-
Simple point charge
- TCF:
-
Transcription cell factor
- Wnt:
-
Wingless Int
- xWNT:
-
Xenopus Wnt
References
Anastas JN, Moon RT (2013) WNT signalling pathways as therapeutic targets in cancer. Nat Rev Cancer 13(1):11
Polakis P (2012) Wnt signaling in cancer. Cold Spring Harb Perspect Biol 4(5):a008052
Myant K, Sansom OJ (2014) Wnt signaling and colorectal cancer. In: Hoppler S, Moon R (eds) Wnt signaling in development and disease. Wiley, New York, pp 357–367
Dimitriadis A, Vincan E, Mohammed IM, Roczo N, Phillips WA, Baindur-Hudson S (2001) Expression of Wnt genes in human colon cancers. Cancer Lett 166(2):185–191
Akiyama T (2000) Wnt/β-catenin signaling. Cytokine Growth Factor Rev 11(4):273–282
Chu ML-H, Ahn VE, Choi H-J, Daniels DL, Nusse R, Weis WI (2013) Structural studies of Wnts and identification of an LRP6 binding site. Structure 21(7):1235–1242
Janda CY, Waghray D, Levin AM, Thomas C, Garcia KC (2012) Structural basis of Wnt recognition by Frizzled. Science 337:59–64
Holcombe R, Marsh J, Waterman M, Lin F, Milovanovic T, Truong T (2002) Expression of Wnt ligands and frizzled receptors in colonic mucosa and in colon carcinoma. Mol Pathol 55(4):220
Bravo DT, You L, Mazieres J, He B, Xu Z, Jablons DM (2008) Targeting Wnt-2 in mesothelioma and lung cancer. Clin Lung Cancer 9(5):289
Mazieres J, You L, He B, Xu Z, Twogood S, Lee AY, Reguart N, Batra S, Mikami I, Jablons DM (2005) Wnt2 as a new therapeutic target in malignant pleural mesothelioma. Int J Cancer 117(2):326–332
Jung Y-S, Jun S, Lee SH, Sharma A, Park J-I (2015) Wnt2 complements Wnt/β-catenin signaling in colorectal cancer. Oncotarget 6(35):37257
You L, He B, Xu Z, Uematsu K, Mazieres J, Fujii N, Mikami I, Reguart N, McIntosh JK, Kashani-Sabet M (2004) An anti-Wnt-2 monoclonal antibody induces apoptosis in malignant melanoma cells and inhibits tumor growth. Cancer Res 64(15):5385–5389
Shi Y, He B, Kuchenbecker KM, You L, Xu Z, Mikami I, Yagui-Beltran A, Clement G, Lin YC, Okamoto J (2007) Inhibition of Wnt-2 and galectin-3 synergistically destabilizes β-catenin and induces apoptosis in human colorectal cancer cells. Int J Cancer 121(6):1175–1181
Ueno K, Hirata H, Hinoda Y, Dahiya R (2013) Frizzled homolog proteins, microRNAs and Wnt signaling in cancer. Int J Cancer 132(8):1731–1740
Ueno K, Hiura M, Suehiro Y, Hazama S, Hirata H, Oka M, Imai K, Dahiya R, Hinoda Y (2008) Frizzled-7 as a potential therapeutic target in colorectal cancer. Neoplasia 10(7):697–705
Webb B, Sali A (2014) Protein structure modeling with MODELLER. Methods Mol Biol 1137:1–15
Willard L, Ranjan A, Zhang H, Monzavi H, Boyko RF, Sykes BD, Wishart DS (2003) VADAR: a web server for quantitative evaluation of protein structure quality. Nucleic Acids Res 31(13):3316–3319
Söding J, Biegert A, Lupas AN (2005) The HHpred interactive server for protein homology detection and structure prediction. Nucleic Acids Res 33(suppl_2):W244–W248
Larkin MA, Blackshields G, Brown N, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23(21):2947–2948
Berjanskii M, Liang Y, Zhou J, Tang P, Stothard P, Zhou Y, Cruz J, MacDonell C, Lin G, Lu P (2010) PROSESS: a protein structure evaluation suite and server. Nucleic Acids Res 38(suppl_2):W633–W640
Laskowski RA, MacArthur MW, Moss DS, Thornton JM (1993) PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Crystallogr 26(2):283–291
Eisenberg D, Lüthy R, Bowie JU (1997) VERIFY3D: Assessment of protein models with three-dimensional profiles. Methods Enzymol 277:396–404
Colovos C, Yeates TO (1993) Verification of protein structures: patterns of nonbonded atomic interactions. Protein Sci 2(9):1511–1519
Wiederstein M, Sippl MJ (2007) ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res 35(suppl_2):W407–W410
Chen VB, Arendall WB, Headd JJ, Keedy DA, Immormino RM, Kapral GJ, Murray LW, Richardson JS, Richardson DC (2010) MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr D Biol Crystallogr 66(1):12–21
Heo L, Park H, Seok C (2013) GalaxyRefine: protein structure refinement driven by side-chain repacking. Nucleic Acids Res 41(W1):W384–W388
Zhang Y, Skolnick J (2005) TM-align: a protein structure alignment algorithm based on the TM-score. Nucleic Acids Res 33(7):2302–2309
Abraham MJ, Murtola T, Schulz R, Páll S, Smith JC, Hess B, Lindahl E (2015) GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1:19–25
Comeau SR, Gatchell DW, Vajda S, Camacho CJ (2004) ClusPro: an automated docking and discrimination method for the prediction of protein complexes. Bioinformatics 20(1):45–50
Van Zundert G, Rodrigues J, Trellet M, Schmitz C, Kastritis P, Karaca E, Melquiond A, van Dijk M, De Vries S, Bonvin A (2016) The HADDOCK2. 2 web server: user-friendly integrative modeling of biomolecular complexes. J Mol Biol 428(4):720–725
DeLano WL (2002) Pymol: an open-source molecular graphics tool. CCP4 newsletter on protein crystallography 40:82–92
Laskowski RA, Swindells MB (2011) LigPlot+: multiple ligand–protein interaction diagrams for drug discovery. J Chem Inf Model 51:2778–2786
Tina K, Bhadra R, Srinivasan N (2007) PIC: protein interactions calculator. Nucleic Acids Res 35(suppl_2):W473–W476
Brenke R, Hall DR, Chuang G-Y, Comeau SR, Bohnuud T, Beglov D, Schueler-Furman O, Vajda S, Kozakov D (2012) Application of asymmetric statistical potentials to antibody–protein docking. Bioinformatics 28(20):2608–2614
Vangone A, Bonvin AM (2015) Contacts-based prediction of binding affinity in protein–protein complexes. Elife 4:PMC4523921
Jo S, Vargyas M, Vasko-Szedlar J, Roux B, Im W (2008) PBEQ-solver for online visualization of electrostatic potential of biomolecules. Nucleic Acids Res 36(suppl_2):W270–W275
Ichiye T, Karplus M (1991) Collective motions in proteins: a covariance analysis of atomic fluctuations in molecular dynamics and normal mode simulations. Proteins 11(3):205–217
Poorebrahim M, Sadeghi S, Rahimi H, Karimipoor M, Azadmanesh K, Mazlomi MA, Teimoori-Toolabi L (2017) Rational design of DKK3 structure-based small peptides as antagonists of Wnt signaling pathway and in silico evaluation of their efficiency. PLoS One 12(2):e0172217
Rismani E, Rahimi H, Arab SS, Azadmanesh K, Karimipoor M, Teimoori-Toolabi L (2017) Computationally Design of Inhibitory Peptides Against Wnt Signaling Pathway: In Silico Insight on Complex of DKK1 and LRP6. Int J Peptide Res Ther 2017:1–12
Janda CY, Dang LT, You C, Chang J, de Lau W, Zhong ZA, Yan KS, Marecic O, Siepe D, Li X (2017) Surrogate Wnt agonists that phenocopy canonical Wnt and β-catenin signalling. Nature 545(7653):234
Katoh M (2001) Frequent up-regulation of WNT2 in primary gastric cancer and colorectal cancer. Int J Oncol 19(5):1003–1007
Kirikoshi H, Sekihara H, Katoh M (2001) Expression profiles of 10 members of frizzled gene family in human gastric cancer. Int J Oncol 19(4):767–771
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31(2):455–461
Hammad MA, Azam SS (2015) Structural dynamics and inhibitor searching for Wnt-4 protein using comparative computational studies. Drug Des, Dev Therapy 9:2449
Kumar S, Žigman M, Patel TR, Trageser B, Gross JC, Rahm K, Boutros M, Gradl D, Steinbeisser H, Holstein T (2014) Molecular dissection of Wnt3a-Frizzled8 interaction reveals essential and modulatory determinants of Wnt signaling activity. BMC Biol 12(1):44
Agostino M, Pohl SÖ-G, Dharmarajan A (2017) Structure-based prediction of Wnt binding affinities for frizzled-type cysteine-rich domains. J Biol Chem 292(27):11218–11229
Espadaler J, Querol E, Aviles FX, Oliva B (2006) Identification of function-associated loop motifs and application to protein function prediction. Bioinformatics 22(18):2237–2243
Azam SS, Mirza AH (2014) Role of thumb index fold in Wnt-4 protein and its dynamics through a molecular dynamics simulation study. J Mol Liq 198:313–321
Gromiha MM, Selvaraj S (2004) Inter-residue interactions in protein folding and stability. Prog Biophys Mol Biol 86(2):235–277
Kumar S (2014) Molecular dissection of mouse Wnt3a-Frizzled8 interaction reveals essential and modulatory determinants of Wnt signaling activity. PhD Thesis, University of Heidelberg
Voronkov A, Baskin I, Palyulin V, Zefirov N (2007) Molecular modeling of the complex between the XWNT8 protein and the CRD domain of the MFZD8 receptor. Doklad Biochem Biophys 412: 8–11
Ain QU, Seemab U, Rashid S, Nawaz MS, Kamal MA (2013) Prediction of structure of human Wnt-CRD (FZD) complex for computational drug repurposing. PLoS One 8(1):e54630
Fiser A (2010) Template-based protein structure modeling. Methods Mol Biol 673:73–94
Acknowledgments
This study was supported by the Semnan University of Medical Sciences (Grant Number: 810) and the Pasteur Institute of Iran (Grant Number: 94/0110/15866).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
There is no conflict of interest.
Electronic supplementary material
ESM 1
(DOCX 30 kb)
Rights and permissions
About this article
Cite this article
Kalhor, H., Poorebrahim, M., Rahimi, H. et al. Structural and dynamic characterization of human Wnt2-Fzd7 complex using computational approaches. J Mol Model 24, 274 (2018). https://doi.org/10.1007/s00894-018-3788-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-018-3788-3