Abstract
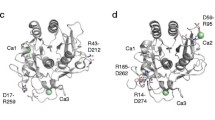
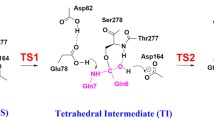
Cysteine protease is ubiquitous in nature. Excess activity of this enzyme causes intercellular proteolysis, muscle tissue degradation, etc. The role of water-mediated interactions in the stabilization of catalytically significant Asp158 and His159 was investigated by performing molecular dynamics simulation studies of 16 three-dimensional structures of plant thiol proteases. In the simulated structures, the hydrophilic W1, W2 and WD1 centers form hydrogen bonds with the OD1 atom of Asp158 and the ND1 atom of His159. In the solvated structures, another water molecule, WE, forms a hydrogen bond with the NE2 atom of His159. In the absence of the water molecule WE, Trp177 (NE1) and Gln19 (NE2) directly interact with the NE2 atom of His159. All these hydrophilic centers (the locations of W1, W2, WD1, and WE) are conserved, and they play a critical role in the stabilization of His–Asp complexes. In the water dynamics of solvated structures, the water molecules W1 and W2 form a water...water hydrogen-bonded network with a few other water molecules. A few dynamical conformations or transition states involving direct (His159 ND1...Asp158 OD1) and water-mediated (His159 ND1...W2...Asp158 OD1) hydrogen-bonded complexes are envisaged from these studies.




Similar content being viewed by others
References
Otto HH, Schirmeister T (1997) Chem Rev 97:133–171
Komatsu K, Tsukuda K, Hosoya J, Satoh S (1986) Exp Neurol 93:642–646
Sloane BF, Moin K, Krepela E, Rozhin J (1990) Cancer Metastasis Rev 9:333–352
Lecaille F, Kaleta J, Bromme D (2002) Chem Rev 102:4459–4488
Huet G, Flip RM, Richet C, Thiebet C, Demeyer D, Balduyck M, Duquesnoy B, Degand P (1992) Clin Chem 38:1694–1697
Polgár L (1974) FEBS Lett 47:15–18
Vernet T, Tessier DC, Chatellier J, Plouffe C, Lee TS, Thomas DY, Storer AC, Menard R (1995) J Biol Chem 270:16645–16652
Harrison MJ, Burton NA, Hillier IH (1997) J Am Chem Soc 119:12285–12291
O’Farrell PA, Joshua-Tor L (2007) Biochem J 401:421–428
Mladenovic M, Fink RF, Thiel W, Schirmeister T, Engels B (2008) J Am Chem Soc 130:8696–8705
Nandi TK, Bairagya HR, Mukhopadhyay BP, Sekar K, Sukul D, Bera AK (2009) J Biosci 34:27–34
Menard R, Khouri HE, Plouffe C, Laflamme P, Durpras R, Vernet T, Tessier DC, Thomas DY, Storer AC (1991) Biochemistry 30:5531–5538
Wang J, Xiang YF, Lim C (1994) Protein Eng Des Sel 7:75–82
Dijkman JP, Van Duijnen PTh (1991) Int J Quant Chem 40:49–59
Welsh WJ, Lin Y (1997) J Mol Struct THEOCHEM 401:315–326
Day RM, Thalhauser CJ, Sudmeier JL, Vincent MP, Torchilin EV, Sanford DG, Bachovchin CW, Bachovchin WW (2003) Protein Sci 12:794–810
Berman HM, Westbrook J, Feng Z, Gilli G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) Nucleic Acids Res 28:235–242
Pickersgill RW, Harris GW, Garman E (1992) Acta Crystallogr Sect B 48:59–67
Kamphuis IG, Kalk KH, Swarte MB, Drenth J (1984) J Mol Biol 179:233–256
Yamamoto D, Matsumoto K, Ohishi H, Ishida T, Inoue M, Kitamura K, Mizuno H (1991) J Biol Chem 266:14771–14777
Kim MJ, Yamamoto D, Matsumoto K, Inoue M, Ishida T, Mizuno H, Sumiya S, Kitamura K (1992) Biochem J 287:797–803
Janowski R, Kozak M, Jankowska E, Grzonka Z, Jaskolski M (2004) J Pept Res 64:141–150
LaLonde JM, Zhao B, Smith WW, Janson CA, DesJarlais RL, Tomaszek TA, Carr TJ, Thompson SK, Oh HJ, Yamashita DS, Veber DF, Abdel-Meguid SS (1998) J Med Chem 41:4567–4576
Varughese KI, Su Y, Cromwell D, Hasnain S, Xuong NH (1992) Biochemistry 31:5172–5176
Baker EN, Dodson EJ (1980) Acta Crystallogr Sect A 36:559–572
Katerelos NA, Taylor MA, Scott M, Goodenough PW, Pickersgill RW (1996) FEBS Lett 392:35–39
Maes D, Bouckaert J, Poortmans F, Wyns L, Looze Y (1996) Biochemistry 35:16292–16298
O’Hara BP, Hemmings AM, Buttle DJ, Pearl LH (1995) Biochemistry 34:13190–13195
Biswas S, Chakrabarti C, Kundu S, Jagannadham MV, Dattagupta JK (2003) Proteins 51:489–497
Ghosh R, Dattagupta JK, Biswas S (2007) Biochem Biophys Res Commun 362:965–970
Choi KH, Laursen RA, Allen KN (1999) Biochemistry 38:11624–11633
Guex N, Diem A, Peitsch MC, Schwede T (2001) The deep view—the Swiss-PdbView program, an environment for comparative protein modeling. GlaxoSmithKline R&D, London
Humphrey W, Dalke A, Schulten K (1996) J Mol Graph 14:33–38
Brooks BR, Bruccoleri RE, Olafson BD, States DJ, Swaminathan S, Karplus M (1983) J Comput Chem 4:187–217
Nelson M, Humphrey W, Gursoy A, Dalke A, Kale L, Skeel RD, Schulten K (1996) Int J Supercomput Appl High Perform Comput 10:251–268
Phillips JC, Braun R, Wang W, Gumbart J, Emad Tajkhorshid E, Villa E, Chipot C, Skeel RD, Kale L, Schulten K (2005) J Comput Chem 26:1781–1802
Fleming PJ, Nicholas C, Fitzkee MM, Rajgopal S, George DR (2005) Protein Sci 14:111–118
Bairagya HR, Mukhopadhyay BP, Sekar K (2009) J Bio Struct Dyn 26:497–507
Bairagya HR, Mukhopadhyay BP, Sekar K (2009) J Bio Struct Dyn 27:149–158
Schymkowitz J, Borg J, Stricher F, Nys R, Rousseau F, Serrano L (2005) Nucleic Acids Res 33:382–388
Mustata G, Briggs JM (2004) Protein Eng Des Sel 17:223–234
Gul S, Hussain S, Thomas MP, Resmini M, Verma CS, Thomas EW, Brocklehurst K (2008) Biochemistry 47:2025–2035
Stollar EJ, Gelpi JL, Velankar S, Golovin A, Orozco M, Luisi BF (2004) Proteins 57:1–8
Sulpizi M, Rothlisberger U, Carloni P (2003) Biophys J 84:2207–2215
Acknowledgments
We thank and acknowledge the National Institute of Technology Durgapur for providing a research facility in the Department of Chemistry. We also thank and acknowledge Dr. K Sekar, Indian Institute of Science, Bangalore, India, for critically reading this manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Nandi, T.K., Bairagya, H.R., Mukhopadhyay, B.P. et al. Conserved water-mediated H-bonding dynamics of catalytic His159 and Asp158: insight into a possible acid–base coupled mechanism in plant thiol protease. J Mol Model 18, 2633–2644 (2012). https://doi.org/10.1007/s00894-011-1277-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-011-1277-z