Abstract
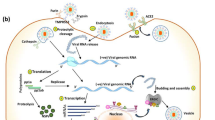
The entry of human immunodeficiency virus (HIV) into cells depends on a sequential interaction of the gp120 envelope glycoprotein with the cellular receptors CD4 and members of the chemokine receptor family. The CC chemokine receptor CCR5 is such a receptor for several chemokines and a major coreceptor for the entry of R5 HIV type-1 (HIV-1) into cells. Although many studies focus on the interaction of CCR5 with HIV-1, the corresponding interaction sites in CCR5 and gp120 have not been matched. Here we used an approach combining protein structure modeling, docking and molecular dynamics simulation to build a series of structural models of the CCR5 in complexes with gp120 and CD4. Interactions such as hydrogen bonds, salt bridges and van der Waals contacts between CCR5 and gp120 were investigated. Three snapshots of CCR5–gp120–CD4 models revealed that the initial interactions of CCR5 with gp120 are involved in the negatively charged N-terminus (Nt) region of CCR5 and positively charged bridging sheet region of gp120. Further interactions occurred between extracellular loop2 (ECL2) of CCR5 and the base of V3 loop regions of gp120. These interactions may induce the conformational changes in gp120 and lead to the final entry of HIV into the cell. These results not only strongly support the two-step gp120–CCR5 binding mechanism, but also rationalize extensive biological data about the role of CCR5 in HIV-1 gp120 binding and entry, and may guide efforts to design novel inhibitors.
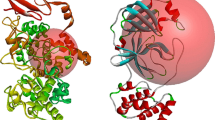
Figure Three structural models of CCR5 in complex with gp120–CD4. These represent snapshots of different intermediates of the complex that characterize the dynamic interaction process of CCR5 with gp120. The ribbon diagram shows gp120 in red, the two N-terminal domains of CD4 in yellow, and CCR5 in blue. The Nt, ECL1, 2, and 3 in CCR5, as well as the bridging sheet and V3 loop base of gp120 are labeled. Model I shows that Nt begins to interact with the bridging sheet, the ECLs of CCR5 do not interact with gp120 and there is a big gap between them. It is a snapshot of the initial recognition of CCR5 with gp120. Model III shows that ECL2 begins to interact with the V3 loop base in addition to the interaction between Nt and the bridging sheet. It is a snapshot of the transition state of the binding process. Model II shows the full interaction between CCR5 and gp120. More residues within Nt interact with the bridging sheet, ECL2 and ECL3 interact with the V3 base loop and bridging sheet, respectively. It is a snapshot of a "fusion-active" state prior to virus entry. The interaction intensities between CCR5 and gp120 are model I<model III<model II




Similar content being viewed by others
Notes
Human immunodeficiency virus
N terminal/N terminus
Extracellular loop
Regulated on activation normal T cell expressed and secreted
Macrophage inflammatory protein
References
Dalgleish AG, Beverley PC, Clapham PR, Crawford DH, Greaves MF, Weiss RA (1984) Nature 312:763–766
Klatzmann D, Champagne E, Chamaret S, Gruest J, Guetard D, Hercend T, Gluckman JC, Montagnier L (1984) Nature 312:767–768
Feng Y, Broder CC, Kennedy PE, Berger EA (1996) Science 272:872–877
Dragic T, Litwin V, Allaway GP, Martin SR, Huang Y, Nagashima KA, Cayanan C, Maddon PJ, Koup RA, Moore JP, Paxton WA (1996) Nature 381:667–673
Deng H, Liu R, Ellmeier W, Choe S, Unutmaz D, Burkhart M, Marzio PD, Marmon S, Sutton RE, Hill CM, Davis CB, Pepier SC, Schall TJ, Littman DR, Landau NR (1996) Nature 381:661–666
Alkhatib G, Combadiere C, Broder CC, Feng Y, Kennedy PE, Murphy PM (1996) Science 272:1955–1958
Strader CD, Fong TM, Tota MR, Underwood D (1994) Annu Rev Biochem 63:101–132
Dean M, Carrington M, Winkler C, Huttley GA, Smith MW, Allikmets R, Goedert JJ, Buchbinder SP, Vittinghoff E, Gomperts E, Donfield S, Vlahov D, Kaslow R, Saah A, Rinaldo C, Detels R, O'Brien SJ (1996) Science 273:1856–1862
Huang Y, Paxton WA, Wolinsky SM, Neumann AU, Zhang L, He T, Kang S, Ceradini D, Jin Z, Yazdanbakhsh K, Kunstman K, Erickson D, Dragon E, Landau NR, Phair J, Ho DD, Koup RA (1996) Nat Med 2:1240–1243
Samson M, Libert F, Doranz BJ, Rucker J, Liesnard C, Farber CM, Saragosti S, Lapoumeroulie C, Cognaux J, Forceille C, Muyldermans G, Verhofstede C, Burtonboy G, Georges M, Imai T, Rana S, Yi Y, Smyth RJ, Collman RG, Doms RW, Vassart G, Parmentier M (1996) Nature 382:722–725
Wu L, Gerard NP, Wyatt R, Choe H, Parolin C, Ruffing A, Borsetti A, Cardoso AA, Desjardin E, Newman W, Gerard C, Sodorski J (1996) Nature 384:179–183
Trkola A, Dragic T, Arthos J, Binley JM, Olson WC, Allaway GP, Cheng-Mayer C, Robinson J, Maddon PJ, Moore JP (1996) Nature 384:184–186
Kwong PD, Wyatt R, Robinson J, Sweet RW, Sodroski J, Hendrickson WA (1998) Nature 393:648–659
Wyatt R, Kwong PD, Desjardins E, Sweet RW, Robinson J, Hendrickson WA, Sodroski JG (1998) Nature 393:705–711
Rizzuto CD, Wyatt R, Hernandez-Ramos N, Sun Y, Kwong PD, Hendrickson WA, Sodroski JA (1998) Science 280:1949–1953
Walker JE, Saraste M (1996) Curr Opin Struct Biol 6:457–459
Gosling J, Monteclaro FS, Atchison RE, Arai H, Tsou CL, Goldsmith MA, Charo IF (1997) Proc Natl Acad Sci USA 94:5061–5066
Atchison RE, Gosling J, Monteclaro FS, Franci C, Digilio L, Charo IF, Goldsmith MA (1996) Science 274:1924–1926
Picard L, Simmons G, Power CA, Meyer A, Weiss RA, Clapham PR (1997) J Virol 71:5003–5011
Alkhatib G, Ahuja SS, Light D, Mummidi S, Berger EA, Ahuja SK (1997) J Biol Chem 272:19771–19776
Samson M, LaRosa G, Libert F, Paindavoine P, Detheux M, Vassart G (1997) J Biol Chem 272:24934–24941
Dragic T, Trkola A, Lin SW, Nagashima KA, Kajumo F, Zhao L, Olson WC, Wu L, Mackay CR, Allaway GP, Sakmar TP, Moore JP, Maddon PJ (1998) J Virol 72:279–285
Rabut GE, Konner JA, Kajumo F, Moore JP, Dragic T (1998) J Virol 72:3464–3468
Farzan M, Choe H, Vaca L, Martin K, Sun Y, Desjardins E, Ruffing N, Wu L, Wyatt R, Gerard N, Gerard C, Sodroski J (1998) J Virol 72:1160–1164
Navenot JM, Wang ZX, Trent JO, Murray JL, Hu QX, DeLeeuw L, Moore PS, Chang Y, Peiper SC (2001) J Mol Biol 313:1181–1193
Lee B, Sharron M, Blanpain C, Doranz BJ, Vakili J, Setoh P, Berg E, Liu G, Guy HR, Durell SR, Parmentier M, Chang CN, Price K, Tsang M, Doms RW (1999) J Biol Chem 274:9617–9626
Genoud S, Kajumo F, Guo Y, Thompson D, Dragic T (1999) J Virol 73:1645–1648
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) Nucleic Acids Research 28:235–242
Guex N, Peitsch MC (1997) Electrophoresis 18:2714–2723
Binley JM, Wyatt R, Desjardins E, Kwong PD, Hendrickson W, Moore JP, Sodroski J (1998) AIDS Res Hum Retrovir 14:191–198
Blanpain B, Doranz BJ, Vakili J, Rucker J, Govaerts C, Baikc SSW, Lorthioir O, Migeotte I, Libert F, Baleux F, Vassart G, Doms RW, Parmentier M (1999) J Biol Chem 274:34719–34727
Doranz BJ, Lu ZH, Rucker J, Zhang TY, Sharron M, Cen YH, Wang ZX, Guo HH, Du JG, Accavitti MA, Doms RW, Peiper SC (1997) J Virol 71:6305–6314
Rucker J, Samson M, Doranz BJ, Libert F, Berson JF, Yi Y, Smyth RJ, Collman RG, Broder CC, Vassart G, Doms RW, Parmentier M (1996) Cell 87:437–446
Kraulis PJ (1991) J Appl Crystallogr 24:946–950
Acknowledgements
This work was supported in part by the National Natural Science Grant (China) (No. 19947006) and partially by the National Key Foundation Research Grant (No. 2002AA231031) in China (863)
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Liu, S., Fan, S. & Sun, Z. Structural and functional characterization of the human CCR5 receptor in complex with HIV gp120 envelope glycoprotein and CD4 receptor by molecular modeling studies. J Mol Model 9, 329–336 (2003). https://doi.org/10.1007/s00894-003-0154-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-003-0154-9