Abstract
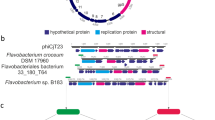
Virion DNA of bacteriophage 11b (Φ11b), which infects a psychrophilic Flavobacterium isolate from Arctic sea-ice, was determined to consist of 36,012 bp. With 30.6% its GC content corresponds to that of host-genus species and is the lowest of all phages of Gram-negative bacteria sequenced so far. Similarities of several of 65 predicted ORFs, genome organization and phylogeny suggest an affiliation to ‘mesophilic’ nonmarine siphoviruses, e.g. to bacteriophages SPP1 and HK97. Early genes presumably encode an essential recombination factor (ERF), a single strand binding (SSB) protein, an endonuclease, and a DNA methylase. The late gene segment is likely to contain a terminase, portal, minor head, protease and a major capsid gene. Five ORFs exhibited similarities to Bacteroidetes species and seem to reflect the host specificity of the phage. Among PAGE-separated virion proteins that were identified by MALDI-ToF mass spectrometry are the portal, the major capsid, and a putative conserved tail protein. The Φ11b genome is the first to be described of a cultivated virus infecting a psychrophilic host as well as a Bacteroidetes bacterium.




Similar content being viewed by others
References
Ackermann H-W, Abedon ST (2001) Bacteriophage names. The Bacteriophage Ecology Group. (Online.) http://www.phage.org/names.htm
Alonso JC, Luder G, Stiege AC, Chai S, Weise F, Trautner TA (1997) The complete nucleotide sequence and functional organization of Bacillus subtilis bacteriophage SPP1. Gene 204:201–212
Andrade MA, Ponting CP, Gibson TJ, Bork P (2000) Homology-based method for identification of protein repeats using statistical significance estimates. J Mol Biol 298:521–537
Arai M, Mitsuke H, Ikeda M, Xia J-X, Kikuchi T, Satake M, Shimizu T (2004) ConPred II: a consensus prediction method for obtaining transmembrane topology models with high reliability. Nucleic Acids Res 32:390–393
Badger JH, Olsen GJ (1999) CRITICA: coding region identification tool invoking comparative analysis. Mol Biol Evol 16:512–524
Bendtsen JD, Nielsen H, von Heijne G, Brunak S (2004) Improved prediction of signal peptides: SignalP 3.0. J Mol Biol 340:783–795
Berger B, Wilson DB, Wolf E, Tonchev T, Milla M, Kim PS (1995) Predicting coiled coils by use of pairwise residue correlations. Proc Natl Acad Sci USA 92:8259–8263
Bernardet J-F, Segers P, Vancanneyt M, Berthe F, Kersters K, Vandamme P (1996) Cutting a Gordian knot: emended classification and description of the genus Flavobacterium, emended description of the family Flavobacteriaceae, and proposal of Flavobacterium hydatis nom. nov. (basonym, Cytophaga aquatilis Strohl and Tait 1978). Int J Syst Bacteriol 46:128–148
Blum H, Beier H, Gross HJ (1987) Improved silver staining of plant proteins, RNA and DNA in polyacrylamide gels. Electrophoresis 8:93–99
Borriss M, Helmke E, Hanschke R, Schweder T (2003) Isolation and characterisation of marine psychrophillic phage-host systems from Arctic sea ice. Extremophiles 7:377–384
Bujnicki JM (2002) Sequence permutations in the molecular evolution of DNA methyltransferases. BMC Evol Biol 2:3
Büttner K, Bernhardt J, Scharf C, Schmid R, Mäder U, Eymann C, Antelmann H, Völker A, Völker U, Hecker M (2001) A comprehensive two-dimensional map of cytosolic proteins of Bacillus subtilis. Electrophoresis 22:2908–2935
Bradford D, Hugenholtz P, Seviour EM, Cunningham MA, Stratton H, Seviour RJ, Blackall LL (1996) 16S rRNA analysis of isolates obtained from gram-negative filamentous bacteria micromanipulated from activated sludge. Syst Appl Microbiol 19:334–343
Brinkmeyer R, Knittel K, Jurgens J, Weyland H, Amann R, Helmke E (2003) Diversity and structure of bacterial communities in Arctic versus Antarctic pack ice. Appl Environ Microbiol 69: 6610–6599
Camacho AG, A. Gual A, Lurz R, Tavares P, Alonso JC (2003) Bacillus subtilis bacteriophage SPP1 DNA packaging motor requires terminase and portal proteins. J Biol Chem 278:23251–23239
Casjens SR, Gilcrease EB, Winn-Stapley DA, Schicklmaier P, Schmieger H, Pedulla ML, Ford ME, Houtz JM, Hatfull GF, Hendrix RW (2005) The generalized transducing Salmonella bacteriophage ES18: complete genome sequence and DNA packaging strategy. J Bacteriol 187:1091–1104
Chai S, Bravo A, Luder G, Nedlin A, Trautner TA, Alonso JC (1992) Molecular analysis of the Bacillus subtilis bacteriophage SPP1 region encompassing genes 1 to 6. The products of gene 1 and gene 2 are required for pac cleavage. J Mol Biol 224:87–102
Clarke GD, Beiko RG, Ragan MA, Charlebois RL (2002) Inferring genome trees by using a filter to eliminate phylogenetically discordant sequences and a distance matrix based on mean normalized BLASTP scores. J Bacteriol 184:2072–2080
Combet C, Blanchet C, Geourjon C, Deléage G (2000) NPS@: Network Protein Sequence Analysis TIBS 25:147–150
Crutz-Le Coq AM, Cesselin B, Commissaire J, Anba J (2002) Sequence analysis of the lactococcal bacteriophage bIL170: insights into structural proteins and HNH endonucleases in dairy phages. Microbiology 148:985–1001
Cserzo M, Wallin E, Simon I, von Heijne G, Elofsson A (1997) Prediction of transmembrane alpha-helices in prokaryotic membrane proteins: the Dense Alignment Surface method. Prot Eng 10:673–676
Delcher AL, Harmon D, Kasif S, White O, Salzberg SL (1999) Improved microbial gene identification with GLIMMER. Nucleic Acids Res 27:4636–4641
Desiere F, Lucchini S, Canchaya C, Ventura M, Brussow H (2002) Comparative genomics of phages and prophages in lactic acid bacteria. Antonie Van Leeuwenhoek 82:73–91
Duda RL, Martincic K, Hendrix RW (1995) Genetic basis of bacteriophage HK97 prohead assembly. J Mol Biol 247:636–647
Engelke DR, Hoener PA, Collins FS (1988) Direct sequencing of enzymatically amplified human genomic DNA. Proc Natl Acad Sci USA 85:544–548
Gardy JL, Laird MR, Chen F, Rey S, Walsh CJ, Ester M, Brinkman FSL (2005) PSORTb v.2.0: expanded prediction of bacterial protein subcellular localization and insights gained from comparative proteome analysis. Bioinformatics 21:617–623
Gasteiger E, Hoogland C, Gattiker A, Duvaud S, Wilkins MR, Appel RD, Bairoch A (2005) Protein identification and analysis tools on the ExPASy server. In Walker JM (eds) The proteomics protocols handbook. Humana Press, pp 571–607
Gruber M, Soding J, Lupas AN (2005) REPPER–repeats and their periodicities in fibrous proteins. Nucleic Acids Res 33:239–243
Gual A, Camacho AG Alonso JC (2000) Functional analysis of the terminase large subunit, G2P, of Bacillus subtilis bacteriophage SPP1. J Biol Chem 275:35311–35299
Heger A, Holm L (2000) Rapid automatic detection and alignment of repeats in protein sequences. Proteins 41:224–237
Hirokawa T, Seah BC, Mitaku S (1998) SOSUI: classification and secondary structure prediction system for membrane proteins. Bioinformatics 14:378–379
Jiang SC, Kellogg CA, Paul JH (1998) Characterization of marine temperate phage-host systems isolated from Mamala Bay, Oahu, Hawaii. Appl Environ Microbiol 64:535–542
Jones DT (1999) Protein secondary structure prediction based on position-specific scoring matrices. J Mol Biol 292:195–202
Juncker AS, Willenbrock H, von Heijne G, Nielsen H, Brunak S, Krogh A (2003) Prediction of lipoprotein signal peptides in Gram-negative bacteria. Protein Sci 12:1652–1662
Kirchman DL (2002) The ecology of cytophaga–flavobacteria in aquatic environments. FEMS Microbiol Ecol 39:91–100
Kisand V, Cuadros R, Wikner J (2002) Phylogeny of culturable estuarine bacteria catabolizing riverine organic matter in the northern Baltic Sea. Appl Environ Microbiol 68:379–388
Kobayashi I (2001) Behavior of restriction-modification systems as selfish mobile elements and their impact on genome evolution. Nucleic Acids Res 29:3742–3756
Krogh A, Larsson B, von Heijne G, Sonnhammer ELL (2001) Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol 305:567–580
Kwan T, Liu J, Dubow M, Gros P Pelletier J (2005) The complete genomes and proteomes of 27 Staphylococcus aureus bacteriophages. Proc Natl Acad Sci USA 102:5174–5179
Leader DP (2004) BugView: a browser for comparing genomes. Bioinformatics 20:129–130
Lowe TM, Eddy SR (1997) tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 25:955–964
Lupas A, Van Dyke M, Stock J (1991) Predicting coiled coils from protein sequences. Science 252:1162–1164
Magrini V, Salmi D, Thomas D, Herbert S, Hartzell P, Youderian P (1997) Temperate Myxococcus xanthus phage Mx8 encodes a DNA adenine methylase, Mox. J Bacteriol 179:4254–4263
McGinnis S, Madden TL (2004) BLAST: at the core of a powerful and diverse set of sequence analysis tools. Nucleic Acids Res 32:20–25
Meyer F, Goesmann A, McHardy AC, Bartels D, Bekel T, Clausen J, Kalinowski J, Linke B, Rupp O, Giegerich R, Puhler A (2003) GenDB-an open source genome annotation system for prokaryote genomes. Nucleic Acids Res 31:2187–2195
Proux C, van Sinderen D, Suarez J, Garcia P, Ladero V, Fitzgerald GF, Desiere F, Brussow H (2002) The dilemma of phage taxonomy illustrated by comparative genomics of Sfi21-like Siphoviridae in lactic acid bacteria. J Bacteriol 184:6026–6036
Rao DN, Eberle H, Bickle TA (1989) Characterization and mutations of the bacteriophage P1 mod gene encoding the recognition subunit of the EcoP1 restriction and modification system. J Bacteriol 171:2347–2352
Roberts MD, Martin NL, Kropinski AM (2004) The genome and proteome of coliphage T1. Virology 318:245–266
Smith MC, Burns RN, Wilson SE, Gregory MA (1999) The complete genome sequence of the Streptomyces temperate phage phiC31: evolutionary relationships to other viruses. Nucleic Acids Res 27:2145–2155
Teeling H., Lombardot T, Bauer M, Ludwig W, Glöckner FO (2004) Evaluation of the phylogenetic position of the planctomycete ‘Rhodopirellula baltica’ SH 1 by means of concatenated ribosomal protein sequences, DNA-directed RNA polymerase subunit sequences and whole genome trees. Int J Syst Evol Microbiol 54:791–801
Tusnady GE, Simon I (2001) The HMMTOP transmembrane topology prediction server. Bioinformatics. 17:849–850
Van Sinderen D, Karsens H, Kok J, Terpstra P, Ruiters MH, Venema G, Nauta A (1996) Sequence analysis and molecular characterization of the temperate lactococcal bacteriophage r1t. Mol Microbiol 19:1343–1355
Wikoff WR, Liljas L, Duda RL, Tsuruta H, Hendrix RW, Johnson JE (2000) Topologically linked protein rings in the bacteriophage HK97 capsid. Science 289:2129–2133
Wolf E, Kim PS, Berger B (1997) MultiCoil: a program for predicting two- and three-stranded coiled coils. Protein Sci 6:1179–1189
Young R, Wang I-N, Roof WD (2000) Phages will out: strategies of host cell lysis. Trends Microbiol 8:120–128
Zimmer M, Sattelberger E, Inman RB, Calendar R, Loessner MJ (2003) Genome and proteome of Listeria monocytogenes phage PSA: an unusual case for programmed +1 translational frameshifting in structural protein synthesis. Mol Microbiol 50:303–317
Acknowledgments
We thank Hansjörg Lehnherr for critical reading of the manuscript and Dr. Steffen Krüger for helpful comments during the sequencing and sequence assembly procedure. We are grateful for the support of Michael Hecker in the analysis of the phage proteome.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by K. Horikoshi
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Borriss, M., Lombardot, T., Glöckner, F.O. et al. Genome and proteome characterization of the psychrophilic Flavobacterium bacteriophage 11b. Extremophiles 11, 95–104 (2007). https://doi.org/10.1007/s00792-006-0014-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-006-0014-5