Abstract
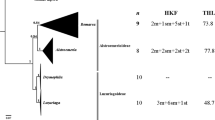
Gnamptogenys includes 138 described species that are widely distributed, with high diversity, in the Neotropics. Some Neotropical species have taxonomic issues, as is the case with Gnamptogenys striatula, for which morphological variations have been observed between different populations. For the ant species with taxonomic issues, classical and molecular cytogenetic studies have assisted in the resolution of these issues. Cytogenetic studies of Gnamptogenys are scarce and have only been reported for 14 taxa. These reports have rarely presented chromosomal morphology. Considering the importance of the taxonomic revision of some species, such as G. striatula, the present study cytogenetically characterized four species of Gnamptogenys: G. striatula, G. moelleri, G. regularis, and G. triangularis, discussing their phylogenetic and biogeographic characteristics. The number of chromosomes ranged from 2n = 26 to 2n = 44, with distinct karyotypes at both species and population levels. All four species presented a pair of 18S rDNA gene markers that coincided with GC-rich regions. In the case of G. striatula from the Atlantic rainforest, a chromosomal polymorphism was observed, with chromosomal translocations being the likely origin of this polymorphism. Two populations of G. striatula showed karyotype differences, thus corroborating previous morphological data indicating the existence of a species complex in this taxon. In addition, G. regularis showed a polymorphism involving a chromosome pair bearing ribosomal genes, possibly caused by unequal crossing-over. Although G. moelleri has a well-defined taxonomy, a population from the eastern Amazon rainforest presented a divergent karyotype from the Atlantic rainforest populations, suggesting the existence of a cryptic species in this taxon.





Similar content being viewed by others
References
Aguiar HJAC, Barros LAC, Alves DR, Mariano CSF, Delabie JHC, Pompolo SG (2017) Cytogenetic studies on populations of Camponotus rufipes (Fabricius, 1775) and Camponotus renggeri Emery, 1894 (Formicidae: Formicinae). PLoS One 12(5):e0177702. https://doi.org/10.1371/journal.pone.0177702
Alves-Silva AP, Barros LAC, Chaul JCM, Pompolo SG (2014) The First cytogenetic data on Strumigenys louisianae Roger, 1863 (Formicidae: Myrmicinae: Dacetini): The lowest chromosome number in the Hymenoptera of the Neotropical region. PLoS One 9(11):e111706. https://doi.org/10.1371/journal.pone.0111706
Barros LAC, Aguiar HJAC, Mariano CSF, Delabie JHC, Pompolo SG (2010) Cytogenetic characterization of the lower-Attine Mycocepurus goeldii (Formicidae: Myrmicinae: Attini). Sociobiology 56:57–66
Barros LAC, Aguiar HJAC, Andrade-Souza V, Mariano CSF, Delabie JHC, Pompolo SG (2012) Occurrence of pre-nucleolar bodies and 45S rDNA location on the chromosomes of the ant Mycocepurus goeldii (Forel) (Formicidae, Myrmicinae, Attini). Hereditas 149:50–54. https://doi.org/10.1111/j.1601-5223.2011.02237.x
Barros LAC, Teixeira GA, Aguiar HJAC, Mariano CSF, Delabie JHC, Pompolo SG (2014a) Banding patterns of three leafcutter ant species of the genus Atta (Formicidae: Myrmicinae) and chromosomal inferences. Fla Entomol 97:1694–1701. https://doi.org/10.1653/024.097.0444
Barros LAC, Aguiar HJAC, Mariano CSF, Delabie JHC, Pompolo SG (2014b) Cytogenetic characterization of the ant Trachymyrmex fuscus Emery, 1934 (Formicidae: Myrmicinae: Attini) with the description of a chromosomal polymorphism. Ann Soc Entomol Fr 49(4):367–373. https://doi.org/10.1080/00379271.2013.856201
Barros LAC, Aguiar HJAC, Mariano CS, Andrade-Souza V, Costa MA, Delabie JH, Pompolo SG (2016) Cytogenetic data on six leafcutter ants of the genus Acromyrmex Mayr, 1865 (Hymenoptera, Formicidae, Myrmicinae): insights into chromosome evolution and taxonomic implications. Comp Cytogenet 10(2):229–243. https://doi.org/10.3897/CompCytogen.v10i2.7612
Blatrix R, Jaisson P (2000) Optional gamergates in the queenright ponerine ant Gnamptogenys striatula Mayr. Insect Soc 47:193–197. https://doi.org/10.1007/PL00001701
Borges DS, Mariano CSF, Delabie JHC, Pompolo SG (2004) Cytogenetics studies in Neotropical ants of the genus Gnamptogenys Roger (Hymenoptera, Formicidae, Ectatominae). Rev Bras Entomol 48(4):481–484. https://doi.org/10.1590/S0085-56262004000400009
Camacho GP (2013) Estudo taxonômico do grupo striatula de Gnamptogenys Roger (Hymenoptera, Formicidae, Ectatomminae) para o Brasil. Dissertation, Universidade Federal de Viçosa, Viçosa
Camacho JPM, Navas-Castillo J, Cabrero J (1986) Extra nucleolar activity associated with presence of a supernumerary chromosome segment in the grasshopper Oedipoda fuscocincta. Heredity 56:237–241
Camacho GP, Feitosa RM (2015) Estado da arte sobre a taxonomia e filogenia de Ectatomminae. In: Delabie JHC, Feitosa RM, Serrao JE, Mariano CSF, Majer JD (eds) As formigas poneromorfas do Brasil, 1st edn. Ilhéus, Brasil, pp 22–32
Correia JPSO, Mariano CSF, Delabie JHCD, Lacau S, Costa MA (2016) Cytogenetic analysis of Pseudoponera stigma and Pseudoponera gilberti (Hymenoptera: Formicidae: Ponerinae): a taxonomic approach. Fla Entomol 99(4):718–721. https://doi.org/10.1653/024.099.0422
Cristiano MP, Cardoso DC, Fernandes-Salomão TM (2013) Cytogenetic and molecular analyses reveal a divergence between Acromyrmex striatus (Roger, 1863) and other congeneric species: taxonomic implications. PLoS One 8(3):e59784. https://doi.org/10.1371/journal.pone.0059784
Feitosa RM (2015) Lista de formigas poneromorfas do Brasil. In: Delabie JHC, Feitosa RM, Serrao JE, Mariano CSF, Majer JD (eds) As formigas poneromorfas do Brasil, 1st edn. Ilhéus, Brasil, pp 94–101
Giraud T, Blatrix R, Poteaux R, Solignac M, Jaisson P (2000) Population structure and mating biology of the polygynous ponerine ant Gnamptogenys striatula in Brazil. Mol Ecol 9:1835–1184. https://doi.org/10.1046/j.1365-294x.2000.01085.x
Goñi B, Imai HT, Kubota M, Kondo M, Yong HS, Tso YP (1982) Chromosome observations of tropical ants in Western Malaysia and Singapore. Annu Rep Natl Inst Genet (Japan) 32:71–73
Imai HT, Crozier RH, Taylor RW (1977) Karyotype evolution in Australian ants. Chromosoma 59:341–393. https://doi.org/10.1007/BF00327974
Imai HT, Brown WL, Kubota M, Yong HS, Tho YP (1983) Chromosome observations on Tropical ants from Western Malaysia. II Annu Rep Natl Inst Genet (Japan) 34:66–69
Imai H, Taylor RW, Crosland MW, Crozier RH (1988) Modes of spontaneous chromossomal mutation and karyotype evolution in ants with reference to the minimum interaction hypothesis. Jpn J Genet 63:159–185. https://doi.org/10.1266/jjg.63.159
Imai HT (1991) Mutability of constitutive heterochromatin (C-bands) during eukaryotic chromosomal evolution ad their cytological meaning. Jpn J Genet 66:635–661. https://doi.org/10.1266/jjg.66.635
Imai HT, Taylor RW, Crozier RH (1994) Experimental bases for the minimum interaction theory. Chromosome evolution in ants of the Myrmecia pilosula species complex (Hymenoptera: Formicidae: Myrmeciinae). Jpn J Genet 69:137–182. https://doi.org/10.1266/jjg.69.137
Kasahara S (2009) Introdução à pesquisa em Citogenética de Vertebrados. Ribeirão Preto, Brazil
Lattke JE (1990) Revisión del género Gnamptogenys Roger en Venezuela (Hymenoptera: Formicidae). Acta Terramaris 2:1–47
Lattke JE (1995) Revision of the ant genus Gnamptogenys in the New World (Hymenoptera: Formicidae). J Hymenopt Res 4:137–193
Lattke JE (2004) A taxonomic revision and phylogenetic analysis of the ant genus Gnamptogenys Roger in Southeast Asia and Australasia (Hymenoptera: Formicidae: Ponerinae). Univ Calif publ entomol 122:1–266. https://doi.org/10.1525/california/9780520098442.001.0001
Lattke JE, Fernández F, Arias-Penna TM, Palacio EE, Mackay W, Mackay E (2008) Género Gnamptogenys Roger. In: Jiménez E, Fernández F, Arias-Penna TM, Lozano-Zambrano FH (eds) Sistemática, Biogeografía y conservación de las hormigas cazadoras de Colombia, 1st edn. Bogotá D.C, Colombia, pp 66–107
Levan A, Fredga K, Sandberg A (1964) Nomenclature for centromeric position on chromosomes. Hereditas 52:201–220. https://doi.org/10.1111/j.1601-5223.1964.tb01953.x
Lorite P, Palomeque T (2010) Karyotype evolution in ants (Hymenoptera: Formicidae), with a review of the known ant chromosome numbers. Myrmecol News 13:89–102
Mariano CSF, Pompolo SG, Delabie JHC, Campos LAO (2001) Estudos cariotípicos de algumas espécies neotropicais de Camponotus Mayr (Hymenoptera: Formicidae). Rev Bras Entomol 45:267–274
Mariano CSF, Pompolo SG, Barros LAC, Mariano-Neto E, Campiolo S, Delabie JCHD (2008) A biogeographical study of the threatened ant Dinoponera lucida Emery (Hymenoptera: Formicidae: Ponerinae) using a cytogenetic approach. Insect Conserv Diver 1:161–168. https://doi.org/10.1111/j.1752-4598.2008.00022.x
Mariano CSF, Santos IS, Silva JG, Costa MA, Pompolo SG (2015) Citogenética e evolução do cariótipo em formigas poneromorfas. In: Delabie JHC, Feitosa RM, Serrao JE, Mariano CSF, Majer JD (eds) As formigas poneromorfas do Brasil, 1st edn. Ilhéus, Brasil, pp 102–125
Moen DS, Irschick DJ, Wiens JJ (2013) Evolutionary conservatism and convergence both lead to striking similarity in ecology, morphology and performance across continents in frogs. P Roy Soc B-Biol Sci es 280(1773):20132156. https://doi.org/10.1098/rspb.2013.2156
Palomeque T, Chica E, Díaz de la Guardia G (1993) Supernumerary chromosome segments in different genera of Formicidae. Genetica. 90:17–29
Pereira JOP (2006) Diversidade genética da abelha sem ferrão Melipona quinquefasciata baseada no sequenciamento das regiões ITS1 parcial e 18S do DNA ribossômico nuclear. Thesis, Universidade Federal do Ceará, Fortaleza, Brazil
Phimpan S, Tanomtong A, Jumrusthanasan S, Supiwong W, Siripiyasing P, Sanoamuang L (2013) First report of NOR polymorphism and chromosome analysis of John’s Snapper, Lutjanus johnii (Perciformes, Lutjanidae) in Thailand. Cytologia 78(4):335–344. https://doi.org/10.1508/cytologia.78.335
Pinkel D, Straume T, Gray JW (1986) Cytogenetic analysis using quantitative, high-sensitivity, fluorescence hybridization. PNAS USA 83:2934–2938. https://doi.org/10.1073/pnas.83.9.2934
Santos IS, Mariano CSF, Delabie JHC, Costa MA, Carvalho AF, Silva JG (2016) "Much more than a neck": karyotype differentiation between Dolichoderus attelaboides (Fabricius, 1775) and Dolichoderus decollatus F. Smith, 1858 (Hymenoptera: Formicidae) and karyotypic diversity of five other Neotropical species of Dolichoderus Lund, 1831. Myrmecol News 23:61–69
Santos RP, Mariano CSF, Delabie JHC, Costa MA, Lima KM, Pompolo SG, Fernandes IO, Miranda EA, Carvalho AF, Silva JG (2018) Genetic characterization of some Neoponera (Hymenoptera: Formicidae) populations within the foetida species complex. J Insect Sci 18(4):14; 1–14; 7. https://doi.org/10.1093/jisesa/iey079
Schubert I, Lysak MA (2011) Interpretation of karyotype evolution should consider chromosome structural constraints. Trends Genet 27(6):207–216. https://doi.org/10.1016/j.tig.2011.03.004
Schweizer D (1980) Simultaneous fluorescent staining of R bands and specific heterocromatic regions (DA/DAPI-bands) in human chromosomes. Cytogenet Cell Genet 27:190–193. https://doi.org/10.1159/000131482
Silva-Rocha PNC, Correia JPSO, Mariano CSF, Delabie JHC, Costa MA (2017) A note on the karyotype and morphology of the ant Platythyrea sinuata (Roger, 1860) (Formicidae, Ponerinae, Plathytyreini). Sociobiology 64(4):484–487. https://doi.org/10.13102/sociobiology.v64i4.1820
Sumner AT (2003) Chromosomes: Organization and Function. North Berwick – United Kingdom
Teixeira GA, Barros LAC, Aguiar HJAC, Pompolo SG (2017) Comparative physical mapping of 18S rDNA in the karyotypes of six leafcutter ant species of the genera Atta and Acromyrmex (Formicidae: Myrmicinae). Genetica 145:351–357. https://doi.org/10.1007/s10709-017-9970-1
Vicari MR, Artoni RF, Bertollo LA (2003) Heterochromatin polymorphism associated with 18S rDNA: a differential pathway among Hoplias malabaricus fish populations. Cytogenet Genome Res 101(1):24–28. https://doi.org/10.1159/000073413
Vicari MR, Almeida MC, Bertollo LAC, Moreira-Filho O, Artoni RF (2006) Cytogenetic analysis and chromosomal characteristics of the polymorphic 18S rDNA in the fish Prochilodus lineatus (Characiformes, Prochilodontidae). Genet Mol Biol 29(4):621–625. https://doi.org/10.1590/S1415-47572006000400008
Acknowledgments
The authors are grateful to Silvia G. Pompolo for valuable advices during this project, Júlio C. M. Chaul for kindly yielding Gnamptogenys triangularis for cytogenetic analysis and to Dr. Jacques H. Delabie for species identification. The authors also thank the Laboratório de Biologia Molecular de Insetos of the Universidade Federal de Viçosa (UFV) for technical support. GAT thanks the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) for the scholarship granted.
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling Editor: Georg Krohne
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Fig. S1
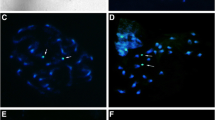
Fluorochrome staining of Gnamptogenys from mordax and rastrata groups. (a, b) Metaphases of G. triangularis (rastrata group), and (c - f) Gnamptogenys regularis (mordax group). Metaphases stained with Chromomycin A3 (a, c, e) and 4’6- diamidino-2-phenylindole (b, d, f), respectively. Arrows indicate regions rich in GC base pairs. Bar = 5 μm. (PNG 4159 kb)
Fig. S2
Fluorochrome staining of Gnamptogenys striatula. Female metaphases of individuals from Rio de Janeiro (2n = 32) and Petrópolis (2n = 34) stained with Chromomycin A3 (a, c) and 4’6- diamidino-2-phenylindole (b, d), respectively. Arrows indicate regions rich in GC base pairs. Bar = 5 μm. (PNG 2942 kb)
Fig. S3
Fluorochrome staining of Gnamptogenys moelleri. Female metaphases of individuals from Petrópolis (2n = 34), Viçosa (2n = 34), and Açailândia (2n = 44) stained with Chromomycin A3 (a, c, e) and 4’6- diamidino-2-phenylindole (b, d, f), respectively. Arrows indicate regions rich in GC base pairs. Bar = 5 μm. (PNG 5513 kb)
Rights and permissions
About this article
Cite this article
Teixeira, G.A., Barros, L.A.C., Lopes, D.M. et al. Cytogenetic variability in four species of Gnamptogenys Roger, 1863 (Formicidae: Ectatomminae) showing chromosomal polymorphisms, species complex, and cryptic species. Protoplasma 257, 549–560 (2020). https://doi.org/10.1007/s00709-019-01451-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-019-01451-6