Abstract
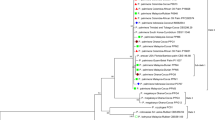
During a survey in June 2011, severe leaf yellow mosaic disease was observed on about 45 % plants of Jatropha curcas growing in the Katerniaghat wildlife sanctuary in India. An association of a begomovirus with disease was detected in 15 out of 20 samples by PCR using begomovirus genus-specific primers and total DNA isolated from symptomatic leaf samples. For identification of the begomovirus, the complete genome was amplified using a Phi-29 DNA-polymerase-based rolling-circle amplification kit and total DNA from five representative samples and then digested with BamHI. The linearized RCA products were cloned and sequenced. Their GenBank accession numbers are JN698954 (SKRK1) and JN135236 (SKRK2). The sequences of the two begomovirus isolates were 97 % identical to each other and no more than 86 % to those of jatropha mosaic India virus (JMIV, HM230683) and other begomoviruses reported worldwide. In phylogenetic analysis, SKRK1 and SKRK2 clustered together and showed distant relationships to jatropha mosaic India virus, Jatropha curcas mosaic virus, Indian cassava mosaic virus, Sri Lankan cassava mosaic virus and other begomoviruses. Based on 86 % sequence identities and distant phylogenetic relationships to JMIV and other begomoviruses and the begomovirus species demarcation criteria of the ICTV (<89 % sequence identity of complete DNA-A genome), the begomovirus isolates associated with leaf yellow mosaic disease of J. curcas were identified as members of a new begomovirus species and provisionally designated as jatropha leaf yellow mosaic Katerniaghat virus (JLYMKV). Agroinfectious clones of the DNA molecule of the begomovirus isolate were also generated, and the fulfillment of Koch’s postulates was demonstrated in J. curcas plants.


Similar content being viewed by others
References
Singh P, Singh S, Mishra SP, Bhatia SK (2010) Molecular characterization of genetic diversity in Jatropha curcas L. Genes Genomes Genomics 4:1–8
Narayana DSA, Shankarappa KS, Govindrappa MR, Prameela HA, Gururaj Rao MR, Rangaswamy KT (2006) Natural occurrence of Jatropha mosaic virus disease in India. Curr Sci 91:584–586
Narayana DSA, Rangaswamy KT, Shankarappa KS, Maruthi MN, Lakshminarayana Reddy CN, Rekha AR, Keshava Murthy KV (2007) Distinct begomoviruses closely related to Cassava mosaic viruses cause Indian jatropha mosaic disease. Int J Virol 3:1–11
Tewari (2007) Jatropha and biodiesel. Ocean Books Ltd
Raj SK, Snehi SK, Kumar S, Khan MS, Pathre U (2008) First molecular identification of a begomovirus in India that is closely related to Cassava mosaic virus and causes mosaic and stunting of Jatropha curcas L. Australas Plant Dis Notes 3:69–71
Gao S, Qu J, Chua NH, Ye J (2010) A new strain of Indian cassava mosaic virus causes a mosaic disease in the biodiesel crop Jatropha curcas. Arch Virol 155:607–612
Ramkat RC, Alberto C, Fatemeh M, Laimer M (2011) Biotechnological approaches to determine the impact of viruses in the energy crop plant Jatropha curcas. Virol J 3:386
Snehi SK, Srivastava A, Raj SK (2012) Biological characterization and complete genome sequence of a possible strain of Indian cassava mosaic virus from Jatropha curcas in India. J Phytopathol 160:547–553
Kashina BD, Alegbejo MD, Banwo OO, Nielsen SL, Nicolaisen M (2013) Molecular identification of a new begomovirus associated with mosaic disease of Jatropha curcas L in Nigeria. Arch Virol 158(2):511–514
Srivastava A, Jaidi M, Kumar S, Raj SK (2014) Molecular identification of a new begomovirus associated with leaf crumple disease of Jatropha curcas L in India. Arch Virol. doi:10.1007/s00705-014-2288-8
Singh R, Raj SK, Prasad V (2008) Molecular characterization of a strain of Squash leaf curl China virus from North India. J Phytopathol 156(4):222–228
Reddy CRV, Colvin J, Muniyappa V, Seal S (2005) Diversity and distribution of begomoviruses infecting tomato in India. Arch Virol 150:845–867
Rojas MR, Gilbertson RL, Russell DR, Maxwell DP (1993) Use of degenerate primers in the polymerase chain reaction to detect whitefly transmitted geminiviruses. Plant Dis 77:340–347
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol Biol Evol 30:2725–2729
Fauquet CM, Briddon RW, Brown JK, Moriones E, Stanley J, Zerbini M, Zhou X (2008) Geminivirus strain demarcation and nomenclature. Arch Virol 153(4):783–821
Fienberg AP, Vogelstein B (1983) A technique for radiolabeling DNA restriction endonuclease fragments to high specific activity. Ann Biochem 132:6
Acknowledgments
The authors are thankful to the Director, CSIR-National Botanical Research Institute, Lucknow, India, for facilities, and the Department of Biotechnology (DBT), India, for funding.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Srivastava, A., Kumar, S., Jaidi, M. et al. Molecular characterization of a new begomovirus associated with leaf yellow mosaic disease of Jatropha curcas in India. Arch Virol 160, 1359–1362 (2015). https://doi.org/10.1007/s00705-015-2375-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-015-2375-5