Abstract
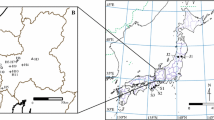
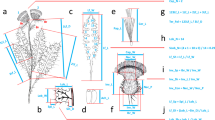
The presence and extent of hybridization within the Chenopodium album aggregate (Amaranthaceae) is still unclear. Although many hybrid combinations have been described, their existence in the field has never been systematically studied and verified. The main aim of this study was to ascertain the extent of interspecific hybridization between the diploid species C. ficifolium and C. suecicum using highly variable nuclear microsatellite markers. Due to the absence of such kind of molecular markers for the whole C. album group, we divided the analysis into two steps: (1) Eleven microsatellite loci designed for the closely related species C. quinoa were cross-amplified in five Eurasian species of the C. album diploid–polyploid complex, i.e. C. album s.s. (6x), C. striatiforme (4x), C. strictum (4x), C. ficifolium (2x) and C. suecicum (2x); (2) For the detection of interspecific hybridization between C. ficifolium and C. suecicum, we sampled 480 individuals from five localities in Central Europe. We also investigated morphological differences between the parental taxa and their hybrid and devised a key for their determination. Analysis of variation in microsatellite loci using Bayesian methods, PCoA and Neighbour-joining tree identified 32 F1 hybrids. These F1 hybrids, described here as C. paradoxum Mandák, formed a cluster between well-differentiated parental species, combining the morphological characters of both their parents. Moreover, genetic analyses also recognized several F2 or backcross hybrids, whose delimitation, mainly from C. suecicum and F1 hybrids, based on morphological characters, is problematic.







Similar content being viewed by others
References
Abbott RJ (1992) Plant invasions, interspecific hybridisation and the evolution of new plant taxa. Trends Ecol Evol 7:401–405. https://doi.org/10.1016/0169-5347(92)90020-C
Abbott RJ, Albach D, Ansell S, Arntzen JW, Baird SJ et al (2013) Hybridization and speciation. J Evol Biol 26:229–246. https://doi.org/10.1111/j.1420-9101.2012.02599.x
Aellen P (1960) Chenopodium. In: Hegi G (ed) Illustrierte Flora von Mitteleuropa, vol. 3(2), 2nd edn. Parey, Berlin
Aellen P (1972) Das vorkommen einer neuen hybride von Chenopodium ficifolium Sm. x Chenopodium viride L. (Chenopodium x gruellii Aellen hybr. nov.) in der ČSSR. Acta Mus Morav 57:167–170
Aellen P, Just T (1943) Key Synopsis of the American Species of the Genus Chenopodium L. Amer Midl Naturalist 30:47–76. https://doi.org/10.2307/2421263
Arnold ML (1997) Natural hybridization and evolution. Oxford University Press, New York
Blom C (1961) Bidrag till kännedomen om Sveriges adventiv och ruderatflora V. Acta Horti Gotob 24:61–133
Clemants S, Mosyakin S (2003) Chenopodiaceae. In: Flora of North America Editorial Committee (ed) Flora of North America North of Mexico, 4th edn. Oxford University Press, Oxford, pp 258–404
Cole MJ (1957) Variation and interspecific relationships of Chenopodium album L. in Britain. PhD Thesis, University of Southampton, United Kingdom
Cole MJ (1961) Interspecific relationships and intraspecific variation of Chenopodium album L. in Britain. I. The taxonomic delimitation of the species. Watsonia 5:47–58
Dostálek J, Hejný S, Husák Š, Schwarzová T, Dvořák F (1990) Chenopodium L., merlík. In: Hejný S, Slavík B (eds) Květena České republiky 2. Academia, Praha, pp 223–265
Dvořák F (1990) Study of Chenopodium interjectum J. Murr, Ch. mixtifolium J. Murr and Ch. laciniatum J. Murr. Feddes Repert 101:347–371. https://doi.org/10.1002/fedr.19901010706
Dvořák F (1992a) Study of Chenopodium purpurascens B. de Juss. ex Jacq. and on some related taxa. Feddes Repert 103:153–173. https://doi.org/10.1002/fedr.19921030302
Dvořák F (1992b) Study of Chenopodium subopulifolium J. Murr emend. D. Feddes Repert 103:49–69. https://doi.org/10.1002/fedr.19921030109
Dvořák F (1993) Relationships and diagnostic characters of Chenopodium striatiforme J. Murr, C. striatum (Krašan) J. Murr and C. strictum Roth. Feddes Repert 104:439–449. https://doi.org/10.1002/fedr.19931040704
Dvořák F (1994) Study of some species subsumed under Chenopodium probstii A. and on C. purpurascens B. de Juss. ex Jacq. Feddes Repert 105:113–139. https://doi.org/10.1002/fedr.19941050302
Ehrich D (2006) AFLPdat: a collection of R functions for convenient handling of AFLP data. Molec Ecol Notes 6:603–604. https://doi.org/10.1111/j.1471-8286.2006.01380.x
Ellstrand NC, Elam DR (1993) Population genetic consequences of small population size: implications for plant conservation. Annual Rev Ecol Evol Syst 24:217–242. https://doi.org/10.1146/annurev.es.24.110193.001245
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Molec Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Excoffier L, Smouse P, Quattro J (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131:479–491
Grant PR, Grant BR (2014) Evolutionary biology: speciation undone. Nature 507:178–179. https://doi.org/10.1038/507178b
Grant V (1981) Plant speciation, 2nd edn. Columbia University Press, New York
Hammer Ø, Harper DAT, Ryan PD (2001) PAST: paleontological statistics software package for education and data analysis. Palaeontol Electronica 4:1–9. http://palaeo-electronica.org/2001_1/past/issue1_01.htm
Jacobsen SE (2011) The situation for Quinoa and its production in southern Bolivia: from economic success to environmental disaster. J Agron Crop Sci 197:390–399. https://doi.org/10.1111/j.1439-037X.2011.00475.x
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806. https://doi.org/10.1093/bioinformatics/btm233
Jalas J, Suominen J (1980) Atlas Florae Europaeae distribution of vascular plants in Europe 5. Chenopodiaceae to Basellaceae. The Committee for Mapping the Flora of Europe and Societas Biologica Fennica Vanamo, Helsinki
Jarvis DE, Kopp OR, Jellen EN, Mallory MA, Pattee J, Bonifacio A, Coleman CE, Stevens MR, Fairbanks DJ, Maughan PJ (2008) Simple sequence repeat marker development and genetic mapping in quinoa (Chenopodium quinoa Willd.). J Genet 87:39–51
Jüttersonke B, Arlt K (1989) Experimentelle Untersuchungen über die infraspezifische Struktur von Chenopodium album L. sowie Untersuchungen an Chenopodium suecicum. J Murr Feddes Repert 100:1–63
Krak K, Vít P, Belyayev A, Douda J, Hreusová L, Mandák B (2016) Allopolyploid origin of Chenopodium album s. str. (Chenopodiaceae): a molecular and cytogenetic insight. PLoS One 11:e0161063. https://doi.org/10.1371/journal.pone.0161063
Leong-Škorničková J, Šída O, Jarolímová V, Sabu M, Fér T, Trávníček P, Suda J (2007) Chromosome numbers and genome size variation in Indian species of Curcuma (Zingiberaceae). Ann Bot (Oxford) 100:505–526. https://doi.org/10.1093/aob/mcm144
Mallet J (2007) Hybrid speciation. Nature 446:279–283. https://doi.org/10.1038/nature05706
Mandák B, Pyšek P, Lysák M, Suda J, Krahulcová A, Bímová K (2003) Variation in DNA-ploidy levels of Reynoutria taxa in the Czech Republic. Ann Bot (Oxford) 92:265–272. https://doi.org/10.1093/aob/mcg141
Mandák B, Trávníček P, Paštová L, Kořínková D (2012) Is hybridization involved in the evolution of the Chenopodium album aggregate? An analysis based on chromosome counts and genome size estimation. Flora 207:530–540. https://doi.org/10.1016/j.flora.2012.03.010
Mandák B, Krak K, Vít P, Pavlíková Z, Lomonosova MN, Habibi F, Lei W, Jellen EN, Douda J (2016) How genome size variation is linked with evolution within Chenopodium sensu lato. Perspect Pl Ecol Evol Syst 23:18–32. https://doi.org/10.1016/j.ppees.2016.09.004
Mason SL, Stevens MR, Jellen EN, Bonifacio A, Fairbanks DJ, Coleman CE, McCarty RR, Rasmussen AG, Maughan PJ (2005) Development and use of microsatellite markers for germplasm characterization in Quinoa (Chenopodium quinoa Willd.). Crop Sci 45:1618–1630. https://doi.org/10.2135/cropsci2004.0295
Murr J (1896) Über einige kritische Chenopodium-Formen. Deutsche Bot Monatsschr 14:32–37
Murr J (1901) Zur Chenopodium-Frage. II. Deutsche Bot Monatsschr 19:37–40
Nei M (1973) The theory and estimation of genetic distance. In: Morton NE (ed) Genetics of population structure. University of Hawai Press, Honolulu, pp 45–54
Nordborg M, Hu TT, Ishino Y, Jhaveri J, Toomajian C et al (2005) The pattern of polymorphism in Arabidopsis thaliana. PLoS Biol 3:1289–1299. https://doi.org/10.1371/journal.pbio.0030196
Olšavská K, Perný M, Španiel S, Šingliarová B (2012) Nuclear DNA content variation among perennial taxa of the genus Cyanus (Asteraceae) in Central Europe and adjacent areas. Pl Syst Evol 298:1463–1482. https://doi.org/10.1007/s00606-012-0650-4
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Pritchard JK, Wen X, Falush D (2009) STRUCTURE ver. 2.3.4. University of Chicago, Chicago. Available at: http://pritch.bsd.uchicago.edu/. Accessed 25 Apr 2015
R Core Team (2014) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. Available at: https://www.R-project.org/
Rahiminejad MR (1995) Taxonomy and biosystematics of the Chenopodium album aggregate. PhD Thesis, University of Leicester, Leicester
Rieseberg LH, Wendel JF (1993) Introgression and its consequences in plants. In: Harrison R (ed) Hybrid zones and the evolutionary process. Oxford University Press, New York, pp 70–109
Rosenberg NA (2004) Distruct: a program for the graphical display of population structure. Molec Ecol Notes 4:137–138. https://doi.org/10.1046/j.1471-8286.2003.00566.x
Schlüter PM, Harris SA (2006) Analysis of multilocus fingerprinting data sets containing missing data. Molec Ecol Notes 6:569–572. https://doi.org/10.1111/j.1471-8286.2006.01225.x
Soltis PS, Soltis DE (2009) The role of hybridization in plant speciation. Annual Rev Pl Biol 60:561–588. https://doi.org/10.1146/annurev.arplant.043008.092039
Štorchová H, Hrdličková R, Chrtek J, Tetera M, Fitze D, Fehrer J (2000) An improved method of DNA isolation from plants collected in the field and conserved in saturated NaCl/CTAB solution. Taxon 49:79–84. https://doi.org/10.2307/1223934
Štorchová H, Drabešová J, Cháb D, Kolář J, Jellen EN (2015) The introns in FLOWERING LOCUS T-LIKE (FTL) genes are useful markers for tracking paternity in tetraploid Chenopodium quinoa Willd. Genet Resources Crop Evol 62:913–925. https://doi.org/10.1007/s10722-014-0200-8
Suda J, Krahulcová A, Trávníček P, Rosenbaumová R, Peckert T, Krahulec F (2007) Genome size variation and species relationships in Hieracium sub-genus Pilosella (Asteraceae) as inferred by flow cytometry. Ann Bot (Oxford) 100:1323–1335. https://doi.org/10.1093/aob/mcm218
Suda J, Trávníček P, Mandák B, Berchová-Bímová K (2010) Genome size as a marker for identifying the invasive alien taxa in Fallopia section Reynoutria. Preslia 82:97–106
Thomas CD (2015) Rapid acceleration of plant speciation during the Anthropocene. Trends Ecol Evol 30:448–455. https://doi.org/10.1016/j.tree.2015.05.009
Uotila P (1977) Chenopodium strictum subsp. striatiforme in the Baltic Sea area. Ann Bot Fenn 14:199–205
Uotila P (1978) Variation, distribution and taxonomy of Chenopodium suecicum and C. album in N Europe. Acta Bot Fenn 108:1–35
Uotila P (1997) Chenopodium L. In: Rechinger KH (ed) Flora Iranica, 172nd edn. Akademische Druck-u. Verlagsanstalt, Graz, pp 24–59
Uotila P (2001a) Chenopodium L. In: Jonsell B (ed) Flora Nordica 2. The Royal Swedish Academy of Sciences, Stockholm, pp 4–31
Uotila P (2001b) Taxonomic and nomenclatural notes on Chenopodium (Chenopodiaceae) for Flora Nordica. Ann Bot Fenn 38:95–97
Vít P, Krak K, Trávníček P, Douda J, Lomonosova MN, Mandák B (2016) Genome size stability across Eurasian Chenopodium species (Amaranthaceae). Bot J Linn Soc 182:637–649. https://doi.org/10.1111/boj.12474
Walsh BM, Adhikary D, Maughan PJ, Emshwiller E, Jellen EN (2015) Chenopodium polyploidy inferred from salt overly sensitive (SOS1) data. Amer J Bot 102:533–543. https://doi.org/10.3732/ajb.1400344
Wilson H (1980) Artificial hybridization among species of Chenopodium sect. Chenopodium. Syst Bot 5:253–263. https://doi.org/10.2307/2418372
Wilson H, Manhart J (1993) Crop/weed gene flow: Chenopodium quinoa Willd. and C. berlandieri Moq. Theor Appl Genet 86:642–648. https://doi.org/10.1007/BF00838721
Acknowledgements
We would like to thank Dagmar Boltíková, Petr Vít, Jan Machač and Pavel Trávníček for their help in the laboratory and field. Fred Rooks is thanked for helping with the language of the manuscript. We also would like to thank Pertti Uotila and Jindřich Chrtek for their discussion concerning nothospecies description.
Funding
This study was supported by the Grant Agency of the Czech Republic (13-02290S) and as part of long-term research development project RVO 67985939.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The authors comply will all rules of the journal following the COPE guidelines; all authors have contributed and approved the final manuscript.
Additional information
Handling editor: Christoph Oberprieler.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Information on Electronic Supplementary Material
Information on Electronic Supplementary Material
Online Resource 1. Samples used for the cross-amplification of microsatellite primers from Chenopodium quinoa to European representatives of C. album aggregate.
Online Resource 2 . Populations used for initial testing, multiplex optimization and estimation of genetic variability at microsatellite loci.
Online Resource 3. Populations used for of the detection of homoploid hybridization between Chenopodium ficifolium and C. suecicum.
Online Resource 4. Numbers and size ranges of alleles for eleven cross-amplified microsatellite loci of five species of the Chenopodium album aggregate collected across Europe.
Online Resource 5. Overall genetic diversity based on 11 microsatellite loci in Chenopodium ficifolium, C. suecicum and their hybrid C. ×paradoxum.
Online Resource 6 . Estimation of the number of genetic clusters (K) based on (A) LnP(D), (B) similarity coefficient and (C) Mean DeltaK based on results of STRUCTURE analysis.
Rights and permissions
About this article
Cite this article
Hodková, E., Mandák, B. An overlooked hybrid between the two diploid Chenopodium species in Central Europe determined by microsatellite and morphological analysis. Plant Syst Evol 304, 295–312 (2018). https://doi.org/10.1007/s00606-017-1477-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00606-017-1477-9