Abstract
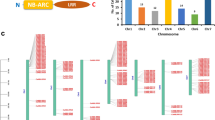
Powdery mildew locus O (Mlo) gene family is one of the largest seven transmembrane protein-encoding gene families. The Mlo proteins act as negative regulators of powdery mildew resistance and a loss-of-function mutation in Mlo is known to confer broad-spectrum resistance to powdery mildew. In addition, the Mlo gene family members are known to participate in various developmental and biotic and abiotic stress response-related pathways. Therefore, a genome-wide similarity search using the characterized Mlo protein sequences of Arabidopsis thaliana was carried out to identify putative Mlo genes in soybean (Glycine max) genome. This search identified 39 Mlo domain containing protein-encoding genes that were distributed on 15 of the 20 G. max chromosomes. The putative promoter regions of these Mlo genes contained response elements for different external stimuli, including different hormones and abiotic stresses. Of the 39 GmMlo proteins, 35 were rich (8.7–13.1 %) in leucine, while five were serine-rich (9.2–11.9 %). Furthermore, all the GmMlo members were localized in the plasma membrane. Phylogenetic analysis of the GmMlo and the AtMlo proteins classified them into three main clusters, and the cluster I comprised two sub-clusters. Multiple sequence alignment visualized the location of seven transmembrane domains, and a conserved CaM-binding domain. Some of the GmMlo proteins (GmMlo10, 20, 22, 23, 32, 36, 37) contained less than seven transmembrane domains. The motif analysis yielded 27 motifs; out of these, motif 2, the only motif present in all the GmMlos, was highly conserved and three amino acid residues were essentially invariant. Five of the GmMlo members were much smaller in size; presumably they originated through deletion following a gene duplication event. The presence of a large number of GmMlo members in the G. max genome may be due to its paleopolyploid nature and the large genome size as compared to that of Arabidopsis. The findings of this study may further help in characterization and isolation of individual GmMlo members.






Similar content being viewed by others
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Bai Y, Pavan S, Zheng Z, Zappel NF, Reinstädler A, Lotti C, De Giovanni C, Ricciardi L, Lindhout P, Visser R, Theres K, Panstruga R (2008) Naturally occurring broad-spectrum powdery mildew resistance in a Central American tomato accession is caused by loss of mlo function. Mol Plant Microbe Interact 21:30–39
Bailey TL, Elkan C (1994) Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc Second Int Conf Intell Syst Mol Biol. AAAI Press, Menlo Park, pp 28–36
Bailey TL, Boden M, Buske FA, Frith M, Grant CE, Clementi L, Ren J, Li WW, Noble WS (2009) MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res 37:W202–W208
Buchanan SG, Gay NJ (1996) Structural and functional diversity in the leucine-rich repeat family of proteins. Prog Biophys Mol Biol 65:1–44
Burge C, Karlin S (1997) Prediction of complete gene structures in human genomic DNA. J Mol Biol 268:78–94
Büschges R, Hollricher K, Panstruga R, Simons G, Wolter M, Frijters A, van Daelen R, van der Lee T, Diergaarde P, Groenendijk J, Topsch S, Vos P, Salamini F, Schulze-Lefert P (1997) The barley MLO gene: a novel control element of plant pathogen resistance. Cell 88:695–705
Chen Z, Hartmann HA, Wu MJ, Friedman EJ, Chen J, Pulley M, Schulze-Lefert P, Panstruga R, Jones AM (2006) Expression analysis of the AtMLO gene family encoding plant-specific seven-transmembrane domain proteins. Plant Mol Biol 60:583–597
Chen L, Shiotani K, Togashi T, Miki D, Aoyama M, Wong HL, Kawasaki T, Shimamoto K (2010) Analysis of the Rac/Rop small GTPase family in rice: expression, subcellular localization and role in disease resistance. Plant Cell Physiol 51:585–595
Devoto A, Piffanelli P, Nilsson I, Wallin E, Panstruga R, Heijne GV, Schulze-Lefert P (1999) Topology, subcellular localization, and sequence diversity of the MLO family in plants. J Biol Chem 274:34993–35004
Devoto A, Hartmann HA, Piffanelli P, Elliott C, Simmons C, Taramino G, Goh CS, Cohen FE, Emerson BC, Schulze-Lefert P, Panstruga R (2003) Molecular phylogeny and evolution of the plant-specific seven-transmembrane MLO family. J Mol Evol 56:77–88
Drummond A, Ashton B, Cheung M, Heled J, Kearse M, Moir R, Stones-Havas S, Thierer T, Wilson A (2008) Geneious v4.0. Biomatters Ltd. Auckland, New Zealand
Dunleavy JM (1980) Yield losses in soybeans induced by powdery mildew. Plant Dis 64:291–292
Elliott C, Uller JM, Miklis M, Bhat RA, Schulze-Lefert P, Panstruga R (2005) Conserved extracellular cysteine residues and cytoplasmic loop–loop interplay are required for functionality of the heptahelical MLO protein. Biochem J 385:243–254
Feechan A, Jermakow AM, Dry IB (2009) Grapevine MLO candidates required for powdery mildew pathogenicity? Plant Signal Behav 4:522–523
Forsthoefel NR, Cutler K, Port MD, Yamamoto T, Vernon DM (2005) PIRLs: a novel class of plant intracellular leucine-rich repeat proteins. Plant Cell Physiol 46:913–922
Gasteiger E, Hoogland C, Gattiker A, Duvaud S, Wilkins MR, Appel RD, Bairoch A (2005) The proteomics protocols. In: Walker JM (ed) Humana Press
Guo AY, Zhu QH, Chen X, Luo JC (2007) GSDS: a gene structure display server. Yi Chuan 29:1023–1026
Higo K, Ugawa Y, Iwamoto M, Korenaga T (1999) Plant cis-acting regulatory DNA elements (PLACE) database. Nucleic Acids Res 27:297–300
Hoefle C, Huesmann C, Schultheiss H, Börnke F, Hensel G, Kumlehn J, Hückelhoven R (2011) A barley ROP GTPase ACTIVATING PROTEIN associates with microtubules and regulates entry of the barley powdery mildew fungus into leaf epidermal cells. Plant Cell 23:2422–2439
Huesmann C, Reiner T, Hoefle C, Preuss J, Jurca ME, Domoki M, Fehér A, Hückelhoven R (2012) Barley ROP binding kinase1 is involved in microtubule organization and in basal penetration resistance to the barley powdery mildew fungus. Plant Physiol 159:311–320
Jørgensen IH (1992) Discovery, characterization and exploitation of Mlo powdery mildew resistance in barley. Euphytica 63:141–152
Jung EH, Jung HW, Lee SC, Han SW, Heu S, Hwang BK (2004) Identification of a novel pathogen-induced gene encoding a leucine-rich repeat protein expressed in phloem cells of Capsicum annuum. Biochim Biophys Acta 1676:211–222
Kêdzierski L, Montgomery J, Curtis J, Handman E (2004) Leucine-rich repeats in host–pathogen interactions. Arch Immunol Ther Exp (Warsz) 52:104–112
Kemmerling B, Schwedt A, Rodriguez P, Mazzotta S, Frank M, Qamar SA, Mengiste T, Betsuyaku S, Parker JE, Müssig C, Thomma BP, Albrecht C, de Vries SC, Hirt H, Nürnberger T (2007) The BRI1-associated kinase 1, BAK1, Has a brassinolide-independent role in plant cell-death control. Curr Biol 17:1116–1122
Kim DS, Hwang BK (2012) The pepper MLO gene, CaMLO2, is involved in the susceptibility cell-death response and bacterial and oomycete proliferation. Plant J 72:843–855
Kim MC, Lee SH, Kim JK, Chun HJ, Choi MS, Chung WS, Moon BC, Kang CH, Park CY, Yoo JH, Kang YH, Koo SC, Koo YD, Jung JC, Kim ST, Schulze-Lefert P, Lee SY, Cho MJ (2002a) MLO, a modulator of plant defense and cell death, is a novel calmodulin-binding protein. J Biol Chem 277:19304–19314
Kim MC, Panstruga R, Elliott C, Müller J, Devoto A, Yoon HW, Park HC, Cho MJ, Schulze-Lefert P (2002b) Calmodulin interacts with MLO protein to regulate defense against mildew in barley. Nature 416:447–450
Konishi S, Sasakuma T, Sasanuma T (2010) Identification of novel Mlo family members in wheat and their genetic characterization. Genes Genet Syst 85:167–175
Kumar J, Hückelhoven R, Beckhove U, Nagarajan S, Kogel KH (2001) A compromised Mlo pathway affects the response of barley to the necrotrophic fungus Bipolaris sorokiniana (teleomorph: Cochliobolus sativus) and its toxins. Phytopathology 91:127–133
Lecourieux D, Ranjeva R, Pugin A (2006) Calcium in plant defense-signalling pathways. New Phytol 171:249–269
Lescot M, Déhais P, Thijs G, Marchal K, Moreau Y, Van de Peer Y, Rouzé P, Rombauts S (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acid Res 30:325–327
Li J, Chory J (1997) A putative leucine-rich repeat receptor kinase involved in brassinosteroid signal transduction. Cell 90:929–938
Liu Q, Zhu H (2008) Molecular evolution of the Mlo gene family in Oryza sativa and their functional divergence. Gene 409:1–10
Lorek J, Panstruga R, Hückelhoven R (2010) The role of seven-transmembrane domain MLO proteins, heterotrimeric G-proteins, and monomeric RAC/ROPs in plant defense. In: Yalovsky S et al (eds) Integrated G proteins signaling in plants, signaling and communication in plants. Springer, Berlin, pp 197–220
Nibau C, Wu H, Cheung AY (2006) RAC/ROP GTPases: ‘hubs’ for signal integration and diversification in plants. Trends Plant Sci 11:309–315
Opalski KS, Schultheiss H, Kogel KH, Hückelhoven R (2005) The receptor-like MLO protein and the RAC/ROP family G-protein RACB modulate actin reorganization in barley attacked by the biotrophic powdery mildew fungus Blumeria graminis f. sp. hordei. Plant J 41:291–303
Osakabe Y, Maruyama K, Seki M, Satou M, Shinozaki K, Yamaguchi-Shinozaki K (2005) Leucine-rich repeat receptor-like Kinase1 is a key membrane-bound regulator of Abscisic acid early signaling in Arabidopsis. Plant Cell 17:1105–1119
Peterhänsel C, Freialdenhoven A, Kurth J, Kolsch R, Schulze-Lefert P (1997) Interaction analyses of genes required for resistance responses to powdery mildew in barley reveal distinct pathways leading to leaf cell death. Plant Cell 9:1397–1409
Piffanelli P, Zhou F, Casais C, Orme J, Schaffrath U, Collins N, Panstruga R, Schulze-Lefert P (2002) The barley MLO modulator of defense and cell death is responsive to biotic and abiotic stress stimuli. Plant Physiol 129:1076–1085
Punta M, Coggill PC, Eberhardt RY, Mistry J, Tate J, Boursnell C, Pang N, Forslund K, Ceric G, Clements J, Heger A, Holm L, Sonnhammer EL, Eddy SR, Bateman A, Finn RD (2012) The Pfam protein families database. Nucleic Acids Res 40:D290–D301
Quevillon E, Silventoinen V, Pillai S, Harte N, Mulder N, Apweiler R, Lopez R (2005) InterProScan: protein domains identifier. Nucleic Acids Res 33:116–120
Reddy VS, Ali GS, Reddy ASN (2003) Characterization of a pathogen-induced calmodulin-binding protein: mapping of four Ca2+-dependent calmodulin-binding domains. Plant Mol Biol 52:143–159
Reinstädler A, Müller J, Czembor JH, Piffanelli P, Panstruga R (2010) Novel induced MLO mutant alleles in combination with site-directed mutagenesis reveal functionally important domains in the heptahelical barley MLO protein. BMC Plant Biol 10:31–43
Schmutz J, Cannon SB, Schlueter J, Ma J, Mitros T, Nelson W, Hyten DL, Song Q, Thelen JJ, Cheng J, Xu D, Hellsten U, May GD, Yu Y, Sakurai T, Umezawa T, Bhattacharyya MK, Sandhu D, Valliyodan B, Lindquist E, Peto M, Grant D, Shu S, Goodstein D, Barry K, Futrell-Griggs M, Abernathy B, Du J, Tian Z, Zhu L, Gill N, Joshi T, Libault M, Sethuraman A, Zhang XC, Shinozaki K, Nguyen HT, Wing RA, Cregan P, Specht J, Grimwood J, Rokhsar D, Stacey G, Shoemaker RC, Jackson SA (2010) Genome sequence of the paleopolyploid soybean (Glycine max). Nature 463:178–183
Schultheiss H, Dechert C, Kogel K, Huckelhoven R (2002) A small GTP-binding host protein is required for entry of powdery mildew fungus into epidermal cells of barley. Plant Physiol 128:1447–1454
Shen Q, Zhao J, Du C, Xiang Y, Cao J, Qin X (2012) Genome-scale identification of MLO domain-containing genes in soybean (Glycine max L. Merr.). Genes Genet Syst 87:89–98
Singh VK, Singh AK, Chand R, Singh BD (2012) Genome wide analysis of disease resistance MLO gene family in sorghum (Sorghum bicolor L. Moench). J Plant Genom 2:18–27
Solovyev V, Kosarev P, Seledsov I, Vorobyev D (2006) Automatic annotation of eukaryotic genes, pseudogenes and promoters. Genome Biol 7(Suppl 1):10.1–10.12
Thompson JD, Higgins DG, Gibson TJ (1994) ClustalW: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Torii KU (2004) Leucine-rich repeat receptor kinases in plants: structure, function, and signal transduction pathways. Int Rev Cytol 234:1–46
Trabanco N, Pérez-Vega E, Campa A, Rubiales D, Ferreira JJ (2012) Genetic resistance to powdery mildew in common bean. Euphytica 186:875–882
Treangen T, Messeguer X (2006) M-GCAT: interactively and efficiently constructing large-scale multiple genome comparison frameworks in closely related species. BMC Bioinform 7:433–448
Tusnády GE, Simon I (1998) Principle governing amino acid composition of integral membrane proteins: application to topology prediction. J Mol Biol 283:489–506
Tusnády GE, Simon I (2001) The HMMTOP transmembrane topology prediction server. Bioinformatics 17:849–850
Wolter M, Hollricher K, Salamini F, Schulze-Lefert P (1993) The mlo resistance alleles to powdery mildew infection in barley trigger a developmentally controlled defense mimic phenotype. Mol Gen Genet 239:122–128
Yu CS, Chen YC, Lu CH, Hwang JK (2006) Prediction of protein subcellular localization. Proteins 64:643–651
Acknowledgments
This work was supported by the UGC research grant. We would also like to thank Centre of Bioinformatics; DBT funded SUB-DIC Centre at School of Biotechnology, Banaras Hindu University, Varanasi for providing us the necessary softwares and technical help.
Ethical standard
The experiments performed in the above article comply with the current laws of the country in which the experiment has been done.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Y. Van de Peer.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Deshmukh, R., Singh, V.K. & Singh, B.D. Comparative phylogenetic analysis of genome-wide Mlo gene family members from Glycine max and Arabidopsis thaliana . Mol Genet Genomics 289, 345–359 (2014). https://doi.org/10.1007/s00438-014-0811-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-014-0811-y