Abstract
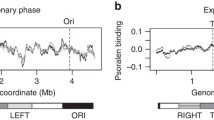
Single stranded chains of biological DNA show a widespread occurrence of parity for complementary nucleotides, i.e., A=T, G=C. This has been referred to as A-T, G-C symmetry. A distinction must be made between this, which this paper calls mirror symmetry, and twofold symmetry, where complementary nucleotide parity occurs between two segments, of the same length and equidistant from a symmetry center, along a single-stranded DNA chain. I have analysed the sequence of Chromosome I of Saccharomyces cerevisiae for the occurrence of complementary nucleotide symmetry. Open reading frame (ORF) sequences made up 63% of the total chromosome length and most of them were asymmetric for both A-T and G-C. The sign of A-T asymmetry was correlated with transcriptional orientation (A>T for sense and A<T for antisense ORFs), whereas G-C asymmetry was not. However, long single-stranded segments of Chromosome I were A-T mirror symmetric because they contained similar frequencies of ORFs in both transcriptional orientations. The same results were obtained with the AA-TT pair of complementary dinucleotides. Profiling of AA-TT symmetry along Chromosome I showed this chromosome to be organized as a succession of five domains that were twofold symmetric for AA-TT, placed between two subtelomeric regions without clear symmetry properties. This pattern was destroyed when ORF sequences were randomly repositioned along the chromosome. Based on the above findings, an architectural model is proposed for Chromosome I, in which the twofold symmetric domains, from 30 to 50 kb long, correspond to chromosome loops.








Similar content being viewed by others
References
Bell SJ, Forsdyke DR (1999a) Accounting units in DNA. J Theor Biol 197:51–61
Bell SJ, Forsdyke DR (1999b) Deviations from Chargaff’s second parity rule correlate with direction of transcription. J Theor Biol 197:63–76
Brewer BJ (1988) When polymerases collide: replication and the transcriptional organization of the E. coli chromosome. Cell 53:679–686
Bradnam KR, Seoighe C, Sharp PM, Wolfe KH (1999) G+C content variation along and among Saccharomyces cerevisiae chromosomes. Mol Biol Evol 16:666–675
Bussey H, Kaback DB, Zhong WW, Vo DT, Clark MW, Fortin N, Hall J, Ouellette BFF, Keng T, Barton AB, Su Y, Davies CJ, Storms RK (1995) The nucleotide sequence of chromosome I from Saccharomyces cerevisiae. Proc Natl Acad Sci USA 92:3809–3813
Calladine CR, Drew HR (1997) Understanding DNA. Academic Press, San Diego
Cebrat S, Dudek MR, Rogowska A (1997) Asymmetry in nucleotide composition of sense and antisense strands as a parameter for discriminating open reading frames as protein coding sequences. J Appl Genet 38:1–9
Fickett JW, Torney DC, Wolf DR (1992) Base compositional structure of genomes. Genomics 13:1056–1064
Forsdyke DR (1995) Relative roles of primary sequences and (G+C)% in determining the hierarchy of frequencies of complementary trinucleotide pairs. J Mol Evol 41:573–581
Francino MP, Chao L, Riley MA, Ochman H (1996) Asymmetries generated by transcription-coupled repair in enterobacterial genes. Science 272:107–109
Gilbert DG (1999) Free software in molecular biology for Macintosh and MS Windows computers. In: Misener S, Krawetz SA (eds) Bioinformatics methods and protocols. Humana Press, Totowa, N.J.
Langkjaer RB, Nielsen ML, Daugaard PR, Liu W, Piskur J (2000) Yeast chromosomes have been significantly reshaped during their evolutionary history. J Mol Biol 304:271–278
Li W, Stolovitzky G, Bernaola-Galvan P, Oliver JL (1998) Compositional heterogeneity within, and uniformity between, DNA sequences of yeast chromosomes. Genome Res 8:916–928
Llorente B, et al (2000) Genomic exploration of the hemiascomycetous yeasts: 18. Comparative analysis of chromosome maps and synteny with Saccharomyces cerevisiae. FEBS Lett 487:101–112
Lobry JR (1995) Properties of a general model of DNA evolution under no-strand bias conditions. J Mol Evol 40:326–330
Lobry JR (1996) Asymmetric substitution patterns in the two DNA strands of bacteria. Mol Biol Evol 13:660–665
Manuelidis L, Chen TL (1990) A unified model of eukaryotic chromosomes. Cytometry 11:8–25
Mewes HW, Frishman D, Güldener U, Mannhaupt D, Mayer K, Mokrejs M, Morgenstern B, Münsterkoetter M, Rudd S, Weil B (2002) MIPS: a database for genomes and protein sequences. Nucleic Acids Res 30:31–4
Prabhu VV (1993) Symmetry observations in long nucleotide sequences. Nucleic Acids Res 21:2797–2800
Raghuraman MK, Winzeler EA, Collingwood D, Hunt S, Wodicka L, Conway A, Lockhart DJ, Davis RW, Brewer BJ, Fangman WL (2001) Replication dynamics of the yeast genome. Science 294:115–121
Rice P, Longden I, Bleasby A (2000) EMBOSS: The European Molecular Biology Open Software Suite. Trends Genet 16:276–277
Rudner R, Karkas JD, Chargaff E (1968) Separation of B. subtilis DNA into complementary strands: III. Direct analysis. Proc Natl Acad Sci USA 60:921–922
Saitoh Y, Laemmli UK (1994) Metaphase chromosome structure: bands arise from a differential folding path of the highly AT-rich scaffold. Cell 76:609–622
Seoighe C, et al (2000) Prevalence of small inversions in yeast gene order evolution. Proc Natl Acad Sci USA 97:14433–14437
Sueoka N (1995) Intrastrand parity rules of DNA base composition and usage biases of synonymous codons. J Mol Evol 40:318–325
Wolfe KH, Shields DC (1997) Molecular evidence for an ancient duplication of the entire yeast genome. Nature 387:708–713
Acknowledgments
I thank J. Navarro for generously programming Cripto
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by E. Cerdà-Olmedo
Rights and permissions
About this article
Cite this article
Conde, J. Twofold symmetries in nucleotide distribution in large domains of Saccharomyces cerevisiae Chromosome I. Mol Genet Genomics 270, 287–295 (2003). https://doi.org/10.1007/s00438-003-0871-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-003-0871-x